Abstract
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic has dramatically expedited global vaccine development efforts1,2,3, most targeting the viral ‘spike’ glycoprotein (S). S localizes on the virion surface and mediates recognition of cellular receptor angiotensin-converting enzyme 2 (ACE2)4,5,6. Eliciting neutralizing antibodies that block S–ACE2 interaction7,8,9, or indirectly prevent membrane fusion10, constitute an attractive modality for vaccine-elicited protection11. However, although prototypic S-based vaccines show promise in animal models12,13,14, the immunogenic properties of S in humans are poorly resolved. In this study, we characterized humoral and circulating follicular helper T cell (cTFH) immunity against spike in recovered patients with coronavirus disease 2019 (COVID-19). We found that S-specific antibodies, memory B cells and cTFH are consistently elicited after SARS-CoV-2 infection, demarking robust humoral immunity and positively associated with plasma neutralizing activity. Comparatively low frequencies of B cells or cTFH specific for the receptor binding domain of S were elicited. Notably, the phenotype of S-specific cTFH differentiated subjects with potent neutralizing responses, providing a potential biomarker of potency for S-based vaccines entering the clinic. Overall, although patients who recovered from COVID-19 displayed multiple hallmarks of effective immune recognition of S, the wide spectrum of neutralizing activity observed suggests that vaccines might require strategies to selectively target the most potent neutralizing epitopes.
Similar content being viewed by others
Main
Plasma and peripheral blood mononuclear cells (PBMCs) were collected from a cross-sectional cohort (n = 41) of Australian adults who recovered from mild-to-moderate SARS-CoV-2 infection a median of 32 d (interquartile range (IQR), 28–35 d) after positive PCR test. Demographic and clinical characteristics of patients with COVID-19, and a control cohort of 27 adults recruited before widespread infection in Australia, are reported in Extended Data Fig. 1. To examine the degree to which baseline cross-reactive coronavirus immunity affected SARS-CoV-2 responses, we pre-screened uninfected individuals for antibodies against the spike protein of beta coronavirus HCoV-HKU1 (HKU1) (Extended Data Fig. 2), selecting individuals with the five highest and five lowest plasma titers as controls for the study.
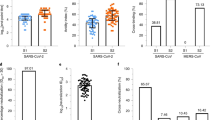
Serological profiles are presented stratified across the cohort based on neutralization activity for each individual. ELISAs were performed using recombinant trimeric S incorporating furin cleavage site removal and stabilizing proline substitutions, or the monomeric receptor-binding domain (RBD) subunit (residues 319–541). Antibodies binding S (Fig. 1a; median endpoint titer, 2,557; IQR, 1,298–7,098) or RBD (Fig. 1b; median, 1,332; IQR, 626–3,445) were observed in all participants with COVID-19 and were highly correlated (Extended Data Fig. 3), with minimal reactivity in controls. Consistent with previous reports15, minimal antibody cross-recognition of the SARS-CoV RBD was observed (Fig. 1c; median, 383; IQR, 226–763). Antibody responses to HKU1 were widely prevalent with endpoint titers generally greater than SARS-CoV-2 titers (Fig. 1d; median, 4,897; IQR, 2,377–8,562), in line with widespread seropositivity to human coronaviruses reported in adults16,17. The capacity of immune plasma to block interaction between recombinant ACE2 and RBD was assessed by ELISA (Methods), with modest inhibition (median, 14.1%; IQR, 10.1–19.3) detected in most subjects, and only 3 of 41 subjects exhibiting more than 50% inhibition activity (Fig. 1e). Although the modest overall signal intensity of this assay might reflect occlusion of some RBD epitopes by plate binding, inhibition activity did track with neutralizing activity (r = 0.53; Extended Data Fig. 3), suggesting that, in most subjects, antibody blockade of ACE2 recognition contributes to neutralization activity. Virus-neutralizing activity in plasma was assessed using a microneutralization assay with live SARS-CoV-2 infection of Vero cells (Methods), as previously described for SARS-CoV and MERS-CoV18,19. Neutralizing antibody titers were generally modest (Fig. 1f; median plasma dilution, 202; IQR, 113–254), consistent with other reports in convalescent cohorts20. S- and RBD-specific antibody responses were consistently elicited by SARS-CoV-2 infection, with serological endpoint titers correlating significantly with neutralizing activity (r = 0.55 and r = 0.61, respectively) and ACE2–RBD binding inhibition (r = 0.72 and r = 0.72, respectively) in the plasma (Extended Data Fig. 3).
a–d, Plasma samples from patients who recovered from SARS-CoV-2 infection (n = 41) and healthy controls (n = 10, black/gray bars) were screened by ELISA for reactivity against S recombinant proteins from SARS-CoV-2 S (a), SARS-CoV-2 RBD (b), SARS-CoV RBD (c) and HCoV-HKU1 (d). e, The capacity of plasma antibodies to inhibit the interaction of RBD with human ACE2 was tested in an ELISA format at a plasma dilution of 1:25. f, Neutralization activity in the plasma was assessed using a microneutralization assay. Data represent individual responses for each individual, and column order for all figures is conserved and stratified on the basis of plasma neutralization activity. SARS-CoV-2-negative donors were stratified based on HKU1 serological titers (black, high titers; gray, low titers).
The frequency and specificity of class-switched (CD19+IgD−) B cells was examined using S or RBD flow cytometric probes (Methods; gating in Supplementary Fig. 1). In contrast to uninfected controls, clear populations of B cells binding spike (S+RBD−), spike and RBD (S+RBD+) or RBD alone (S−RBD+) could be resolved in recovered COVID-19 patients (Fig. 2a and Extended Data Fig. 4) and comprised a respective 0.38% (IQR, 0.24–0.52), 0.047% (IQR, 0.023–0.084) and 0.033% (IQR, 0.015–0.045) of the CD19+IgD− population (Fig. 2b). The very low frequencies of S−RBD+ B cells were likely a mix of background staining and binding to RBD epitopes usually occluded in S. Immunoglobulin isotypes were determined by IgM and IgG surface staining, with IgM−IgG− class-switched B cells previously established as almost exclusively IgA+ (ref. 21). Most S+RBD− B cells were IgG+ (median, 57.5%; IQR, 46.8–64.8), with smaller proportions of IgM+ (20.9%; IQR, 17.4–29.1) and IgA+ (IgM−IgG−, 17.4%; IQR, 13.2–25.9; Fig. 2c). Isotype distribution of S+RBD+ B cells was variable due to low event counts, with median frequencies of 45.5% IgG (IQR, 27.8–70.7), 13.6% IgM (IQR, 0–30.3) and 20.0% IgA (IQR, 4.73–41.9). The activation phenotype of antigen-specific B cells was assessed using CD21 and CD27 (ref. 22) (Extended Data Fig. 5), with S+RBD− (median, 58.5%; IQR, 52.2–66.6) or S+RBD+ (median, 68.7%; IQR, 54.4–80) B cells predominantly exhibiting a resting memory phenotype (CD21+CD27+), consistent with the median time since infection. However, notable proportions of activated memory B cells (CD21−CD27+) were still evident for both S+RBD− (18.9%; IQR, 13.2–25.7) or S+RBD+ populations (13.2%; IQR, 5.88–20), with minimal CD21−CD27− and CD21+CD27− phenotypes observed.
a, Co-staining class-switched B cells (CD19+IgD−) with SARS-CoV-2 spike and RBD probes allows resolution of antigen-specific cells in individuals previously infected with SARS-CoV-2 relative to uninfected controls. b, Frequencies of spike+RBD−, spike+RBD+ and spike+RBD− B cells as a proportion of CD19+IgD− B cells in PBMCs from patients previously infected with SARS-CoV-2 (n = 41) and uninfected controls (n = 10). c, Isotype distribution of spike+RBD− and spike+RBD+ CD19+IgD− B cells; ND, not detected due to absent probe+ cells (n = 41). d, Co-staining spike+CD19+IgD− B cells with SARS-CoV-2 and SARS-CoV RBD probes allows discrimination of cross-reactive specificities versus those unique to SARS-CoV-2. e–g, Frequency of SARS-CoV-2 RBD+/SARS-CoV RBD− cells as a proportion of spike+CD19+IgD−IgG+ B cells (n = 15) (e) and their correlation (Spearmanʼs, two-tailed) to ACE2–RBD binding inhibition (f) and neutralizing activity in plasma (g). b, e, Data are shown as the median with IQR.
The RBD of SARS-CoV and SARS-CoV-2 share significant homology but with marked diversity within the ACE2 binding motif4,23. To better understand individuals with notable RBD-specific antibody titers but limited neutralization or RBD–ACE2 inhibition, we tested differential staining with SARS-CoV and SARS-CoV-2 RBD probes to more precisely quantify B cells recognizing the unique ACE2 binding site of SARS-CoV-2. Cells from a subset of COVID-19 convalescent subjects (n = 15), selected across a range of serum neutralization activities but with prominent RBD-specific B cell populations, were stained as before with S, RBD and SARS-CoV RBD probes (Fig. 2d). Both SARS-CoV-2 RBD-specific and SARS-CoV/SARS-CoV-2 RBD cross-reactive IgG+ B cell populations could be resolved (gating in Supplementary Fig. 2), with the former comprising the major fraction of the S+RBD+ response (Fig. 2e). However, although increased proportions of B cells uniquely binding SARS-CoV-2 RBD does broadly track with plasma blockade of RBD–ACE2 binding (Fig. 2f; r = 0.46), we found no correlation with neutralization activity (Fig. 2g; r = 0.05).
To better understand the immunoglobulins expressed by S- or RBD-specific B cells, we sorted probe+IgG+ B cells from five subjects with high plasma neutralizing titers. A total of 612 (512 S-specific and 100 RBD-specific) productive recombined heavy chain sequences were recovered, 98 (16%) from clonally expanded families, with 302 κ and 89 λ light chain pairs. IGHV, IGKV and IGLV gene use was diverse, with a normal human distribution of CDR-H3 lengths (Extended Data Fig. 6). IGHV somatic mutation of S- and RBD-specific immunoglobulins was similar between subjects, with a median of 1.7% (IQR, 1.0–2.8) and 2.0% (IQR, 1.1−3.4), respectively, in line with other primary infections24,25 but lower than influenza B HA-specific B cells after immunization (6.25%)26 or chronic infections27.
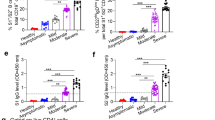
TFH cells provide critical cognate help to antigen-specific B cells to initiate and maintain humoral immune responses28. cTFH cells (CD3+CD4+CD45RA−CXCR5+, gating in Supplementary Fig. 3) in the blood constitute a surrogate of TFH activity in lymphoid tissues29,30. Total unstimulated cTFH frequencies were similar across SARS-CoV-2 convalescent and uninfected donors (Extended Data Fig. 7a). Antigen specificity of the cTFH population was determined using an activation induced marker assay31 after stimulation with S or RBD proteins (Fig. 3a) or overlapping peptide pools (Methods and Extended Data Fig. 7b). Recovered patients exhibited robust cTFH responses to S (median, 0.92% S-specific, cTFH; IQR, 0.42–1.52), whereas RBD-specific cTFH (median, 0.12%; IQR, 0.04–0.38) was significantly less frequent (P < 0.0001) (Fig. 3b). Consistent with high HKU1 seropositivity (Fig. 1d), cTFH responses to HKU1 spike were detected in 97.5% of donors (median, 0.52% of cTFH cells; IQR, 0.32–0.99), with higher frequencies generally observed within COVID-19 convalescent donors than uninfected controls (Fig. 3b), suggestive of boosting of cross-reactive T cells by SARS-CoV-2 infection.
a, Representative staining of CD25 and CD134 co-expression on cTFH (CD3+CD4+CD45RA−CXCR5+) cells after stimulation with 5 μg ml−1 BSA (negative control), SARS-CoV-2 S, SARS-CoV-2 RBD or HKU1 spike protein or SEB (positive control). b, Antigen-specific cTFH (n = 10 SARS-CoV-2− donors, n = 41 SARS-CoV-2+ donors) frequencies were calculated as the proportion of CD25+CD134+ cTFH cells in each stimulation condition after background subtraction using the negative control. c, Representative expression of CCR6 and CXCR3 on bulk cTFH, SARS-CoV-2 spike-specific, HKU1 spike-specific or SEB-responsive (CD25+OX-40+) cTFH. d, Quantification of CCR6+CXCR3−, CCR6+CXCR3+, CCR6−CXCR3+ or CCR6−CXCR3− cTFH populations among SARS-CoV-2+ donors (n = 41). b,d, Data are shown as the median with IQR.
cTFH cells have classically been divided using surface expression of chemokine receptors CCR6 and CXCR3 (refs. 30,32,33). However, the functional capacities of putative cTFH1 (CCR6−CXCR3+), cTFH2 (CCR6−CXCR3−) and TH17-like cTFH (CCR6+CXCR3−) subsets are still unclear and might vary with anatomical or pathogenic context29,30,33. S-specific (median, 52.6%) or HKU1 spike-specific (median, 62.9%) cTFH cells were significantly enriched for CCR6+CXCR3− cells (P < 0.0001; Fig. 3c,d) compared to the unstimulated total cTFH population (median, 21.1%) or SEB stimulated controls. Although TFH31 and cTFH are poor cytokine producers relative to conventional CD4 memory (Tmem) cells (CD3+CD4+CD45RA−CXCR5−) (Extended Data Fig. 7c), CCR6+ and CXCR3+ cTFH exhibited Th17 and Th1-like cytokine profiles, respectively (Extended Data Fig. 7d). We identified an immunogenic peptide derived from S2, LLQYGSFCTQLNRAL, which elicited cTFH responses in three donors (Extended Data Fig. 8a). In each case, the CCR6/CXCR3 phenotype of peptide-specific cTFH was similar to the phenotype of the bulk S2-responding cells (Extended Data Fig. 8b).
Tmem responses were similar to cTFH—namely, strong reactivity to S and HKU1 spike proteins (median, 0.53% and 0.54% of CD4 memory cells, respectively) and lower frequencies of RBD-specific T cells (median, 0.24% of CD4 memory cells) (Extended Data Fig. 9a,b). Consistent with other reports34,35, S-specific Tmem produced interferon (IFN)-γ and tumor necrosis factor (TNF) in response to both S1 and S2 peptide pools, with interleukin (IL)-17 responses approximately ten-fold lower in magnitude (Extended Data Fig. 9c).
Serological neutralization activity will be a critical endpoint for upcoming SARS-CoV-2 vaccine trials. A co-correlation matrix of subject characteristics and immunological parameters highlighted broad co-correlation of immune parameters related to S immunogenicity, including plasma antibody titers and frequencies of S-specific B and cTFH cell populations (Fig. 4a). Notably, neutralization activity positively correlated with S and RBD antibody titers, S- and RBD-specific B cell frequencies and S-specific cTFH (including CCR6−CXCR3+/− cTFH frequencies) but inversely correlated with CCR6+CXCR3− S-specific cTFH. Principal component analysis (PCA) on immunological variables alone revealed clustering of subjects with stronger and weaker plasma neutralization activity (Fig. 4b). Using a multiple regression approach, we identified high titers of S-specific antibody and low proportions of S-specific cTFH with a TH17-like phenotype (CCR6+CXCR3−) as the two most significant predictive factors related to neutralization activity (Fig. 4c). In line with recent reports, we found that neutralizing activity20,36, S-specific antibody20,36 and S-specific cTFH responses, but not cTFH phenotypes, associated with participants reporting greater symptom severity (Extended Data Fig. 10).
a, Co-correlation matrix of characteristics, infection and immunological dynamics in patients who recovered from SARS-CoV-2 infection. Correlations (Spearmanʼs, two-tailed) that did not reach significance of P < 0.05 were not displayed in the matrix. b, PCA of immune parameters showing individuals colored in proportion to plasma neutralization titers. c, Spearmanʼs correlation (two-tailed) of the plasma neutralization activity with S-specific antibody titers and the proportion of CCR6+CXCR3− cells within the S-specific cTFH population (n = 41).
Efficient elicitation of potent neutralizing antibodies is likely to be critical for effective vaccines against SARS-CoV-2. However, despite near universal detection of antibodies, B cells and T cells specific for S and/or RBD in patients with COVID-19 (refs. 20,34,37), we found that neutralization activity in the plasma ranges widely from relatively potent to negligible, consistent with other reports20,36. Similarly, B and T cell responses targeting the RBD, and in particular the ACE2 interaction site, were markedly less frequent than total responses to S, to the point of being undetectable in some participants. Overall, our data suggest that for some individuals, precise antibody recognition and blockade of the RBD–ACE2 binding site is the principal pathway to generating neutralizing antibody. However, the disconnect between plasma neutralizing titers and anti-RBD immunity in many individuals suggests that sufficient non-RBD epitope targets exist to constitute an alternative pathway to similar virus neutralization outcomes.
Direct assessment of key immunological events in lymphoid tissues after immunization is challenging in humans, making surrogate biomarkers of effective immunization in the blood especially informative. Circulating S-specific cTFH elicited by infection correlated with both S-specific B cell and antibody responses, with the limited CD4 T cell epitopes localized to the RBD a potential limitation for RBD-based vaccine strategies. S-specific cTFH exhibits a clear phenotypic bias toward a CCR6+CXCR3− phenotype (cTFH17), previously reported for other viral glycoproteins38 and similarly observed within cTFH specific for HKU1 spike in our cohort. Notably, increased frequencies of both cTFH1 (CCR6−CXCR3+) and cTFH2 (CCR6−CXCR3−) were associated with patients with the highest plasma neutralizing activity. Promotion of cTFH1 responses is well characterized for influenza vaccines21,32,39 and in some chronic infections40,41, where elevated frequencies correlate with robust neutralizing antibody responses. However, the functional significance of CXCR3+ cTFH in SARS-CoV-2 infection is currently unclear. Given the considerable scientific interest in predicting the biogenesis of protective immunity against SARS-CoV-2, our study suggests that B cell and cTFH frequencies and phenotypes could constitute informative biomarkers of immune function for assessment of upcoming clinical trials of new vaccines targeting S. The small cohort size and lack of epitope-level resolution are limitations of the current study. However, future studies in expanded cohorts will facilitate in-depth phenotypic definition of S-specific B and cTFH populations using single-cell sequencing technologies, clarify connections between cTFH and lymph node counterparts and explore interplay between cTFH phenotype and neutralizing outcomes in response to different vaccine platforms and adjuvants.
Methods
Ethics statement
The study protocols were approved by the University of Melbourne Human Research Ethics Committee (2056689), and all associated procedures were carried out in accordance with approved guidelines. All participants provided written informed consent in accordance with the Declaration of Helsinki.
Subject recruitment and sample collection
Patients who had recovered from COVID-19 and healthy controls were recruited through contacts with the investigators and invited to provide a blood sample. Characteristics of SARS-CoV-2 convalescent patients and healthy controls are provided in Extended Data Fig. 1. For all participants, whole blood was collected with sodium heparin anticoagulant. Plasma was collected and stored at −80 °C, and PBMCs were isolated via Ficoll–Paque separation, cryopreserved in 10% DMSO/FCS and stored in liquid nitrogen.
Expression of coronavirus proteins and hACE2
A set of proteins was generated for serological and flow cytometric assays. All unique materials are readily available from the authors. The ectodomain of SARS-CoV-2 (isolate WHU1; residues 1–1,208) or HKU1 S protein (isolate N5; residues 1–1,290) were synthesised with the furin cleavage site removed and P986 and P987 stabilization mutations42, a C-terminal T4 trimerisation domain, Avitag and His-tag, expressed in Expi293 cells and purified by Ni-NTA affinity and size-exclusion chromatography using a Superose 6 16/70 column (GE Healthcare) (Supplementary Fig. 4). SARS-CoV S was biotinylated using BirA (Avidity). The SARS-CoV-2 RBD43 with a C-terminal His-tag (residues 319–541; provided by Florian Krammer) was similarly expressed and purified. SARS-CoV RBD (residues N321–P513) with a C-terminal Avitag and His-tag was expressed in Expi293 cells and purified by Ni-NTA, biotinylated using Bir-A (Avidity) and purified by size-exclusion chromatography using a Superdex 75 (GE Healthcare). The human (residues 19–613) and mouse (residues 19–615) ACE2 ectodomain with C-terminal His-tag (provided by Merlin Thomas) were expressed in Expi293 cells and purified using Ni-NTA and size-exclusion chromatography (Supplementary Fig. 5). Antigenicity of coronaviral proteins was assessed by binding to immune sera, anti-RBD mAbs CR3022 and 4B or human and mouse ACE2 (Supplementary Fig. 6). The glycosylation profile of recombinant S proteins (Supplementary Fig. 6) was assessed using mass spectrometry as previously described44 by SP3 protein clean-up44 and trypsin in-solution digestion. Purified peptides were de-salted and then separated using a two-column chromatography setup comprising a PepMap100 C18 20 mm × 75 μm trap and a PepMap C18 500 mm × 75 μm analytical column on Dionex Ultimate 3000 UPLC (Thermo Fisher). Samples were concentrated onto the trap column at 5 μl min−1 with Buffer A (2% acetonitrile and 0.1% formic acid) for 6 min and infused into a Q-Exactive Plus Mass Spectrometer (Thermo Fisher) at 300 nl min−1 via the analytical column. Next, 125-min gradients were used altering the buffer composition from 2% Buffer B (80% acetonitrile and 0.1% formic acid) to 28% B over 95 min, then from 28% B to 40% B over 10 min and then from 40% B to 100% B over 2 min. The composition was held at 100% B for 3 min, then dropped to 3% B over 5 min and held at 3% B for another 10 min. The Q-Exactive Plus Mass Spectrometer was operated in a data-dependent mode automatically switching between the acquisition of a single Orbitrap MS1 scan (70,000 resolution, AGC of 3 × 106) followed by 15 data-dependent HCD MS2 events (35,000 resolution; stepped NCE 28, 30 and 32; maximum injection time of 125 ms and AGC of 2 × 105) with 30-s dynamic exclusion enabled.
The identification of glycoforms was accomplished using Byonic (Protein Metrics, version 3.5.3 (ref. 45)). MS raw files were searched with an MS1 tolerance of ±5 ppm, and a tolerance of ±20 ppm was allowed for HCD MS2 scans. Searches were performed using cysteine carbamidomethylation as a fixed modification and methionine oxidation as a variable modification, in addition to allowing N-linked glycosylation on asparagine. The default Byonic N-linked glycan database, which is composed of 309 mammalian N-glycans compiled, was used. The protease specificity was set to full trypsin specificity, and a maximum of two miss-cleavage events was allowed. Data were searched against the expected protein sequence. The search was filtered to a 1% protein false discovery rate as set in the Byonic parameters, with the final results filtered to remove glycopeptide assignments with Byonic score below 300 (double the cutoff score suggested by Lee et al.46) to remove low-quality glycopeptide assignments. Data are available via ProteomeXchange with identifier PXD019163.
ELISA
Antibody binding to coronavirus S or RBD proteins was tested by ELISA. Next, 96-well Maxisorp plates (Thermo Fisher) were coated overnight at 4 °C with 2 μg ml−1 recombinant S or RBD proteins. After blocking with 1% FCS in phosphate-buffered saline (FBS), duplicate wells of serially diluted plasma were added and incubated for 2 h at room temperature. Plates were washed before incubation with 1:20,000 dilution of HRP-conjugated anti-human IgG (Sigma) for 1 h at room temperature. Plates were washed and developed using TMB substrate (Sigma), stopped using sulphuric acid and read at 450 nm. Endpoint titers were calculated as the reciprocal serum dilution giving signal 2× background using a fitted curve (4 parameter log regression).
ACE2–RBD inhibition ELISA
An ELISA was performed to measure the ability of plasma antibodies to block interaction between recombinant human ACE2 and RBD proteins. Next, 96-well Maxisorp plates (Thermo Fisher) were coated overnight at 4 °C with 8 µg ml−1 of recombinant RBD protein in carbonate-bicarbonate coating buffer (Sigma). After blocking with PBS containing 1% bovine serum albumin (BSA), duplicate wells of serially diluted plasma (1:25 to 1:1,600) were added and incubated for 1 h at room temperature. Plates were then incubated with 1.5 µg ml−1 of biotinylated recombinant ACE2 protein for 1 h at room temperature followed by incubation with HRP-conjugated streptavidin (Thermo Fisher) for 1 h at room temperature. Plates were developed with TMB substrate (Sigma), stopped with 0.15 M sulphuric acid and read at 450 nm.
Microneutralization assay
SARS-CoV-2 isolate CoV/Australia/VIC01/2020 (ref. 47) was passaged in Vero cells and stored at −80C. Plasma was heat inactivated at 56 °C for 30 min. Plasma was serially diluted 1:20 to 1:10,240 before the addition of 100 TCID50 of SARS-CoV-2 in MEM/0.5% BSA and incubation at room temperature for 1 h. Residual virus infectivity in the plasma/virus mixtures was assessed in quadruplicate wells of Vero cells incubated in serum-free media containing 1 μg ml−1 of TPCK trypsin at 37 °C and 5% CO2; viral cytopathic effect was read on day 5. The neutralizing antibody titer was calculated using the Reed–Muench method, as previously described18,19.
Flow cytometric detection of SARS-CoV-2-reactive B cells
Probes for delineating SARS-CoV-2 S-specific B cells within cryopreserved human PBMCs were generated by sequential addition of streptavidin-phycoerythrin (PE) (Thermo Fisher) to trimeric S protein biotinylated using recombinant Bir-A (Avidity). Biotinylated SARS-CoV RBD was similarly conjugated to streptavidin-BV421 (BD Biosciences). SARS-CoV-2 RBD protein was directly labeled to APC using an APC Conjugation Lightning-Link Kit (Abcam). Cells were stained with Aqua viability dye (Thermo Fisher). Monoclonal antibodies for surface staining included CD19-ECD (J3-119, 1:150) (Beckman Coulter), IgM BUV395 (G20-127, 1:150), CD21 BUV737 (B-ly4, 1:150), IgD PE-Cy7 (IA6-2, 1:500), IgG BV786 (G18-145, 1:75), streptavidin-BV510 (1:600) (BD Biosciences), CD20 APC-Cy7 (2H7, 1:150), CD14 BV510 (M5E2, 1:300), CD3 BV510 (OKT3, 1:600), CD8a BV510 (RPA-T8, 1:1500), CD16 BV510 (3G8, 1:500), CD10 BV510 (HI10a, 1:750) and CD27 BV605 (O323, 1:150) (BioLegend). Cells were washed, fixed with 1% formaldehyde (Polysciences) and acquired on a BD LSR Fortessa or BD Aria II.
Recovery and analysis of B cell receptor sequences
Single S- (S+RBD−) and RBD-specific (S+RBD+) IgG+ B cells were stained as before and sorted using a BD Aria II into 96-well plates, subject to cDNA generation and multiplex PCR and Sanger sequencing, as previously described48. Productive, recombined heavy (V-D-J) and light (V-J) chain immunoglobulin sequences were analyzed using IMGT V-quest49.
Flow cytometric detection of antigen-specific cTFH and CD4 T cells
Cryopreserved human PBMCs were thawed and rested for 4 h at 37 °C. Cells were cultured in 96-well plates at 1 × 106 cells per well and stimulated for 20 h with 5 μg ml−1 of protein (BSA, SARS-CoV-2 S, SARS-CoV-2 RBD and HKU1 S). Selected donors were also stimulated with SEB (5 μg ml−1) as a positive control. After stimulation, cells were washed, stained with Live/Dead blue viability dye (Thermo Fisher) and incubated in a cocktail of monoclonal antibodies: CD27 BUV737 (L128, 1:50), CCR7 Alexa700 (150503, 1:50), CD45RA PE-Cy7 (HI100, 1:200), CD20 BUV805 (2H7, 1:50), CD14 BUV395 (MOP9, 1:100) (BD Biosciences), CD3 BV510 (SK7, 1:100), CD4 BV605 (RPA-T4, 1:100), CD8 BV650 (RPA-T8, 1:400), CD25 APC (BC96, 1:50), OX-40 PerCP-Cy5.5 (ACT35, 1:50), PD-1 BV421 (EH12.2H7, 1:50), CCR6 BV785 (G034E3, 1:100), CXCR3 PE-Dazzle 594 (G02H57, 1:50) (BioLegend) and CXCR5 PE (MU5UBEE, Thermo Fisher, 1:50). Cells were washed, fixed with 1% formaldehyde and acquired on a BD LSR Fortessa using BD FACS Diva.
Cytokine production after stimulation with overlapping peptide pools (15mers overlapping by 11) was measured by intracellular cytokine staining. Thawed PBMCs were rested for 6 h and then stimulated with DMSO, S1 or S2 peptide pool (2 µg ml−1 per peptide), individual peptide LLQYGSFCTQLNRAL (2 µg ml−1) or SEB for 16 h in combination with anti-CD49d/CD28 co-stimulatory antibodies (1:100, BD Biosciences). Golgi block and Golgi plug (BD Biosciences) were added 1 h after stimulation. After stimulation, cells were washed, stained with Live/Dead blue viability dye (Thermo Fisher) and incubated in a cocktail of monoclonal antibodies: CD27 BUV737 (L128, 1:50), CD45RA PE-Cy7 (HI100, 1:200), CD20 BUV805 (2H7, 1:50), CXCR5 BB515 (RF8B2, 1:33) (BD Biosciences); CD3 BV510 (SK7, 1:100), CD4 BV421 (RPA-T4, 1:100), CD8 BV650 (RPA-T8, 1:400), CCR6 BV785 (G034E3, 1:100) and CXCR3 PE-Dazzle 594 (G02H57, 1:50) (BioLegend). Cells were washed, permeabilized with Cytofix/Cytoperm (BD Biosciences) and then stained with IFN-γ APC (B27, 1:100), IL-17A (BL168, 2:25) and TNF APC-Cy7 (Mab11, 1:25) (BioLegend). Cells were washed and acquired on a BD LSR Fortessa using BD FACS Diva.
Statistical analyses
Grouped data are generally presented as median ± IQR, with groups compared by Wilcoxon or Mann–Whitney U tests using Prism 8.0 (GraphPad). Pairwise correlations were assessed using Spearmanʼs tests in Prism 8.0 (GraphPad). A multiple linear regression was used to determine which factors can be used to predict the values of microneutralization across 41 patients. A backward model selection method was used by first fitting a model with all variables of interest and gradually removing one variable with the least significant P value. For each step, the nested F test was used to compare the models (ie, if the removed variable contributed significantly to the fit). This procedure was stopped when a model with only significant predictors was obtained. Confirmatory forward selection methods were also employed, arriving at the same final model. This analysis was performed using the lm function in R (version 3.6.3).
Spearmanʼs correlation matrix was calculated using the rcorr function in R (version 3.6.3). PCA was performed in R (version 3.6.3) using the princomp function by first scaling the variables to have a unit variance. For analysis of disease severity and immunological outputs, a nonparametric Kruskal–Wallis test was used to assess the difference in the mean of the dependent variables (ie, neutralizing activities and immune parameters) across categorical variables (ie, disease severity; all analyses were performed in R (version 3.63)). Throughout the manuscript, significance was defined as P < 0.05.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The mass spectrometry data that support the findings of this study are available via ProteomeXchange with identifier PXD019163. The authors declare that all other data supporting the findings of this study are available in the paper and the Supplementary Information files.
References
Amanat, F. & Krammer, F. SARS-CoV-2 vaccines: status report. Immunity 52, 583–589 (2020).
Thanh, Le,T. et al. The COVID-19 vaccine development landscape. Nat. Rev. Drug Discov. 19, 305–306 (2020).
Lurie, N., Saville, M., Hatchett, R. & Halton, J. Developing Covid-19 vaccines at pandemic speed. N. Engl. J. Med. 382, 1969–1973 (2020).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280 (2020).
Yan, R. et al. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 367, 1444–1448 (2020).
Lan, J. et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581, 215–220 (2020).
Rogers, T. F. et al. Rapid isolation of potent SARS-CoV-2 neutralizing antibodies and protection in a small animal model. Science https://doi.org/10.1126/science.abc7520 (2020).
Wu, Y. et al. A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science 368, 1274–1278 (2020).
Cao, Y. et al. Potent neutralizing antibodies against SARS-CoV-2 identified by high-throughput single-cell sequencing of convalescent patients’ B cells. Cell https://doi.org/10.1016/j.cell.2020.05.025 (2020).
Brouwer, P. J. M. et al. Potent neutralizing antibodies from COVID-19 patients define multiple targets of vulnerability. Science https://doi.org/10.1126/science.abc5902 (2020).
Jiang, S., Hillyer, C. & Du, L. Neutralizing antibodies against SARS-CoV-2 and other human coronaviruses. Trends Immunol. 41, 355–359 (2020).
Gao, Q. et al. Rapid development of an inactivated vaccine candidate for SARS-CoV-2. Science https://doi.org/10.1126/science.abc1932 (2020).
Yu, J. et al. DNA vaccine protection against SARS-CoV-2 in rhesus macaques. Science https://doi.org/10.1126/science.abc6284 (2020).
van Doremalen, N. et al. ChAdOx1 nCoV-19 vaccination prevents SARS-CoV-2 pneumonia in rhesus macaques. Preprint at https://www.biorxiv.org/content/10.1101/2020.05.13.093195v1 (2020).
Lv, H. et al. Cross-reactive antibody response between SARS-CoV-2 and SARS-CoV infections. Cell Rep. 31, 107725 (2020).
Zhou, W., Wang, W., Wang, H., Lu, R. & Tan, W. First infection by all four non-severe acute respiratory syndrome human coronaviruses takes place during childhood. BMC Infect. Dis. 13, 433 (2013).
Okba, N. M. A. et al. Severe acute respiratory syndrome coronavirus 2-specific antibody responses in coronavirus disease 2019 patients. Emerg. Infect. Dis. 26, 1478–1488 (2020).
Subbarao, K. et al. Prior infection and passive transfer of neutralizing antibody prevent replication of severe acute respiratory syndrome coronavirus in the respiratory tract of mice. J. Virol. 78, 3572–3577 (2004).
Houser, K. V. et al. Prophylaxis with a Middle East respiratory syndrome coronavirus (MERS-CoV)-specific human monoclonal antibody protects rabbits from MERS-CoV infection. J. Infect. Dis. 213, 1557–1561 (2016).
Robbiani, D. F. et al. Convergent antibody responses to SARS-CoV-2 in convalescent individuals. Nature https://doi.org/10.1038/s41586-020-2456-9 (2020).
Koutsakos, M. et al. Circulating TFH cells, serological memory, and tissue compartmentalization shape human influenza-specific B cell immunity. Sci. Trans. Med. 10, eaan8405 (2018).
Lau, D. et al. Low CD21 expression defines a population of recent germinal center graduates primed for plasma cell differentiation. Sci. Immunol. 2, eaai8153 (2017).
Ou, X. et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat. Commun. 11, 1620 (2020).
Stettler, K. et al. Specificity, cross-reactivity, and function of antibodies elicited by Zika virus infection. Science 353, 823–826 (2016).
Godoy-Lozano, E. E. et al. Lower IgG somatic hypermutation rates during acute dengue virus infection is compatible with a germinal center-independent B cell response. Genome Med. 8, 23 (2016).
Liu, Y. et al. Cross-lineage protection by human antibodies binding the influenza B hemagglutinin. Nat. Commun. 10, 324 (2019).
Wu, X. et al. Focused evolution of HIV-1 neutralizing antibodies revealed by structures and deep sequencing. Science 333, 1593–1602 (2011).
Crotty, S. T follicular helper cell biology: a decade of discovery and diseases. Immunity 50, 1132–1148 (2019).
Brenna, E. et al. CD4+ T follicular helper cells in human tonsils and blood are clonally convergent but divergent from non-Tfh CD4+ cells. Cell Rep. 30, 137–152 (2020).
Morita, R. et al. Human blood CXCR5+CD4+ T cells are counterparts of T follicular cells and contain specific subsets that differentially support antibody secretion. Immunity 34, 108–121 (2011).
Dan, J. M. et al. A cytokine-independent approach to identify antigen-specific human germinal center T follicular helper cells and rare antigen-specific CD4+ T cells in blood. J. Immunol. 197, 983–993 (2016).
Bentebibel, S. E. et al. Induction of ICOS+CXCR3+CXCR5+ TH cells correlates with antibody responses to influenza vaccination. Sci. Transl. Med. 5, 176ra132 (2013).
Koutsakos, M., Nguyen, T. H. O. & Kedzierska, K. With a little help from T follicular helper friends: humoral immunity to influenza vaccination. J. Immunol. 202, 360–367 (2019).
Grifoni, A. et al. Targets of T cell responses to SARS-CoV-2 coronavirus in humans with COVID-19 disease and unexposed individuals. Cell 181, 1489–1501 (2020).
Weiskopf, D. et al. Phenotype of SARS-CoV-2-specific T-cells in COVID-19 patients with acute respiratory distress syndrome. Sci. Immunol. 26, eabd2071 (2020).
Wu, F. et al. Neutralizing antibody responses to SARS-CoV-2 in a COVID-19 recovered patient cohort and their implications. Preprint at https://www.medrxiv.org/content/10.1101/2020.03.30.20047365v2 (2020).
Braun, J. et al. Presence of SARS-CoV-2 reactive T cells in COVID-19 patients and healthy donors. Preprint at https://www.medrxiv.org/content/10.1101/2020.03.30.20047365v2 (2020).
Farooq, F. et al. Circulating follicular T helper cells and cytokine profile in humans following vaccination with the rVSV-ZEBOV Ebola vaccine. Sci. Rep. 6, 27944 (2016).
Bentebibel, S. E. et al. ICOS+PD−1+CXCR3+ T follicular helper cells contribute to the generation of high-avidity antibodies following influenza vaccination. Sci. Rep. 6, 26494 (2016).
Niessl, J. et al. Persistent expansion and Th1-like skewing of HIV-specific circulating T follicular helper cells during antiretroviral therapy. EBioMedicine 54, 102727 (2020).
Zhang, J. et al. Circulating CXCR3+ Tfh cells positively correlate with neutralizing antibody responses in HCV-infected patients. Sci. Rep. 9, 10090 (2019).
Wrapp, D. et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367, 1260–1263 (2020).
Amanat, F. et al. A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat. Med. https://doi.org/10.1038/s41591-020-0913-5 (2020).
Hughes, C. S. et al. Single-pot, solid-phase-enhanced sample preparation for proteomics experiments. Nat. Protoc. 14, 68–85 (2019).
Bern, M., Kil, Y. J. & Becker, C. Byonic: advanced peptide and protein identification software. Curr. Protoc. Bioinforma. 13, 13.20 (2012).
Lee, L. Y. et al. Toward automated N-glycopeptide identification in glycoproteomics. J. Proteome Res. 15, 3904–3915 (2016).
Caly, L. et al. Isolation and rapid sharing of the 2019 novel coronavirus (SARS-CoV-2) from the first patient diagnosed with COVID-19 in Australia. Med. J. Aust. 212, 459–462 (2020).
Tiller, T. et al. Efficient generation of monoclonal antibodies from single human B cells by single cell RT–PCR and expression vector cloning. J. Immunol. Methods 329, 112–124 (2008).
Brochet, X., Lefranc, M. P. & Giudicelli, V. IMGT/V-QUEST: the highly customized and integrated system for IG and TR standardized V-J and V-D-J sequence analysis. Nucleic Acids Res. 36, W503–W508 (2008).
Acknowledgements
We thank the generous participation of the trial individuals for providing samples. The SARS-CoV-2 RBD expression plasmids were kindly provided by F. Krammer (Icahn School of Medicine at Mt. Sinai). The human and mouse ACE2 expression plasmids were kindly provided by M. Thomas (Monash University). We acknowledge V. Jameson and staff at the Melbourne Cytometry Platform (Melbourne Brain Centre node) for provision of flow cytometry services. We thank the Melbourne Mass Spectrometry and Proteomics Facility of the Bio21 Molecular Science and Biotechnology Institute at the University of Melbourne for the support of mass spectrometry analysis. This study was supported by the Victorian Government and Medical Research Future Fund (MRFF) GNT2002073 (to W.-H.T., D.I.G., M.P.D., S.J.K. and A.K.W.), the ARC Centre of Excellence in Convergent Bio-Nano Science and Technology (to S.J.K.), National Health and Medical Research Council (NHMRC) program grant APP1149990 (to S.J.K. and M.P.D.), NHMRC project grant GNT1162760 (to A.K.W), the Jack Ma Foundation (to D.I.G., N.A.G. and K.S.) and the A2 Milk Company (to K.S.). W-H.T. is a Howard Hughes Medical Institute–Wellcome Trust International Research Scholar (208693/Z/17/Z). J.A.J., D.I.G., M.P.D., W.-H.T., S.J.K. and A.K.W. are supported by NHMRC fellowships. The Melbourne WHO Collaborating Centre for Reference and Research on Influenza is supported by the Australian Government Department of Health.
Author information
Authors and Affiliations
Contributions
J.A.J., H.-X.T., W.S.L., S.J.K. and A.K.W. designed the study and experiments. J.A.J., H.-X.T., W.S.L., A.R., H.G.K., K.W., R.E., H.E.K, C.J.B., F.L.M., N.A.G., P.P., M.H.D., N.E.S. and A.K.W. performed experiments. W.-H.T., N.A.G., D.I.G. and K.S. contributed unique reagents. J.A.J., H.-X.T., W.S.L., A.R., K.S., M.P.D., S.J.K. and A.K.W. analyzed the experimental data. J.A.J., H.-X.T., W.S.L., A.R., M.P.D., S.J.K. and A.K.W. wrote the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Saheli Sadanand was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
Demographics and clinical characteristics of recruited COVID-19 patients and healthy control subjects.
Extended Data Fig. 2 Pre-screen of HCoV-HKU1 serum endpoint titres among a cohort of healthy subjects (N = 27) bled prior to the SARS-CoV-2 pandemic.
Data is shown as the median.
Extended Data Fig. 3 Correlations between antibody binding titres, ACE2/RBD binding inhibition and neutralization activity in plasma from subjects recovered from SARS-CoV-2 infection.
a, Spearman’s Correlation between endpoint titres of S-specific plasma antibody and the extent of ACE2/RBD binding inhibition, plasma neutralization titres or endpoint titres of RBD-specific plasma antibody (N = 41). b, Correlation between endpoint titres of RBD-specific plasma antibody and the extent of ACE2/RBD binding inhibition or plasma neutralziation activity (N = 41). c, Correlation between plasma ACE2/RBD binding inhibition and neutralization activity (N = 41). All correlations are two-tailed.
Extended Data Fig. 4 Representative staining of S- and RBD-specific IgD-IgG+ B cells.
3 uninfected subjects (left panels) and 6 subjects after recovery from SARS-CoV-2 infection (middle and right panels). CD19+IgD-IgG+ B cells cells were identified using gating strategy shown in Supplementary Fig. 1. Binding to SARS-CoV-2 spike and/or SARS-CoV-2 RBD probes was assessed.
Extended Data Fig. 5 Memory B cell phenotypes in subjects after SARS-CoV-2 infection.
a, b, Representative memory B cell phenotypes identified by CD21 and CD27 co-stain of probe+CD19+IgD- cells (blue) overlaid on CD19+IgD- cells (grey) (a) and the corresponding frequencies of the four populations in subjects previously infected with SARS-CoV-2 (b) (Resting memory—CD21+CD27+; activated memory—CD21−CD27+; naïve/CD27lo memory—CD21+CD27−; atypical B cells—CD21−CD27−); n.d—not detected due to absent probe+ cells. Memory B cell phenotypes were identified using gating strategy shown in Supplementary Fig. 2.
Extended Data Fig. 6 Analysis of V gene distribution, somatic mutation and CDR-H3 lengths for S- and RBD-specific BCR sequences.
a,b,c, Frequency distributions of human IGHV (a), IGKV (b) and IGLV (c) genes from BCR sequences recovered from S- (red) and RBD-binding (blue) IgG+ B cells from recovered COVID-19 patients (N = 5) with reference to distribution in cord blood (grey)49. Few IGLV sequences for RBD-binding B cells were recovered and are not shown. d, e, CDR-H3 (d) and somatic mutation (e) in S- (red) and RBD-binding (blue) IgG+ B cells compared to influenza B HA-specific IgG+ B cells recovered after immunisation23. f, Somatic mutation in S- (red) and RBD-binding (blue) IgG+ B cells for each individual subject. d,e,f, Data are shown as the median.
Extended Data Fig. 7 cTFH populations in SARS-CoV-2 positive and negative donors.
a, cTFH frequency (as a proportion of CD4+CD45RA– cells) among SARS-CoV-2 negative (n = 10) and SARS-CoV-2 positive (n = 39) donors. b, Frequency of antigen-specific cTFH following stimulation with peptide pools spanning S1 (without RBD peptides), S2 or RBD regions of SARS-CoV-2 S in SARS-CoV-2 positive donors (n = 20). c, Frequency of IFNγ+, IL-17+ or TNF+ cells following SEB stimulation among cTFH or Tmem (CD4+CD45RA–CXCR5−) populations in SARS-CoV-2 positive donors (n = 20; Wilcoxon two-tailed test). d, CCR6 and CXCR3 expression among bulk cTFH (grey), IFNγ+ (blue) or IL-17+ (red) cTFH cells following SEB stimulation in SARS-CoV-2 positive donors (n = 20). a, b, d, Data are shown as the median with interquartile range.
Extended Data Fig. 8 Peptide-specific cTFH responses in SARS-CoV-2 positive donors.
a, A 15-mer peptide derived from SARS-CoV-2 S2, LLQYGSFCTQLNRAL, was found to be immunogenic in 3 SARS-CoV-2 positive subjects (CP39, CP02, CP36). CP12 shown as representative non-responder. b, Plots indicate the CCR6/CXCR3 expression among S2 peptide pool- or peptide-specific cTFH for each donor.
Extended Data Fig. 9 CD4+ Tmem responses in COVID patients and controls.
a, Representative staining of CD134 and CD25 expression following protein or SEB stimulation among CD4 Tmem (CD4+CD45RA-CXCR5-) cells. b, Background-subtracted frequencies of SARS-CoV-2 S, SARS-CoV-2 RBD or HCoV HKU1 spike-specific CD4 Tmem cells among SARS-CoV-2 negative (n = 10) or SARS-CoV-2 positive (n = 39) donors. c, IFNγ, IL-17 and TNF responses following stimulation with overlapping peptide pools covering SARS-CoV-2 S1 or S2 domains, or SEB in SARS-CoV-2 positive donors (n = 20). d, Graphs indicate background-subtracted responses; responses not above background are indicated as 0.001% (n = 20). b, d, Data are shown as the median with interquartile range.
Extended Data Fig. 10 Association of immunological features with disease severity.
a, Analysis of the relationship between disease severity (mild, moderate or severe) and immunological parameters within the SARS-CoV-2 convalescent cohort. p < 0.05 indicated in bold (Kruskal-Wallis test, two-tailed). b, Relationship between cTFH, S-specific antibody titre, or microneutralization (MN) titre with disease severity (n = 26 mild, n = 10 moderate, n = 5 severe). Data are shown as the median with interquartile range.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6
Rights and permissions
About this article
Cite this article
Juno, J.A., Tan, HX., Lee, W.S. et al. Humoral and circulating follicular helper T cell responses in recovered patients with COVID-19. Nat Med 26, 1428–1434 (2020). https://doi.org/10.1038/s41591-020-0995-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-020-0995-0
This article is cited by
-
Primary prophylaxis with mTOR inhibitor enhances T cell effector function and prevents heart transplant rejection during talimogene laherparepvec therapy of squamous cell carcinoma
Nature Communications (2024)
-
Measurement of IFN-γ and IL-2 for the assessment of the cellular immunity against SARS-CoV-2
Scientific Reports (2024)
-
Impact of the immune profiles of hypertensive patients with and without obesity on COVID-19 severity
International Journal of Obesity (2024)
-
Efficacy of COVID-19 mRNA vaccination in patients with autoimmune disorders: humoral and cellular immune response
BMC Medicine (2023)
-
T cells in health and disease
Signal Transduction and Targeted Therapy (2023)