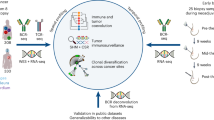
Abstract
Disruption of systemic homeostasis by either chronic or acute stressors, such as obesity1 or surgery2, alters cancer pathogenesis. Patients with cancer, particularly those with breast cancer, can be at increased risk of cardiovascular disease due to treatment toxicity and changes in lifestyle behaviors3,4,5. While elevated risk and incidence of cardiovascular events in breast cancer is well established, whether such events impact cancer pathogenesis is not known. Here we show that myocardial infarction (MI) accelerates breast cancer outgrowth and cancer-specific mortality in mice and humans. In mouse models of breast cancer, MI epigenetically reprogrammed Ly6Chi monocytes in the bone marrow reservoir to an immunosuppressive phenotype that was maintained at the transcriptional level in monocytes in both the circulation and tumor. In parallel, MI increased circulating Ly6Chi monocyte levels and recruitment to tumors and depletion of these cells abrogated MI-induced tumor growth. Furthermore, patients with early-stage breast cancer who experienced cardiovascular events after cancer diagnosis had increased risk of recurrence and cancer-specific death. These preclinical and clinical results demonstrate that MI induces alterations in systemic homeostasis, triggering cross-disease communication that accelerates breast cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data generated are included in the article and in its Supplementary Information. Gene-expression data that support the findings of this study have been deposited in the Gene Expression Omnibus under accession number GSE137790. All data are also available from the corresponding authors on reasonable request.
References
Quail, D. F. & Dannenberg, A. J. The obese adipose tissue microenvironment in cancer development and progression. Nat. Rev. Endocrinol. 15, 139–154 (2019).
Krall, J. A. et al. The systemic response to surgery triggers the outgrowth of distant immune-controlled tumors in mouse models of dormancy. Sci. Transl. Med. 10, eaan3464 (2018).
Hershman, D. L. et al. Association of cardiovascular risk factors with cardiac events and survival outcomes among patients with breast cancer enrolled in SWOG clinical trials. J. Clin. Oncol. 36, 2710–2717 (2018).
Jones, L. W., Haykowsky, M. J., Swartz, J. J., Douglas, P. S. & Mackey, J. R. Early breast cancer therapy and cardiovascular injury. J. Am. Coll. Cardiol. 50, 1435–1441 (2007).
Hooning, M. J. et al. Long-term risk of cardiovascular disease in 10-year survivors of breast cancer. J. Natl Cancer Inst. 99, 365–375 (2007).
McAllister, S. S. & Weinberg, R. A. The tumour-induced systemic environment as a critical regulator of cancer progression and metastasis. Nat. Cell Biol. 16, 717–727 (2014).
Libby, P., Nahrendorf, M. & Swirski, F. K. Leukocytes link local and systemic inflammation in ischemic cardiovascular disease: an expanded “Cardiovascular Continuum”. J. Am. Coll. Cardiol. 67, 1091–1103 (2016).
Engblom, C., Pfirschke, C. & Pittet, M. J. The role of myeloid cells in cancer therapies. Nat. Rev. Cancer 16, 447–462 (2016).
Shipp, C., Speigl, L., Janssen, N., Martens, A. & Pawelec, G. A clinical and biological perspective of human myeloid-derived suppressor cells in cancer. Cell. Mol. Life Sci. 73, 4043–4061 (2016).
Ugel, S., De Sanctis, F., Mandruzzato, S. & Bronte, V. Tumor-induced myeloid deviation: when myeloid-derived suppressor cells meet tumor-associated macrophages. J. Clin. Invest. 125, 3365–3376 (2015).
Nahrendorf, M. et al. The healing myocardium sequentially mobilizes two monocyte subsets with divergent and complementary functions. J. Exp. Med. 204, 3037–3047 (2007).
Biswas, S. et al. CXCL13-CXCR5 co-expression regulates epithelial to mesenchymal transition of breast cancer cells during lymph node metastasis. Breast Cancer Res. Treat. 143, 265–276 (2014).
Panse, J. et al. Chemokine CXCL13 is overexpressed in the tumour tissue and in the peripheral blood of breast cancer patients. Br. J. Cancer 99, 930–938 (2008).
Ginestier, C. et al. CXCR1 blockade selectively targets human breast cancer stem cells in vitro and in xenografts. J. Clin. Invest. 120, 485–497 (2010).
Hohl, T. M. et al. Inflammatory monocytes facilitate adaptive CD4 T cell responses during respiratory fungal infection. Cell Host Microbe 6, 470–481 (2009).
Bronte, V. et al. Recommendations for myeloid-derived suppressor cell nomenclature and characterization standards. Nat. Commun. 7, 12150 (2016).
Nam, S. et al. Interferon regulatory factor 4 (IRF4) controls myeloid-derived suppressor cell (MDSC) differentiation and function. J. Leukoc. Biol. 100, 1273–1284 (2016).
Netea, M. G. et al. Trained immunity: a program of innate immune memory in health and disease. Science 352, aaf1098 (2016).
Christ, A. et al. Western diet triggers NLRP3-dependent innate immune reprogramming. Cell 172, 162–175 (2018).
Kurotaki, D. et al. Transcription factor IRF8 governs enhancer landscape dynamics in mononuclear phagocyte progenitors. Cell Rep. 22, 2628–2641 (2018).
Waight, J. D. et al. Myeloid-derived suppressor cell development is regulated by a STAT/IRF-8 axis. J. Clin. Invest. 123, 4464–4478 (2013).
Mitroulis, I. et al. Modulation of myelopoiesis progenitors is an integral component of trained immunity. Cell 172, 147–161 (2018).
Franklin, R. A. et al. The cellular and molecular origin of tumor-associated macrophages. Science 344, 921–925 (2014).
Nelson, E. R. et al. 27-Hydroxycholesterol links hypercholesterolemia and breast cancer pathophysiology. Science 342, 1094–1098 (2013).
Peinado, H. et al. Pre-metastatic niches: organ-specific homes for metastases. Nat. Rev. Cancer 17, 302–317 (2017).
Caan, B. et al. Life after cancer epidemiology (LACE) study: a cohort of early stage breast cancer survivors (United States). Cancer Causes Control 16, 545–556 (2005).
Kwan, M. L. et al. The pathways study: a prospective study of breast cancer survivorship within Kaiser Permanente Northern California. Cancer Causes Control 19, 1065–1076 (2008).
Hasin, T. et al. Heart failure after myocardial infarction is associated with increased risk of cancer. J. Am. Coll. Cardiol. 68, 265–271 (2016).
Hasin, T. et al. Patients with heart failure have an increased risk of incident cancer. J. Am. Coll. Cardiol. 62, 881–886 (2013).
Meijers, W. C. et al. Heart failure stimulates tumor growth by circulating factors. Circulation 138, 678–691 (2018).
Wendeln, A. C. et al. Innate immune memory in the brain shapes neurological disease hallmarks. Nature 556, 332–338 (2018).
Moorlag, S., Roring, R. J., Joosten, L. A. B. & Netea, M. G. The role of the interleukin-1 family in trained immunity. Immunol. Rev. 281, 28–39 (2018).
Cheng, S. C. et al. Broad defects in the energy metabolism of leukocytes underlie immunoparalysis in sepsis. Nat. Immunol. 17, 406–413 (2016).
Amend, S., Valkenburg, K. C. & Pienta, K. J. Murine hind limb long bone dissection and bone marrow isolation. J. Vis. Exp. 14, 53936 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Ramkhelawon, B. et al. Netrin-1 promotes adipose tissue macrophage retention and insulin resistance in obesity. Nat. Med. 20, 377–384 (2014).
Cesano, A. nCounter((R)) PanCancer immune profiling panel (NanoString Technologies, Inc., Seattle, WA). J. Immunother. Cancer 3, 42 (2015).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
Tissue sectioning and histological analyses were provided by the New York University (NYU) Langone’s Experimental Pathology Research Laboratory. Cell sorting/flow cytometry technologies were provided by NYU Langone’s Cytometry and Cell Sorting Laboratory, which is supported in part by grant P30CA016087 from the National Institutes of Health/National Cancer Institute. This work was supported by funding from the National Institutes of Health (R35HL135799 to K.J.M., P01HL131478 and P01HL131481 to K.J.M. and E.A.F., T32HL098129 to C.v.S., K23HL125991 to J.D.N., R01CA234025 to E.R.N., R01CA129059 to B.J.C. and R01HL132073 to D.S.P.); NYU Cancer Institute Center Support Grant NCIP30CA16087; NYU Shared Instrumentation Grant S10 OD021747; the Memorial Sloan Kettering Cancer Center Support Grant/Core Grant (P30 CA008748)), the American Heart Association (19CDA34630066 to C.v.S., 19POST34380010 to M.S., 18CDA34110203 to T.J.B. and 20POST35080180 to N.Y.), the Canadian Institutes of Health Research (Doctoral Foreign Study Award to G.J.K. and PJT159742 to D.F.Q.), AKTIV Against Cancer (L.W.J.) and the Susan G. Komen Foundation (CCR18548032 to D.F.Q.). D.F.Q. is also supported by the Brain Tumor Funders’ Collaborative, Canada Foundation for Innovation and a Tier II Canada Research Chair in tumor microenvironment research.
Author information
Authors and Affiliations
Contributions
G.J.K., L.W.J. and K.J.M. conceptualized the study; G.J.K., A.A.C.N., E.M.C., C.v.S., M.S.A., M.S., M.S., L.S., T.J.B., K.R., N.Y., D.N. and V.M. performed investigative studies; E.J.B., K.A., J.D.N. and B.J.C. performed data analysis; L.W.J., E.A.F., D.S.P., D.F.Q. and E.R.N. provided input on study design and interpretation; K.J.M. supervised the study; G.J.K. and K.J.M. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Michael Basson was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cardiac function is not altered by the presence of cancer following surgical MI.
(a) Representative triphenyl-tetrazolium chloride (TTC) staining to confirm surgical myocardial infarction through ligation of the left anterior descending coronary artery. (b) Echocardiography examination 16 days following MI or sham surgery (19 days post tumor implantation). Values are mean ± SD. BW Body weight, LV area d, left ventricular area at diastole; LV area s, left ventricular area at systole; EF, ejection fraction; LV volume d, left ventricular volume at diastole; LV volume s, left ventricular volume at systole; SV, stroke volume; FS, longitudinal fractional shortening; CI, cardiac index; LVAWd, left ventricular anterior wall thickness at diastole; LVPWd, left ventricular posterior wall thickness at diastole; LV mass cor, left ventricular mass corrected. P values in data determined as non-parametric were analyzed by a two-tailed Mann–Whitney test, while data determined to be parametric were analyzed by a two-tailed unpaired Student’s t-test; *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001 for sham (no tumor vs tumor) and MI (no tumor vs tumor) comparisons only.
Extended Data Fig. 2 Proliferation of immune (CD45+) and non-immune (CD45-) cells in tumors of mice following MI or sham surgery.
Quantification of Ki67 and CD45 co-staining of tumors to detect proliferating CD45 a, and CD45+ b, cells in the tumor border (n = 5/group). Data are the mean ± s.e.m and P values determined using a two-tailed unpaired Student’s t-test.
Extended Data Fig. 3 Flow cytometric analysis of intratumoral immune cells following MI or sham surgery.
a, Flow cytometry gating strategy for myeloid (top) and lymphoid (bottom) cells in E0771 tumors. b, Flow cytometric analysis showing the relative fold change in immune cell proportions in tumors from mice exposed to MI (n = 11, red) vs sham (n = 10, grey) surgery (day 20); two independent experiments were conducted. All populations were gated based on live/dead stain and CD45+. c, Flow cytometric analysis of CD11b+Gr1-Cd11c+ dendritic cell-like exposed to MI (n = 7) or sham (n = 7) surgery and CD11b+ Gr1+ F4/80+ macrophage-like cells in E0771 tumors from mice exposed to MI (n = 5) or sham (n = 4) surgery. d, Flow cytometric analysis of tumor immune cells (% total live cells) at day 20 to identify CD3+ T cells (n = 11 MI, 10 sham) and CD11b + myeloid (n = 14 MI, 11 sham) subsets: CD11b+ Ly6G+, neutrophils; CD11b+ Gr1–, macrophage-like cells; CD11b+ Ly6Chi, monocytes; Data are the mean ± s.e.m and P values in data were analyzed by a two-tailed Mann–Whitney test (b [FoxP3+ cells]), or a two-tailed unpaired Student’s t-test (b,c,d).
Extended Data Fig. 4 Circulating leukocytes and bone marrow progenitors in tumor bearing mice following MI or sham surgery.
a, Numbers of circulating leukocytes and relative proportion of neutrophils, eosinophils, lymphocytes and basophils in E0771 tumor bearing mice after MI (n = 8) or sham (n = 8) surgery. b,c, Flow cytometric analysis of bone marrow hematopoietic and stem cell populations in E0771 tumor bearing mice 9 days post-MI or sham surgery (12 days post-tumor cell implantation). LSK: Lineage–Sca1–cKit+ cells; CMP: common myeloid progenitor; GMP: granulocytic myeloid progenitor; MEP: megakaryocyte–erythroid progenitor. (a-c) two independent experiments were conducted. Data are the mean ± s.e.m. P values were calculated using a two-way analysis of variance (ANOVA), with results not significant (p > 0.05) (a), or a two-tailed unpaired Student’s t-test (c).
Extended Data Fig. 5 Monocyte adoptive transfer experiments into tumor bearing mice; RNA-seq analyses of tumor and bone marrow monocytes in mice exposed to MI or sham surgery.
a–c, Monocytes were adoptively transferred from CD45.1 mice 9 days after exposure to MI or sham surgery into E0771 tumor bearing CD45.2 mice (a), and tumor weight (b), and CD45.1 Ly6Chi monocytes recruited to the tumor (c) were determined 16 h later (n = 9 sham; n = 8 MI). d–e, Monocytes were adoptively transferred from CD45.1 mice into E0771 tumor-bearing CD45.2 mice 9 days following MI or sham surgery (d), and tumor weight was determined 16 h later (n = 9 sham; n = 10 MI) (e). f, Fold change (FC) in gene expression in E0771 tumors from mice 8 days post-MI compared to sham surgery (n = 3 per group). g, Heatmap of genes significantly up- or down-regulated (p < 0.05, two-sided Wald test) in bone marrow monocytes 9-10 days after MI vs sham treatment specifically in tumor bearing mice. MI (n = 6, 3 pools of 2 mice), sham (n = 10, 5 pools of 2 mice), MI + tumor (n = 5), or sham+tumor (n = 6). (a-e) two independent experiments were conducted. Data are the mean ± S.D. and P values were analyzed by two-tailed unpaired Student’s t-test (b, c, e, f), with comparisons not significant unless indicated. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001.
Extended Data Fig. 6 CCR2 inhibition mitigates MI-induced accelerated tumor growth.
a, Outline of CCR2 inhibition study in mice implanted with E0771 tumor cells and exposed to MI or sham surgery. CCR2 inhibitor (CCR2i) was administered starting at 7 days after MI or sham surgery. b, Tumor volume over the course of the study (left) and at sacrifice (day 20 day) (right) in mice exposed to MI (n = 5 CCR2 inhibitor, n = 7 DMSO control) or sham surgery (n = 8 CCR2 inhibitor, n = 7 DMSO control). Two independent experiments were conducted (a-c). P values were calculated using a repeated measures analysis of variance (ANOVA) with Bonferroni’s multiple comparisons test (b) or one way ANOVA (c).
Extended Data Fig. 7 Monocytic myeloid-derived suppressor cells isolated from tumors of mice exposed to MI or sham surgery do not alter the proliferation of T cells in vitro.
a, Representative flow plots of CD11b-Ly6C-CD8+ T cells from MDSC:T cell suppression assay, representing % IFN𝛾+, TNFα+, Granzyme B (GrB)+ populations. b,c, Purified splenic CD8+ T cells from naïve mice were stimulated with αCD3/αCD28 for 72 hours in the presence of mMDSCs isolated from tumors from mice exposed to MI (n = 5) or sham surgery (n = 7), and CD8+ T cell proliferation was assessed by % Ki67 (b) and replication and division index (c) using a cell trace proliferation dye. (a,b,c) two independent experiments were conducted. Data are the mean ± s.e.m. P values were calculated using or two-tailed unpaired Student’s t-test (b,c), where relevant comparisons were not significant (p > 0.05).
Extended Data Fig. 8 RNA-seq and ATAC-seq analyses of mMDSCs in tumor and bone marrow of mice exposed to MI or sham surgery.
a, Heat map showing differential gene expression (log2FC) of select immune-related genes from RNA-Seq of tumoral CD11b+ Ly6Chigh mMDSCs from mice 17 days post-MI (n = 5) or sham (n = 6) surgery (Padj < 0.1). b, Gene set concordance analysis showing that the top 1000 differentially expressed genes up- and down-regulated in tumor CD11b + Ly6Chigh mMDSCs (x axis) are enriched in genes up- and down-regulated in bone marrow Ly6Chigh monocytes (y axis) from the same mice 17 days post-MI, compared to background gene expression. c, Gene ontology (GO) analyses of more accessible chromatin regions (n = 942) in bone marrow Ly6Chigh monocytes from mice exposed to MI vs. sham surgery. d, List of genes differentially expressed in tumor Ly6Chigh monocytes whose chromatin loci that are also less accessible in bone marrow Ly6Chigh monocytes after MI compared to sham surgery, grouped by transcription factors identified in Fig. 3e. e, ATAC-Seq (top) and RNA-Seq (bottom) reads in bone marrow and tumor Ly6Chigh monocytes at selected gene loci. P(adj) values were calculated using the Benjamini-Hochberg method (a). P values determined by two-sided Student’s T-test (b) or hypergeometric distribution (c).
Extended Data Fig. 9 The effect of systemic CD8+ cell depletion on tumor growth in mice following MI or sham surgery.
a, E0771 tumor-bearing mice were exposed to MI or sham surgery and randomly allocated to either intraperitoneal IgG or anti-CD8 injections 10, 15, and 19 days following tumor implantation. b, Intratumoral T cell content measured by flow cytometry in sham+IgG (n = 8), sham+anti-CD8 (n = 7), MI + IgG (n = 8) and MI + anti-CD8 (n = 9). c, Tumor volume was followed over 20 days and at sacrifice (d) in sham+IgG (n = 8), sham+anti-CD8 (n = 7), MI + IgG (n = 8) and MI + anti-CD8 (n = 7). a–d, two independent experiments were conducted. Data are the mean ± s.e.m. P values were calculated using a repeated measures analysis of variance (ANOVA) with Bonferroni’s multiple comparisons test (c) or a two-tailed (b) or one-tailed (d) Mann–Whitney test.
Extended Data Fig. 10 Tumor growth, circulating monocytes and intratumoral innate immune flow cytometry gating strategy in MMTV-PyMT mice following surgical MI or sham surgery.
a, Mean tumor volume at time of MI (n = 8) or sham surgery (n = 8) in MMTV-PyMT mice. b, Tumor growth in MMTV-PyMT mice after MI or sham surgery (n = 8/group). c, Tumor volume at sacrifice in MMTV-PyMT mice used for the metastasis subgroup analysis (n = 4/group). d, Circulating monocytes in MMTV mice exposed to MI or sham surgery (n = 8/group). e, Flow cytometry gating strategy for myeloid cells from MMTV-PyMT tumors. Mammary tissue macrophages (MTM: CD11bhighMHCII+); granulocytic myeloid derived suppressor cell (gMDSC: CD11bhighMHCII-Ly6CloLy6Ghigh); monocytic myeloid derived suppressor cell (mMDSC: CD11bhighMHCII-Ly6ChiLy6Glo); tumor-associated macrophage (TAM: CD11bloMHCIIhigh). (a-e) three independent experiments were conducted. Data are the mean ± s.e.m and P values were calculated using a repeated measures analysis of variance (ANOVA) (d) with Bonferroni’s multiple comparisons test (b) or by a two-tailed unpaired Student’s t-test (a, c).
Supplementary information
Supplementary Information
Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Koelwyn, G.J., Newman, A.A.C., Afonso, M.S. et al. Myocardial infarction accelerates breast cancer via innate immune reprogramming. Nat Med 26, 1452–1458 (2020). https://doi.org/10.1038/s41591-020-0964-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-020-0964-7
This article is cited by
-
Current evidence regarding the cellular mechanisms associated with cancer progression due to cardiovascular diseases
Journal of Translational Medicine (2024)
-
Effect of inflammation on association between cancer and coronary artery disease
BMC Cardiovascular Disorders (2024)
-
CVD phenotyping in oncologic disorders: cardio-miRNAs as a potential target to improve individual outcomes in revers cardio-oncology
Journal of Translational Medicine (2024)
-
Made to order: emergency myelopoiesis and demand-adapted innate immune cell production
Nature Reviews Immunology (2024)
-
Linking immune modulation to cardiac fibrosis
Nature Cardiovascular Research (2024)