Abstract
The global burden of diabetes is rapidly increasing, from 451 million people in 2019 to 693 million by 20451. The insidious onset of type 2 diabetes delays diagnosis and increases morbidity2. Given the multifactorial vascular effects of diabetes, we hypothesized that smartphone-based photoplethysmography could provide a widely accessible digital biomarker for diabetes. Here we developed a deep neural network (DNN) to detect prevalent diabetes using smartphone-based photoplethysmography from an initial cohort of 53,870 individuals (the ‘primary cohort’), which we then validated in a separate cohort of 7,806 individuals (the ‘contemporary cohort’) and a cohort of 181 prospectively enrolled individuals from three clinics (the ‘clinic cohort’). The DNN achieved an area under the curve for prevalent diabetes of 0.766 in the primary cohort (95% confidence interval: 0.750–0.782; sensitivity 75%, specificity 65%) and 0.740 in the contemporary cohort (95% confidence interval: 0.723–0.758; sensitivity 81%, specificity 54%). When the output of the DNN, called the DNN score, was included in a regression analysis alongside age, gender, race/ethnicity and body mass index, the area under the curve was 0.830 and the DNN score remained independently predictive of diabetes. The performance of the DNN in the clinic cohort was similar to that in other validation datasets. There was a significant and positive association between the continuous DNN score and hemoglobin A1c (P ≤ 0.001) among those with hemoglobin A1c data. These findings demonstrate that smartphone-based photoplethysmography provides a readily attainable, non-invasive digital biomarker of prevalent diabetes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the authors and Azumio, but restrictions apply to the availability of these data, which were used under license for the current study, and so are not publicly available. Data are however available from the authors upon reasonable request and with permission of Azumio.
Code availability
The code that supports this work is copyright of the Regents of the University of California and can be made available through license.
References
Cho, N. H. et al. IDF Diabetes Atlas: global estimates of diabetes prevalence for 2017 and projections for 2045. Diabetes Res. Clin. Pract. 138, 271–281 (2018).
Harris, M. I., Klein, R., Welborn, T. A. & Knuiman, M. W. Onset of NIDDM occurs at least 4–7 yr before clinical diagnosis. Diabetes Care 15, 815–819 (1992).
Bertoni, A. G., Krop, J. S., Anderson, G. F. & Brancati, F. L. Diabetes-related morbidity and mortality in a national sample of U.S. elders. Diabetes Care 25, 471–475 (2002).
Allen, J. Photoplethysmography and its application in clinical physiological measurement. Physiol. Meas. 28, R1–R39 (2007).
Elgendi, M. et al. The use of photoplethysmography for assessing hypertension. npj Digit. Med. 2, 60 (2019).
Alty, S. R., Angarita-Jaimes, N., Millasseau, S. C. & Chowienczyk, P. J. Predicting arterial stiffness from the digital volume pulse waveform. IEEE Trans. Biomed. Eng. 54, 2268–2275 (2007).
Otsuka, T., Kawada, T., Katsumata, M. & Ibuki, C. Utility of second derivative of the finger photoplethysmogram for the estimation of the risk of coronary heart disease in the general population. Circ. J. 70, 304–310 (2006).
Smartphone Ownership Is Growing Rapidly around the World, but Not Always Equally (Pew Research Center, 2019).
Coravos, A., Khozin, S. & Mandl, K. D. Developing and adopting safe and effective digital biomarkers to improve patient outcomes. npj Digit. Med. 2, 14 (2019).
Singh, J. P. et al. Association of hyperglycemia with reduced heart rate variability (The Framingham Heart Study). Am. J. Cardiol. 86, 309–312 (2000).
Avram, R. et al. Real-world heart rate norms in the Health eHeart study. npj Dig. Med. 2, 58 (2019).
Carnethon, M. R., Golden, S. H., Folsom, A. R., Haskell, W. & Liao, D. Prospective investigation of autonomic nervous system function and the development of type 2 diabetes. Circulation 107, 2190–2195 (2003).
Guo, Y. et al. Genome-wide assessment for resting heart rate and shared genetics with cardiometabolic traits and type 2 diabetes. J. Am. Coll. Cardiol. 74, 2162–2174 (2019).
Lilia, C.-M. et al. Endothelial dysfunction evaluated using photoplethysmography in patients with type 2 diabetes. J. Cardiovasc. Dis. Diagn. 3, 219 (2015).
Pilt, K., Meigas, K., Ferenets, R., Temitski, K. & Viigimaa, M. Photoplethysmographic signal waveform index for detection of increased arterial stiffness. Physiol. Meas. 35, 2027–2036 (2014).
Schönauer, M. et al. Cardiac autonomic diabetic neuropathy. Diabetes Vasc. Dis. Res. 5, 336–344 (2008).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Gulshan, V. et al. Development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. J. Am. Med. Assoc. 316, 2402–2410 (2016).
Zhang, H. et al. Comparison of physician visual assessment with quantitative coronary angiography in assessment of stenosis severity in China. JAMA Intern. Med. 178, 239–247 (2018).
Hannun, A. Y. et al. Cardiologist-level arrhythmia detection and classification in ambulatory electrocardiograms using a deep neural network. Nat. Med. 25, 1–11 (2019).
Dixit, S. et al. Secondhand smoke and atrial fibrillation: data from the Health eHeart Study. Heart Rhythm 13, 3–9 (2016).
Fawcett, T. An introduction to ROC analysis. Pattern Recognit. Lett. 27, 861–874 (2006).
Noble, D., Mathur, R., Dent, T., Meads, C. & Greenhalgh, T. Risk models and scores for type 2 diabetes: systematic review. Br. Med. J. 343, d7163–d7163 (2011).
Moreno, E. M. et al. Type 2 diabetes screening test by means of a pulse oximeter. IEEE Trans. Biomed. Eng. 64, 341–351 (2017).
Nirala, N., Periyasamy, R., Singh, B. K. & Kumar, A. Detection of type-2 diabetes using characteristics of toe photoplethysmogram by applying support vector machine. Biocybern. Biomed. Eng. 39, 38–51 (2019).
Selvin, E., Steffes, M. W., Gregg, E., Brancati, F. L. & Coresh, J. Performance of A1C for the classification and prediction of diabetes. Diabetes Care 34, 84–89 (2010).
Camacho, J. E., Shah, V. O., Schrader, R., Wong, C. S. & Burge, M. R. Performance of A1C versus OGTT for the diagnosis of prediabetes in a community-based screening. Endocr. Pract. 22, 1288–1295 (2016).
Karakaya, J., Akin, S., Karagaoglu, E. & Gurlek, A. The performance of hemoglobin A1c against fasting plasma glucose and oral glucose tolerance test in detecting prediabetes and diabetes. J. Res. Med. Sci. 19, 1051–1057 (2014).
Pisano, E. D. et al. Diagnostic performance of digital versus film mammography for breast-cancer screening. N. Engl. J. Med. 353, 1773–1783 (2005).
Mathews, W. C., Agmas, W. & Cachay, E. Comparative accuracy of anal and cervical cytology in screening for moderate to severe dysplasia by magnification guided punch biopsy: a meta-analysis. PLoS ONE 6, e24946 (2011).
Use of Glycated Haemoglobin (HbA1c) in the Diagnosis of Diabetes Mellitus: Abbreviated Report of a WHO Consultation (World Health Organization, 2011).
Kim, D.-I. et al. The association between resting heart rate and type 2 diabetes and hypertension in Korean adults. Heart 102, 1757–1762 (2016).
Lindström, J. & Tuomilehto, J. The diabetes risk score: a practical tool to predict type 2 diabetes risk. Diabetes Care 26, 725–731 (2003).
American Diabetes Association Classification and diagnosis of diabetes: standards of medical care in diabetes—2018. Diabetes Care 41, S13–S27 (2018).
Elgendi, M. On the analysis of fingertip photoplethysmogram signals. Curr. Cardiol. Rev. 8, 14–25 (2012).
Emami, S. New methods for computing interpolation and decimation using polyphase decomposition. IEEE Trans. Educ. 42, 311–314 (1999).
Ioffe, S. & Szegedy, C. Batch normalization: accelerating deep network training by reducing internal covariate shift. in Proc. 32nd International Conference on Machine Learning 448–456 (2015).
Nair, V. & Hinton, G. E. Rectified linear units improve restricted Boltzmann machines. in Proc. 27th International Conference on Machine Learning 807–814 (2010).
Srivastava, N., Hinton, G. E., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Goodfellow, I., Bengio, Y. & Courville, A. Deep Learning (MIT Press, 2016).
He, K., Zhang, X., Ren, S. & Sun, J. Delving deep into rectifiers: surpassing human-level performance on ImageNet classification. in Proc. IEEE International Conference on Computer Vision 2015, 1026–1034 (2015).
Liu, L. et al. On the variance of the adaptive learning rate and beyond. Preprint at https://arxiv.org/abs/1908.03265 (2019).
Yosinski, J., Clune, J., Nguyen, A., Fuchs, T. & Lipson, H. Understanding neural networks through deep visualization. Preprint at https://arxiv.org/abs/1506.06579 (2015).
Ferri, C., Hernández-Orallo, J. & Modroiu, R. An experimental comparison of performance measures for classification. Pattern Recognit. Lett. 30, 27–38 (2009).
Glas, A. S., Lijmer, J. G., Prins, M. H., Bonsel, G. J. & Bossuyt, P. M. M. The diagnostic odds ratio: a single indicator of test performance. J. Clin. Epidemiol. 56, 1129–1135 (2003).
Benichou, T. et al. Heart rate variability in type 2 diabetes mellitus: a systematic review and meta-analysis. PLoS ONE 13, e0195166 (2018).
Acknowledgements
We acknowledge A. Markowitz for editorial support made possible by CTSI grant KL2 TR001870. R.A. received support from the “Fonds de la recherche en santé du Québec” (grant 274831). G.H.T. received support from the National Institutes of Health NHLBI K23HL135274. J.E.O., M.J.P. and G.M.M. received support from the National Institutes of Health (U2CEB021881). M.J.P. is partially supported by a PCORI contract supporting the Health eHeart Alliance (PPRN-1306-04709). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
J.E.O., R.A., G.H.T. and K.A. contributed to the study design. P.K., J.E.O., R.A., K.A. and G.H.T. contributed to data collection. R.A. and G.H.T. performed data cleaning and analysis, ran experiments and created tables and figures. R.A., J.E.O., P.K., J.W.H., G.M.M., M.J.P., K.A. and G.H.T. contributed to data interpretation and writing. G.H.T., J.E.O. and K.A. supervised. G.H.T. and K.A. contributed equally as co-senior authors. All authors read and approved the submitted manuscript.
Corresponding author
Ethics declarations
Competing interests
J.E.O. has received research funding from Samsung and iBeat. G.M.M. has received research funding from Medtronic, Jawbone and Eight. K.A. received funding from Jawbone Health Hub. P.K. is an employee of Azumio. G.H.T has received research grants from Janssen Pharmaceuticals and Myokardia and is an advisor to Cardiogram, Inc. None of the remaining authors have potential conflicts of interest. Azumio provided no financial support for this study and provided only access to the data. Data analysis, interpretation and decision to submit the manuscript were performed independently from Azumio.
Additional information
Peer review information Michael Basson was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Baseline characteristics of the primary cohort by diabetes status.
Primary cohort sample size was 53,870 individual people. Where data was only available for subgroups of the full cohort, subgroup sample size is denoted by N. Differences in means of continuous variables between 2 groups were compared using the two-sample t-test. Differences in proportions of categorical variables between 2 groups were compared using the Chi-Squared test. Tests of significance were 2 sided. Abbreviations: bpm: beats per minute; CAD: Coronary artery disease; CHF: Congestive heart failure; COPD: Chronic obstructive pulmonary disease; HR: Heart rate, MI: Myocardial Infarction; PVD: Peripheral Vascular Disease.
Extended Data Fig. 2 Baseline characteristics in the primary cohort training, development and test datasets.
Primary cohort sample size was 53,870 individual people. Where data was only available for subgroups of the full cohort, subgroup sample size is denoted by N. Differences in means of continuous variables between 2 groups were compared using two-sample t-test. Differences in means of continuous variables between 3+ groups were compared using one-way ANOVA. Differences in proportions of categorical variables between the 2+ groups were compared using Chi-Squared. Tests of significance were 2 sided. a, b, c: Each subscript letter denotes a subset of dataset categories whose column proportions do not differ significantly from each other at the 0.05 level. Post-hoc analysis was performed using Fisher’s least significant differences to compare means of continuous variables between groups. Abbreviations: SD: Standard deviation; CAD: Coronary artery disease; CHF: Congestive heart failure; COPD: Chronic obstructive pulmonary disease; HR: Heart rate, MI: Myocardial Infarction; PVD: Peripheral Vascular Disease.
Extended Data Fig. 3 Baseline characteristics of the primary, contemporary and clinic cohorts.
Where data was only available for subgroups of the full cohorts, subgroup sample size is denoted by N. Differences in means of continuous variables between 2 groups were compared using two-sample t-test. Differences in means of continuous variables between 3+ groups were compared using one-way ANOVA. Differences in proportions of categorical variables between the 2+ groups were compared using Chi-Squared. Tests of significance were 2 sided. a, b, c: Each subscript letter denotes a subset of dataset categories whose column proportions do not differ significantly from each other at the 0.05 level. Post-hoc analysis was performed using Fisher’s least significant differences to compare means of continuous variables between groups. Abbreviations: SD: Standard deviation; CAD: Coronary artery disease; CHF: Congestive heart failure; COPD: Chronic obstructive pulmonary disease; HR: Heart rate, MI: Myocardial Infarction; PVD: Peripheral Vascular Disease.
Extended Data Fig. 4 Confusion matrices for DNN performance in three validation datasets.
Confusion matrices for the predictions of the DNN in the Test Dataset (a, b), Contemporary Cohort (c, d), and Clinic Cohort (e, f), at both the recording and user-level. Total number of patients are presented in parentheses. The DNN Score cutoff used was 0.427.
Extended Data Fig. 5 DNN performance to predict diabetes according to time of day, recording length and heart rate in the test dataset.
DNN sensitivity, specificity, diagnostic odds-ratio and AUC to detect prevalent diabetes are presented across strata of age, gender and number of recordings. The Test Dataset sample size is 11,313 individuals. Counts are provided in parentheses for all subgroup metrics. The diagnostic odds-ratio is the ratio of positive likelihood ratio (sensitivity / (1–specificity)) to the negative likelihood ratio ((1–sensitivity)/specificity). The diagnostic odds-ratio is presented at the recording-level with the associated 95% confidence interval. Interaction p-values are two-sided Wald tests for interaction between the DNN Score and the respective covariates for diabetes. Abbreviations: DNN: deep neural network; OR: diagnostic odds ratio; AUC: area under the curve; CI: confidence interval; BPM: beats per minute.
Extended Data Fig. 6 Activation maps from several hidden convolutional layers of the trained deep neural network (DNN) for one photoplethysmography (PPG) record.
a, An example of a PPG recording which serves as the input into the DNN. b, The activation map of one example filter (out of 16) from the first convolutional layer of the neural network. This activation map is obtained after the example PPG recording is fed into the trained DNN. Each lighter colored band illustrates “activation” of a model parameter. At this early layer of the neural network, the lighter colored bands correspond directly to each cardiac cycle of the PPG waveform. Thicker lines likely indicate morphological features of the waveform. c, Visualization of the activation maps of the 16 filters from the first convolutional layer of the neural network, obtained after the input PPG is fed into the trained DNN. Each of the 16 filters can learn different sets of “features” from the input PPG recording. Filters with more purple bands have more inactive neurons, as compared to those with lighter colors (green being the strongest activation and dark purple being the weakest activation). Six filters appear completely inactivated (all purple), suggesting that the features these filters focus on are not present in this example input PPG. d, Visualization of the activation maps of the 7th convolutional layer of the DNN, comprised of 32 filters. Broadly, these activation maps from the 7th layer of the DNN are more complex compared to those from the 1st layer (b, c), demonstrating how deeper layers of the DNN encode increasingly abstract information representing higher level interactions and complex features.
Extended Data Fig. 7 Activation maps from hidden convolutional layers of the trained deep neural network (DNN) for an example photoplethysmography (PPG) recording with artifacts.
a, An example PPG recording with 2 artifacts (blue and orange rectangles) which serves as the input into the DNN. b, Activation maps of the 16 filters from the first convolutional layer of the DNN. Each lighter colored band illustrates “activation” of a model parameter. Orange and blue arrow are placed on filters denoting the location of artifacts, highlighted by orange and blue rectangles (a), respectively. Some filters, such as the 4th image in the top row, seem to not have activation at the location of the artifactual beats (hollow orange and blue arrows), suggesting that the DNN is “ignoring” data from these artifact locations. Whereas other filters are have activation, suggested by lighter color bars, in the locations of the artifacts (full orange and blue arrows), such as the 2nd filter from the left in the top row, suggesting that the DNN is using data from these artifact locations. Some filters, such as the 2nd from the left in the bottom row “ignore” the artifactual beats by having uniform activation throughout the signal length (except where there are artifacts) likely representing the cardiac cycle. These findings suggest that the DNN is able to identify artifactual beats and differentiate them from good quality waveforms.
Extended Data Fig. 8 Example photoplethysmography (PPG) waveforms.
a, Examples of raw PPG recordings from individuals with and without diabetes (red/green recordings, respectively), which serve as inputs to the deep neural network. DNN Scores predicted for each recording are shown. PPG recordings are either cropped or zero-padded to the same fixed length (~20.3 seconds) before being input into the DNN. The “flat line” in three examples is a demonstration of zero-padding shorter records to the fixed length. DNN: Deep Neural Network; ms: milliseconds.
Extended Data Fig. 9 Deep neural network architecture.
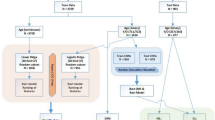
The neural network had 39 layers organized in a block structure, consisting of convolutional layers with an initial filter size of 15 and filter number (N) of 16. The size of the filters decreased, and the number of filters increased as network depth increased, as shown. After each convolutional layer, we applied batch normalization, rectified linear activation and dropout with a probability of 0.2. The final flattened and fully connected softmax layer produced an output distribution across the classes of diabetes/no diabetes. This output distribution is referred to as the DNN Score. PPG: photoplethysmography; DNN: Deep Neural Network; Hz: Hertz.
Supplementary information
Supplementary Information
Supplementary Tables 1–4.
Rights and permissions
About this article
Cite this article
Avram, R., Olgin, J.E., Kuhar, P. et al. A digital biomarker of diabetes from smartphone-based vascular signals. Nat Med 26, 1576–1582 (2020). https://doi.org/10.1038/s41591-020-1010-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-020-1010-5
This article is cited by
-
Dermal features derived from optoacoustic tomograms via machine learning correlate microangiopathy phenotypes with diabetes stage
Nature Biomedical Engineering (2023)
-
Novel Artificial Intelligence Applications in Cardiology: Current Landscape, Limitations, and the Road to Real-World Applications
Journal of Cardiovascular Translational Research (2023)
-
Detection of signs of disease in external photographs of the eyes via deep learning
Nature Biomedical Engineering (2022)
-
A computational framework for discovering digital biomarkers of glycemic control
npj Digital Medicine (2022)
-
Pandemic-proof recruitment and engagement in a fully decentralized trial in atrial fibrillation patients (DeTAP)
npj Digital Medicine (2022)