Abstract
Tumor protein p53 (TP53) is the most frequently mutated gene in cancer1,2. In patients with myelodysplastic syndromes (MDS), TP53 mutations are associated with high-risk disease3,4, rapid transformation to acute myeloid leukemia (AML)5, resistance to conventional therapies6,7,8 and dismal outcomes9. Consistent with the tumor-suppressive role of TP53, patients harbor both mono- and biallelic mutations10. However, the biological and clinical implications of TP53 allelic state have not been fully investigated in MDS or any other cancer type. We analyzed 3,324 patients with MDS for TP53 mutations and allelic imbalances and delineated two subsets of patients with distinct phenotypes and outcomes. One-third of TP53-mutated patients had monoallelic mutations whereas two-thirds had multiple hits (multi-hit) consistent with biallelic targeting. Established associations with complex karyotype, few co-occurring mutations, high-risk presentation and poor outcomes were specific to multi-hit patients only. TP53 multi-hit state predicted risk of death and leukemic transformation independently of the Revised International Prognostic Scoring System (IPSS-R)11. Surprisingly, monoallelic patients did not differ from TP53 wild-type patients in outcomes and response to therapy. This study shows that consideration of TP53 allelic state is critical for diagnostic and prognostic precision in MDS as well as in future correlative studies of treatment response.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Clinical, copy number and mutation data are available at https://github.com/papaemmelab/MDS-TP53-state. The data underlying Figs. 1–4 are provided as Source Data.
Databases used in the study are gnomAD (https://gnomad.broadinstitute.org), COSMIC (https://cancer.sanger.ac.uk/cosmic), cBioPortal for Cancer Genomics (https://www.cbioportal.org), OncoKB Precision Oncology Knowledge Base (https://www.oncokb.org), ClinVar (https://www.ncbi.nlm.nih.gov/clinvar) and the IARC TP53 Database (https://p53.iarc.fr).
Code availability
The NGS-based, allele-specific copy number algorithm CNACS7 is available as a python toil workflow engine at https://github.com/papaemmelab/toil_cnacs, where release v.0.2.0 was used in this study. Source code to reproduce figures from the manuscript is available at https://github.com/papaemmelab/MDS-TP53-state.
Change history
04 May 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41591-021-01367-w
References
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Zehir, A. et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat. Med. 23, 703–713 (2017).
Haase, D. et al. TP53 mutation status divides myelodysplastic syndromes with complex karyotypes into distinct prognostic subgroups. Leukemia 33, 1747–1758 (2019).
Bejar, R. et al. Clinical effect of point mutations in myelodysplastic syndromes. N. Engl. J. Med. 364, 2496–2506 (2011).
Kitagawa, M., Yoshida, S., Kuwata, T., Tanizawa, T. & Kamiyama, R. p53 expression in myeloid cells of myelodysplastic syndromes. Association with evolution of overt leukemia. Am. J. Pathol. 145, 338–344 (1994).
Lindsley, R. C. et al. Prognostic mutations in myelodysplastic syndrome after stem-cell transplantation. N. Engl. J. Med. 376, 536–547 (2017).
Yoshizato, T. et al. Genetic abnormalities in myelodysplasia and secondary acute myeloid leukemia: impact on outcome of stem cell transplantation. Blood 129, 2347–2358 (2017).
Jädersten, M. et al. TP53 mutations in low-risk myelodysplastic syndromes with del(5q) predict disease progression. J. Clin. Oncol. 29, 1971–1979 (2011).
Haferlach, T. et al. Landscape of genetic lesions in 944 patients with myelodysplastic syndromes. Leukemia 28, 241–247 (2014).
Kastenhuber, E. R. & Lowe, S. W. Putting p53 in context. Cell 170, 1062–1078 (2017).
Greenberg, P. L. et al. Revised international prognostic scoring system for myelodysplastic syndromes. Blood 120, 2454–2465 (2012).
Schanz, J. et al. New comprehensive cytogenetic scoring system for primary myelodysplastic syndromes (MDS) and oligoblastic acute myeloid leukemia after MDS derived from an international database merge. J. Clin. Oncol. 30, 820–829 (2012).
Breems, D. A. et al. Monosomal karyotype in acute myeloid leukemia: a better indicator of poor prognosis than a complex karyotype. J. Clin. Oncol. 26, 4791–4797 (2008).
Donehower, L. A. et al. Integrated analysis of TP53 gene and pathway alterations in the cancer genome atlas. Cell Rep. 28, 1370–1384 (2019).
Rucker, F. G. et al. TP53 alterations in acute myeloid leukemia with complex karyotype correlate with specific copy number alterations, monosomal karyotype, and dismal outcome. Blood 119, 2114–2121 (2012).
Papaemmanuil, E. et al. Genomic classification and prognosis in acute myeloid leukemia. N. Engl. J. Med. 374, 2209–2221 (2016).
Sallman, D. A. et al. Impact of TP53 mutation variant allele frequency on phenotype and outcomes in myelodysplastic syndromes. Leukemia 30, 666–673 (2016).
Goel, S. et al. High prevalence and allele burden-independent prognostic importance of p53 mutations in an inner-city MDS/AML cohort. Leukemia 30, 1793–1795 (2016).
Montalban-Bravo, G. et al. Genomic context and TP53 allele frequency define clinical outcomes in TP53-mutated myelodysplastic syndromes. Blood Adv. 4, 482–495 (2020).
Lausen, B. & Schumacher, M. Maximally selected rank statistics. Biometrics 48, 73–85 (1992).
Boettcher, S. et al. A dominant-negative effect drives selection of TP53 missense mutations in myeloid malignancies. Science 365, 599–604 (2019).
Levine, A. J. The many faces of p53: something for everyone. J. Mol. Cell Biol. 11, 524–530 (2019).
Lang, G. A. et al. Gain of function of a p53 hot spot mutation in a mouse model of li-Fraumeni syndrome. Cell 119, 861–872 (2004).
Olive, K. P. et al. Mutant p53 gain of function in two mouse models of Li-Fraumeni syndrome. Cell 119, 847–860 (2004).
Loizou, E. et al. A gain-of-function p53-mutant oncogene promotes cell fate plasticity and myeloid leukemia through the pluripotency factor FOXH1. Cancer Discov. 9, 962–979 (2019).
Wong, T. N. et al. Role of TP53 mutations in the origin and evolution of therapy-related acute myeloid leukaemia. Nature 518, 552–555 (2015).
Platzbecker, U. Treatment of MDS. Blood 133, 1096–1107 (2019).
Roman, E. et al. Myeloid malignancies in the real-world: occurrence, progression and survival in the UK’s population-based Haematological Malignancy Research Network 2004–15. Cancer Epidemiol. 42, 186–198 (2016).
Smith, A. et al. Cohort profile: the Haematological Malignancy Research Network (HMRN); a UK population-based patient cohort. Int. J. Epidemiol. 47, 700–700g (2018).
Welch, J. S. et al. TP53 and decitabine in acute myeloid leukemia and myelodysplastic syndromes. N. Engl. J. Med. 375, 2023–2036 (2016).
Malcovati, L. et al. Clinical significance of SF3B1 mutations in myelodysplastic syndromes and myelodysplastic/myeloproliferative neoplasms. Blood 118, 6239–6246 (2011).
Papaemmanuil, E. et al. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood 122, 3616–3627 (2013). quiz 3699.
International Standing Committee on Human Cytogenetic Nomenclature. ISCN 2013: An International System for Human Cytogenetic Nomenclature (Karger, 2013).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
Ye, K., Schulz, M. H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Karczewski, K. J. et al. Variation across 141,456 human exomes and genomes reveals the spectrum of loss-of-function intolerance across human protein-coding genes. Nature 581, 434–443 (2020).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinformatics 14, 178–192 (2013).
Tate, J. G. et al. COSMIC: the catalogue of somatic mutations in cancer. Nucleic Acids Res. 47, D941–D947 (2019).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Chang, M. T. et al. Identifying recurrent mutations in cancer reveals widespread lineage diversity and mutational specificity. Nat. Biotechnol. 34, 155–163 (2016).
Chang, M. T. et al. Accelerating discovery of functional mutant alleles in cancer. Cancer Discov. 8, 174–183 (2018).
Landrum, M. J. et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 42, D980–D985 (2014).
Chakravarty, D. et al. OncoKB: a precision oncology knowledge base. JCO Precis. Oncol. 2017, 10.1200 (2017).
Grinfeld, J. et al. Classification and personalized prognosis in myeloproliferative neoplasms. N. Engl. J. Med. 379, 1416–1430 (2018).
Bouaoun, L. et al. TP53 variations in human cancers: new lessons from the iarc tp53 database and genomics data. Hum. Mutat. 37, 865–876 (2016).
Giacomelli, A. O. et al. Mutational processes shape the landscape of TP53 mutations in human cancer. Nat. Genet. 50, 1381–1387 (2018).
Talevich, E., Shain, A. H., Botton, T. & Bastian, B. C. CNVkit: genome-wide copy number detection and visualization from targeted DNA sequencing. PLoS Comput. Biol. 12, e1004873 (2016).
Acknowledgements
This work was supported in part by grants from the Celgene Corporation through the MDS Foundation. It was also supported by grants-in-aid from the Japan Agency for Medical Research and Development (AMED) (JP19cm0106501, JP19ck0106250 and 15H05909 (S.O.) and JP18ck0106353 (Y.N.)), from the Japan Society for the Promotion of Science (JSPS) (KAKEN JP26221308, JP19H05656 (S.O.)) and from the Ministry of Education, Culture, Sports, Science and Technology (hp160219 (S.O.)). J.B. and A.P. acknowledge funding from Blood Cancer UK (grant 13042). P.V. was supported by the Austrian Science Fund (grant F4704-B20). M.Y.F. was supported by Italian MIUR-PRIN grants. L.M. was supported by the Associazione Italiana per la Ricerca sul Cancro (AIRC, Milan, Italy) 5 per Mille project (21267 and IG 20125). M.T.V. was supported by AIRC 5 per Mille project (21267). M.T.V. recruited patients through the GROM-L clinical network. E.B. was supported by the Francois Wallace Monahan Fellowship and an EvansMDS Young Investigator award. E.P. is a Josie Robertson Investigator and is supported by the European Hematology Association, the American Society of Hematology, Gabrielle’s Angels Foundation, V Foundation and The Geoffrey Beene Foundation. We thank T. Iraca for logistical support.
Author information
Authors and Affiliations
Contributions
E.B. and E.P. designed the study. E.B. and Y.N. performed statistical analysis. S.D. and E.P. supervised statistical analysis. L.M., B.L.E., R.B., P.L.G., M. Cazzola, E.H.-L., S.O. and E.P. supervised research. P.L.G. and E.P. coordinated the study. L.M., F.S., C.A.C., M. Creignou, U.G., A.A.L., M.J., M.T., O.K., M.Y.F., F.T., R.F.P., V.S., I.K., J.B., F.P.S.S., S.K., T.I., T.H., A.T.-K., T.K., C.P., V.M.K., M.R.S., M.B., C.G., L.P., L.A., M.G.D.P., P.F., A.P., U.P., M.H., P.V., S.C., Y.M., C.F., M.T.V., L.-Y.S., M.F., J.H.J., J.C., Y.A., N.G., M. Cazzola, E.H.-L. and S.O. provided clinical data and DNA specimens. E.B., Y.W., M.P. and E.P. coordinated sample acquisition. A.V. and K.V. performed sample preparation and sequencing. E.B., R.P.H., H.T. and M. Creignou curated clinical data. R.P.H. and J.M.B. performed pathology review. E.B. and H.T. processed cytogenetic data. F.S., D.H. and J.S. performed cytogenetic review. E.B., Y.N., J.S.M.-M., T.Y., A.S. and G.G. performed bioinformatic analysis. J.S.M.-M., M.F.L., J.E.A. and J.Z. supported sequence data pipelines. Y.S. and R.S. developed copy number algorithm CNACS. M.F.L. generated copy number profiles. Y.Z. performed SNP array analysis. E.B. and Y.N. prepared figures and tables. E.B., S.O. and E.P. wrote the manuscript. All authors reviewed the manuscript during its preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interests. U.G. has received honoraria from Celgene, Novartis, Amgen, Janssen, Roche and Jazz and research funding from Celgene and Novartis. C.A.C. has received research funding from Celgene. A.A.L. is in advisory boards of Celgene, Amgen, Roche, Novartis and Alexion and has received research funding from Celgene. F.T. is on the advisory boards of Jazz, Pfizzer and Abbvie and has received research funding from Celgene. I.K. is on the advisory board of Genesis Pharma and has received research funding from Celgene and Janssen Hellas. F.P.S.S. has received honoraria from Janssen-Cilag, Bristol-Myers-Squibb, Novartis, Amgen, Abbvie and Pfizer, is on the advisory boards of Novartis, Amgen and Abbvie and has received research funding from Novartis. A.T.-K. has received honoraria from Novartis, Bristol-Myers-Squibb and MSD and has received research funding from Celgene, Ono Pharmaceutical and Cognano. T.K. has received research funding from Bristol-Myers-Squibb, Otsuka Pharmaceutical, Kyowa Kirin, MSD, Astellas Pharmaceutical, Nippon Shinyaku, Novartis Pharmaceutical, Sumitomo Dainippon Pharmaceutical, Janssen Pharmaceutical, Celgene, SymBio Pharmaceutical, Taiho Pharmaceutical, Tejin, Sanofi K.K. and Celltrion. M.R.S. is on the advisory boards of Abbvie, Astex, Celgene, Karyopharm, Selvita and TG Therapeutic, has equity in Karyopharm and has received research funding from Astex, Incyte, Sunesis, Takeda and TG Therapeutics. G.S. is on the advisory boards of AbbVie, Amgen, Astellas, Böehringer-Ingelheim, Celgene, Helsinn Healthcare, Hoffmann-La Roche, Janssen-Cilag, Novartis and Onconova and has received research funding from Celgene, Hoffmann-La Roche, Janssen-Cilag and Novartis. L.A. is on the advisory boards of Abbvie, Astex, Celgene and Novartis and has received research funding from Celgene. D.S.N. has equity in Madrigal Phamaceuticals and has received research funding from Celgene and Pharmacyclics. K.L.B. has received research funding from GRAIL. M.H. has received honoraria from Novartis, Pfizer and PriME Oncology, is on the advisory boards of Abbvie, Bayer Pharma, Daiichi Sankyo, Novartis and Pfizer and has received institutional research funding from Astellas, Bayer Pharma, BergenBio, Daiichi Sankyo, Karyopharm, Novartis, Pfizer and Roche. P.V. has received honoraria and research funding from Celgene. S.C. has received research funding from Kyowa Kirin, Chugai Pharmaceutical, Takeda Pharmaceutical, Astellas Pharmaceutical, Sanofi KK and Ono Pharmaceutical. Y.M. has received honoraria from Ohtsuka, Novartis, Nippon Shinyaku, Dainippon-Sumitomo and Kyowa Kirin and research funding from Chugai. C.F. is on the advisory boards of, and has received honoraria from, Celgene, Novartis and Janssen and has received research funding from Celgene. M.T.V. is on the advisory board of Celgene, has received honoraria from Celgene and Novartis and has received research funding from Celgene. Y.A. has received honoraria from Mochida, Meiji, Chugai and Kyowa Kirin. N.G. is on the advisory board of, and has received honoraria from, Novartis and has received research funding from Alexion. B.L.E. has received research funding from Celgene and Deerfield. R.B. is on the advisory boards of Celgene, AbbVie, Astex, NeoGenomics and Daiichi Sankyo and has received research funding from Celgene and Takeda. E.H.-L. has received research funding from Celgene. E.B. has received research funding from Celgene. E.P. has received research funding from Celgene and has served on scientific advisory boards for Novartis. E.P. is the founder and CEO of Isabl, a company offering analytics for cancer whole-genome sequencing data.
Additional information
Peer review information Javier Carmona was the primary editor on this article, and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Study cohort characteristics.
Table describing the baseline characteristics of the study cohort. 1Q: first quartile; 3Q: third quartile; OS: overall survival; #: AML classification per WHO 2016 and previously RAEB-T cases. $: Median follow-up time is calculated for censored patients.
Extended Data Fig. 2 Validation cohort characteristics.
Table describing the baseline characteristics of the validation cohort. 1Q: first quartile; 3Q: third quartile; OS: overall survival; $: Median follow-up time is calculated for censored patients.
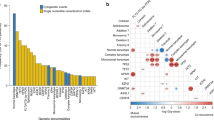
Extended Data Fig. 3 Landscape of chromosomal aberrations in MDS.
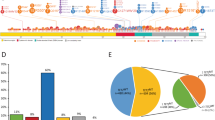
a, Landscape of chromosomal arm-level aberrations across 3,324 patients. Aberrations include copy-neutral loss of heterozygosity (cnloh), deletion (del) and gain. Chromosomes or chromosome arms with more than 5 aberrations are depicted on the x-axis. Aberrations were assessed using the integration of conventional G-banding analysis (CBA) data and NGS derived allele specific copy-number profiles (see Methods). NGS aberrant segments were restricted to segments larger than 3 megabases. b, Frequency distribution of chromosomal aberrations ordered by type of aberrations. First top three plots represent arm-level copy-neutral loss of heterozygosity (cnloh), deletion (del) and gain. Fourth bottom plot represents other types of aberrations to include the presence of marker chromosome (mar), rearrangements where r_i_j denotes a rearrangement between chromosome i and j, isochromosome 17q (iso17q), whole genome amplification (WGA) and presence of ring chromosome (ring). All aberrations observed in more than 3 patients are depicted. Of note, cnloh is detectable with NGS but not with CBA. On the opposite, rearrangements, presence of marker or ring chromosome and WGA were only assessed from CBA data. In 393 cases with missing CBA data, those specific aberrations were imputed from other molecular markers.
Extended Data Fig. 4 Evidence of biallelic TP53 targeting in the cases with multiple TP53 hits.
a, Scatter plot of the two maximum TP53 variant allele frequency (VAF) values from cases with multiple TP53 mutations and no copy-neutral LOH or deletion at TP53 locus (n=90). Points are annotated according to the level of information of the mutation pairs. In 67% (n=60) of pairs the sum of the two VAFs exceeded 50% so that the mutations were considered to be in the same cells as per the pigeonhole principle (triangle and diamond points). In 18 cases, the genomic distance between two mutations was within sequencing read length and it was therefore possible to phase the mutations. In all those cases the mutations were observed to be unphased, that is, in trans (square and diamond points). Within those 18 pairs of unphased mutations, 10 pairs had a sum of VAFs above 50%, that is, mutations were necessarily on different alleles and in the same cells, implying biallelic targeting (diamond points). b, c, Scatter plots of the VAF of TP53 mutations and minor allele frequency of 17p heterozygous SNPs from cases with one TP53 mutation and 17p deletion (b., n=69) or 17p copy-neutral LOH (c., n=61). The high correlations in (a.), (b.) and (c.) (R2 of 0.77, 0. 94 and 0.97, respectively) are indicative of biallelic targeting of TP53. d, Table of pairs of TP53 mutations from the same patients that could be phased. All pairs were in trans, that is, mutations were supported by different alleles. e, Representative IGV example of unphased mutations (patient p12 from table (d.)).
Extended Data Fig. 5 Heatmap of chromosomal aberrations per TP53 allelic state.
Each column represents a patient from the TP53 subgroups of monoallelic mutation (top orange band, 1mut), multiple mutations (top light blue band, >1mut), mutation(s) and deletion (top blue band, mut+del) and mutation(s) and copy-neutral loss of heterozygosity (top dark blue band, mut+cnloh). Aberrations observed at a frequency higher than 2% in either monoallelic or multi-hit TP53 state are depicted on the y-axis. Aberrations include from top to bottom the annotation of complex karyotype (complex), the presence of marker chromosome (mar), deletion (del), gain (plus), rearrangement (with r_i_j rearrangement between chromosome i and j), copy-neutral loss of heterozygosity (cnloh), whole genome amplification (WGA) and the presence of ring chromosome (ring). The deletions of 17p of two cases from the 1mut TP53 subgroup did not affect the TP53 locus.
Extended Data Fig. 6 TP53 allelic state segregates patient outcomes across WHO subtypes and IPSS-R risk groups.
a, Proportion of WHO subtypes per TP53 allelic state of monoallelic mutation (1mut) and multiple hits (multi). t-MDS: therapy-related MDS; SLD: single lineage dysplasia; RS: ring sideroblast; MLD: multiple lineage dysplasia; EB: excess blasts; AML-MRC: AML with myelodysplasia-related changes; U: unclassified. Multi-hit TP53 is enriched for t-MDS compared to monoallelic TP53 state (21% vs. 8%, OR=2.9, p=0.002 two-sided Fisher exact test) and for MDS-EB2 (31% vs. 13%, OR=3.1, p=5x10−5 two-sided Fisher exact test). Contrarily, monoallelic TP53 is enriched for MDS-del5q (15% vs. 2%, OR=8.4, p=6x10-6 two-sided Fisher exact test). b, Proportion of IPSS-R risk groups per TP53 allelic state. Multi-hit TP53 is strongly enriched for the very-poor category compared to monoallelic TP53 state (74% vs. 9%, OR=28, p=2x10-35 two-sided Fisher exact test). c, Kaplan-Meier probability estimates of overall survival (OS) across main WHO subtypes per TP53 allelic state of wild-type TP53 (WT), monoallelic TP53 (1mut) and multiple TP53 hits (multi). WHO subtypes MDS-SLD and MDS-MLD are merged together as MDS-SLD/MLD and WHO subtypes MDS-EB1 and MDS-EB2 are merged together as MDS-EB1/2. d, Kaplan-Meier probability estimates of overall survival across IPSS-R risk groups per TP53 allelic state. IPSS-R very-good and good risk groups are merged together (leftmost panel), and IPSS-R very-poor and poor risk groups are merged together as well (rightmost panel). In (c.) and (d.), annotated p-values are from two-sided log-rank tests and numbers indicate cases with OS data per allelic state.
Extended Data Fig. 7 Outcomes across TP53 subgroups and VAF strata.
a, b, Kaplan-Meier probability estimates of overall survival (a.) and cumulative incidence of AML transformation (AMLt) (b.) across TP53 subgroups of wild-type TP53 (WT), single TP53 mutation (1mut), multiple TP53 mutations (>1mut), TP53 mutation(s) and deletion (mut+del), TP53 mutation(s) and copy-neutral loss of heterozygosity (mut+cnloh). c-d, Kaplan-Meier probability estimates of overall survival (c.) and cumulative incidence of AMLt (d.) per TP53 allelic state and range of variant allele frequency (VAF) of TP53 mutations. Annotated p-values are from two-sided log-rank tests in (a.) and (c.) and from two-sided Gray’s tests in (b.) and (d.). The number of cases with outcome data per group is indicated in parentheses.
Extended Data Fig. 8 Maintained differences in genome instability levels and outcomes between TP53 states per mutation type.
a, Proportion of different types of mutation per TP53 subgroup. Truncated mutations (pink) include frameshift indels, nonsense or nonstop mutations and splice-site variants. Mutations annotated as hotspot (purple) are missense mutations at amino acid positions 273, 248, 220 and 175. Mutations annotated as other-missense (green) are additional missense mutations or inframe indels. Odds ratio and two-sided Fisher’s test p-values for the proportion of truncated versus non-truncated mutations between the multi-hit TP53 subgroups and the monoallelic TP53 subgroup (1mut) are indicated on the right side. b, Number per patient of unique chromosomes other than 17 with aberrations per TP53 subgroup of single gene mutation (1mut), mutation and deletion (mut+del) and mutation and copy-neutral loss of heterozygosity (mut+cnloh) and across mutation types. Note that 5 patients with both several mutations and deletion or cnloh with ambiguity between the mutation type categories have been excluded for this analysis. The number of patients within each category is indicated in parentheses. In boxplots, the median is indicated by the tick horizontal line, and the first and third quartiles by the box edges. The lower and upper whiskers extend from the hinges to the smallest and largest values, respectively, no further than 1.5x the interquartile range from the hinges. Data beyond the whiskers are plotted individually as dots. The annotated p-values are derived from the two-sided Wilcoxon rank-sum test, each compared to the 1mut group within the same mutation type. c. Kaplan-Meier probability estimates of overall survival (OS) per TP53 subgroup across mutation types. Annotated p-values are from two-sided log-rank tests. The number of cases per subgroup with OS data is indicated in parentheses.
Extended Data Fig. 9 Characteristics of treated cohort subsets.
Table describing the baseline characteristics of the subset of patients that i) received hypomethylating agent (HMA), ii) received Lenalidomide in the context of del(5q) or iii) underwent hematopoietic stem cell transplantation (HSCT).
Extended Data Fig. 10 Clinical workflow for the assessment of TP53 allelic state.
Schematic of a simple clinical workflow based on the number of TP53 mutations, the presence or absence of deletion 17p per cytogenetic analysis, and the presence or absence of cnLOH or focal deletion at 17p per NGS based assay or SNP array. Mutations were considered if VAF≥2%. VAF: variant allele frequency; CK: complex karyotype; OS: overall survival; AML: transformation to acute myeloid leukemia.
Supplementary information
Supplementary Information
Supplementary Tables 1–3 and Figs. 1–22.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Rights and permissions
About this article
Cite this article
Bernard, E., Nannya, Y., Hasserjian, R.P. et al. Implications of TP53 allelic state for genome stability, clinical presentation and outcomes in myelodysplastic syndromes. Nat Med 26, 1549–1556 (2020). https://doi.org/10.1038/s41591-020-1008-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-020-1008-z
This article is cited by
-
p53 biology and reactivation for improved therapy in MDS and AML
Biomarker Research (2024)
-
Pharmacological reactivation of p53 in the era of precision anticancer medicine
Nature Reviews Clinical Oncology (2024)
-
TP53-mutated acute myeloid leukemia and myelodysplastic syndrome: biology, treatment challenges, and upcoming approaches
Annals of Hematology (2024)
-
The role of next-generation sequencing in hematologic malignancies
Blood Research (2024)
-
Role of the STING pathway in myeloid neoplasms: a prospero-registered systematic review of principal hurdles of STING on the road to the clinical practice
Medical Oncology (2024)