Abstract
The rapid increase in telemedicine coupled with recent advances in diagnostic artificial intelligence (AI) create the imperative to consider the opportunities and risks of inserting AI-based support into new paradigms of care. Here we build on recent achievements in the accuracy of image-based AI for skin cancer diagnosis to address the effects of varied representations of AI-based support across different levels of clinical expertise and multiple clinical workflows. We find that good quality AI-based support of clinical decision-making improves diagnostic accuracy over that of either AI or physicians alone, and that the least experienced clinicians gain the most from AI-based support. We further find that AI-based multiclass probabilities outperformed content-based image retrieval (CBIR) representations of AI in the mobile technology environment, and AI-based support had utility in simulations of second opinions and of telemedicine triage. In addition to demonstrating the potential benefits associated with good quality AI in the hands of non-expert clinicians, we find that faulty AI can mislead the entire spectrum of clinicians, including experts. Lastly, we show that insights derived from AI class-activation maps can inform improvements in human diagnosis. Together, our approach and findings offer a framework for future studies across the spectrum of image-based diagnostics to improve human–computer collaboration in clinical practice.
Similar content being viewed by others
Main
Image-based AI has the potential to improve visual diagnostic accuracy. Limited physical access to health-care providers during the recent COVID-19 pandemic is prompting changes in health-care delivery and accelerating the adoption of telemedicine1. AI-based triage and decision support could assist readers in managing workloads and expanding their performance. Most research to date has been predicated on head-to-head comparisons of the diagnostic accuracy of AI-based systems with that of humans2,3,4. Similarly, recent studies in dermatology demonstrate that AI for selected lesions is equivalent or even superior to human experts in image-based diagnosis under experimental conditions5,6,7,8,9. This competitive view of AI is evolving based on studies suggesting that a more promising approach is human–AI cooperation10,11,12,13,14,15. The role of human–computer collaboration in health-care delivery, the appropriate settings in which it can be applied and its impact on the quality of care have yet to be evaluated16. To this end, we studied the use case of skin cancer diagnosis to address the effects of varied representations of AI-based support across different levels of clinical expertise and multiple clinical workflows.
To explore the impact of different representations of current state-of-the-art AI on diagnostic accuracy of clinicians in different scenarios, we first trained a 34-layer residual network (ResNet34), a particular type of convolutional neural network (CNN), on the training dataset of a publicly available image benchmark of pigmented lesions containing seven diagnostic categories, including malignant (melanomas (MELs), basal cell carcinomas (BCCs) and actinic keratoses and intraepithelial carcinomas (AKIECs)) and benign (melanocytic nevi (NVs), benign keratinocytic lesions (BKLs), dermatofibromas (DFs) and vascular lesions (VASCs)) proliferations17. When tested on the corresponding publicly available benchmark test set, the mean recall of our CNN across all disease categories was 77.7% (95% confidence interval (CI) 70.3% to 85.1%), and the accuracy was 80.3%. When compared with the results of a recently published reader study, this CNN outperforms most human raters and ranks in the top quartile of machine-learning algorithms that were developed and tested with the same image dataset18. To examine whether human–computer collaboration is influenced by the way that the output from the CNN is presented to humans, we developed a web-based user interface for comparing three forms of output from the CNN as decision support to human raters (Fig. 1).
a, Schematic overview of the interaction modalities offered: (I) AI-based multiclass probabilities, (II) AI-based probability of malignancy, (III) AI-based CBIR and (IV) crowd-based multiclass probabilities. b, Raters needed significantly more time to engage with CBIR support (n = 302 ratings; mean 16.5 s, 95% CI 14.5 to 18.6 s) than with multiclass probabilities (n = 302 ratings; mean 4.6 s, 95% CI 4.3 to 4.9 s; P = 2.6 × 10−24), malignancy probability (n = 301 ratings; mean 5.2 s, 95% CI 4.6 to 5.7 s; P = 1.0 × 10−22), crowd-based multiclass probabilities (n = 301 ratings, mean 4.5 s, 95% CI 4.1 to 4.9 s; P = 2.0 × 10−25) or without support (n = 302 ratings; mean 5.6 s, 95% CI 5.4 to 5.8 s; P = 4.5 × 10−22). All P values were derived from two-sided paired t-tests with Holm–Bonferroni correction for multiple comparisons. In the CBIR group, one outlier of >200 s is not shown on the plot. The bars denote means, and error bars represent 95% CIs. c, The number of interactions with CBIR-based support, as measured by enlarged thumbnails, is low and decreases further with the number of interactions, indicating that this type of support is not appreciated over time.
The representations of AI that we selected derive from the literature and differ in key characteristics, including simplicity, granularity and concreteness. Because our task was a multiclass classification problem, one obvious approach was to provide AI-based multiclass probabilities. The second approach was motivated by solutions already implemented in currently available AI-based support for skin cancer diagnosis6; we dichotomized the disease categories into a benign and a malignant class and displayed the AI-predicted probability of malignancy. For the third and fundamentally different approach, we used the same CNN to implement a form of AI-based CBIR that supports physicians in the interpretation of images by searching databases to retrieve similar images with known diagnoses11,19,20. As an alternative to AI-based decision support, we also provided previously collected9 rating frequencies of 511 human raters for each disease category (crowd-based multiclass probabilities).
Next, we invited human raters to participate in a reader study. A total of 302 raters from 41 countries participated, including 169 (56.0%) board-certified dermatologists, 77 (25.5%) dermatology residents and 38 (12.6%) general practitioners. The raters’ task was to diagnose batches of images, first without and then with one type of decision support. We recorded the time needed to reach a diagnosis, normalized this time over all individual ratings for each user and interaction modality, and used this as a surrogate marker for confidence.
We collected 512 tests and 13,428 ratings. Our results show that decision support with AI-based multiclass probabilities improves the accuracy of human raters from 63.6% to 77.0% (increase of 13.3%, 95% CI 11.5% to 15.2%; P = 4.9 × 10−35, two-sided paired t-test, t = 14.5, d.f. = 301; n = 302 raters), but no improvement was observed for decision support with AI-based prediction of malignancy or with our representation of AI-based CBIR (Fig. 2a–d and Supplementary Tables 1 and 2).
a–d, AI-based multiclass probabilities (a), AI-based probability of malignancy (b), AI-based CBIR (c) and crowd-based multiclass probabilities (d). In a multiclass classification problem, humans show a net gain from support by AI-based and crowd-based multiclass probabilities but not from other less granular or less explicit types of decision support. e,f, Net gain with respect to the frequency of correct diagnoses decreases with experience and confidence. Experts who are confident in a given diagnosis do not benefit from AI-based support. Bars denote means, whiskers represent 95% CIs and dots denote individual raters. g, AI-estimated rank of a diagnosis for final rater decisions, grouped by whether the rater changed their initial diagnosis. While changes occurred almost exclusively for the top class (class 1; left), a substantial number of decisions remained unchanged in cases where the AI evaluated them as second or third ranked (right). h, When in disagreement with the top AI predictions (class 1) before interaction, raters changed their opinion to these predictions if the AI multiclass probabilities were large. Bars denote means, and error bars represent 95% CIs. i, Raters were susceptible to faulty AI-based support. The significant gain in accuracy (left, n = 155 raters; median 9.5%; P = 1.2 × 10−12, two-sided paired Wilcoxon signed-rank test) turned into a significant loss (right, n = 155 raters; median −6.3%; P = 6.0 × 10−13, two-sided paired Wilcoxon signed-rank test) when AI-based multiclass probabilities of the top predictions (class 1) were changed to a random incorrect answer. Thick central lines denote the medians, lower and upper box limits denote the first and third quartiles and whiskers extend from the box to the outermost extreme value but no further than 1.5 times the interquartile range (IQR).
This suggests that the form of decision support should be in accordance with the given task. The probability of malignancy may be useful for simple binary management decisions, such as whether to perform a biopsy or not, but not for a multiclass diagnostic problem. The studied form of AI-based CBIR is neither simple nor concrete; it needs more extensive cognitive engagement in terms of time and decision-making, because the rater needs to extrapolate the diagnosis from similarities between the test image and images with known diagnoses. The raters needed significantly more time to interact with AI-based CBIR decision support than with other types of support (Fig. 1b). Over time, human raters also tended to ignore the AI-based CBIR decision support (Fig. 1c). However, given that a large spectrum of CBIR approaches are described in the literature, another form of CBIR may still provide benefit. It has been shown that human-centered refinement tools improve the end user experience of CBIR in pathology and increase trust and utility21. Future work should, therefore, study a broader variety of layouts and combinations of collaborations between AI and humans.
After we established that multiclass probabilities were the best form of CNN output for the given task, we focused on this form to explore the impact of AI-based support on human performance in more detail. We show an inverse relationship between the net gain from AI-based support and rater experience (Pearson’s r = −0.18, 95% CI −0.28 to −0.07, P = 1.5 × 10−2; n = 302 raters). Raters in the least experienced group changed their initial diagnosis more often than experts (mean 26.0%, 95% CI 21.3% to 30.7% versus mean 14.7%, 95% CI 9.9% to 19.6%). Expert raters benefited only marginally (net gain 13.4%, 95% CI 6.3% to 20.6%) and only if they were not confident with their initial diagnosis, but not if they were confident (−0.7%, 95% CI −6.8% to 5.4%; Fig. 2e,f). If experts were confident, they were usually correct and did not need support. This finding suggests that, if experts have high confidence in their initial diagnosis, they should ignore AI-based support or not use it at all. This simple heuristic corresponds to what we observed in our experiments; if their initial diagnosis was not in agreement with the top class predicted by the CNN, the experts changed their initial diagnosis less often if they were confident (29.8%, 95% CI 14.1% to 45.4%) and more often if they were not confident (53.9%, 95% CI 33.2 % to 74.7%). The least experienced raters tended to accept AI-based support that contradicted their initial diagnosis even if they were confident. In general, raters changed their initial diagnosis less often if they were confident than if they were not confident in their decision (14.7%, 95% CI 12.6% to 16.8% versus 37.5%, 95% CI 34.0% to 41.0%; P = 1.9 × 10−25, two-sided paired t-test; n = 302 raters).
Having established a positive impact of good quality AI-based support on diagnostic accuracy, we tested the impact of ‘faulty’ AI on diagnostic accuracy. Faulty AI could result from the application of AI algorithms on examples beyond the domain of images on which the AI was trained7,9,22 or the more remote possibility of adversarial attacks23,24,25. To represent faulty AI, we intentionally generated misleading AI-based multiclass probabilities. If the top class probability of the CNN favored the correct diagnosis, we switched the probabilities in such a way that the CNN output favored a random incorrect diagnosis. We demonstrate that any previously observed gains in accuracy with AI-based support turn into a loss when that AI support is faulty. Figure 2i shows that all groups of raters are susceptible to underperforming in this scenario. Our results suggest that, if raters build up the trust that is necessary to benefit from AI-based support, they are also vulnerable to perform below their expected ability if there is a fault with the AI. Whether techniques to facilitate interpretability or explainability mitigate the risk of this negative impact remains an open topic of research21,26.
Another finding of importance is that the benefit of human–computer collaboration is asymmetrically distributed across disease categories. Our data showed that the net gain was higher for the class of pigmented actinic keratoses and intraepithelial carcinoma (increase of 31.5%, 95% CI 22.9% to 40.1%; n = 43 images) than for other categories (Supplementary Table 3). This suggests that the benefit of AI-based support needs to be adapted to the given task and the expected prevalence of target conditions.
We further demonstrate that AI-based multiclass ranking and probabilities have an impact on the raters’ tendency to change their initial diagnosis. Most changes occurred in favor of the AI-predicted top category. Raters typically maintained their decisions that were in disagreement with the AI prediction only if that decision was ranked by AI prediction as at least the second or third option (Fig. 2g). Furthermore, raters tended to change their assessments more frequently when the difference in the AI-predicted probability between the initially selected category and the AI top category was high (Fig. 2h). This suggests that the distribution of class probabilities affects the behavior of raters. Big winners and top-ranked classes are preferred to small winners, and categories with low probabilities will barely affect the decision of raters.
Additionally, we demonstrate that aggregated AI-based multiclass probabilities and crowd wisdom significantly increased the number of correct diagnoses in comparison to individual raters or AI in isolation (Fig. 3a). The disadvantage of crowd wisdom is that it is not readily and instantly available; in contrast to software, raters cannot be cloned.
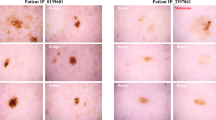
a, Single human raters (top) achieve the lowest mean accuracy (64.8%, 95% CI 62.4% to 67.3%; n = 600 images). Ratings of bootstrapped human collectives (middle) show significantly higher accuracy (73.7%, 95% CI 70.9% to 76.6%; P = 1.5 × 10−35; n = 600 images), similar to the raw top class predictions of the CNN (blue line; 76.9%). The highest accuracy is achieved by combining AI-based multiclass probabilities and human collectives (bottom), which is significantly higher than for collectives alone (81.0%, 95% CI 78.2% to 83.9%; P = 8.6 × 10−9; n = 600 images). Bars denote means, whiskers represent 95% CIs and dots represent the mean correct rating of the corresponding group of a single image; groups were compared using a two-sided paired Wilcoxon signed-rank test. b, Performance of CNN predictions used as a filter in a screening setting of high-risk patients who provided self-made dermoscopic photographs of their skin lesions over 3 months. Top bars denote whether the CNN predicted malignancy on an image, lesion or patient level, and bottom bars denote the corresponding ground truth. While the CNN shows low sensitivity for single images, it detects the majority of skin cancer cases from multiple images (lesion level) and almost every patient with skin cancer (patient level). c, Changes of raters’ decisions with AI-based support in a telemedical setting with dermoscopic images of pigmented lesions taken by patients. P values were derived from two-sided paired t-tests with Holm–Bonferroni correction for multiple comparisons. Colored dots and whiskers denote means and 95% CIs, and gray dots represent correct answers of raters. d, Switch of management decisions using CNN predictions as a second opinion. Raters’ decisions before (top bar) and after (bottom bar) seeing CNN predictions are shown, grouped by ground truth. e, Change of correct answers after explainable AI-guided teaching about chronic sun damage in the background of pigmented actinic keratoses. The overall percentage of correct answers increased with teaching (left), mostly as a result of improved recognition of actinic keratoses (right). P values were derived from two-sided paired t-tests with Holm–Bonferroni correction for multiple comparisons. Colored dots and whiskers denote means and 95% CIs, and gray dots represent correct answers of raters.
Next, we analyzed the impact of AI-based support in clinically relevant scenarios. To examine the potential of AI-based support in telemedicine, we reused prospectively collected images of a randomized controlled trial on self-examinations in high-risk patients27. Ninety-three participants submitted 1,521 self-made photographs of 596 suspicious lesions for telediagnosis. While the CNN was trained only on curated images of pigmented lesions, this sample also included non-pigmented variants of keratinocyte cancer, mucosal lesions and low-quality images. Although the proportion of correct specific diagnoses was significantly lower for these images (53.9% versus 76.2%; P = 8.9 × 10−14, chi-squared test; n = 1,430 images), the CNN was able to recognize 95.2% of patients with skin cancer at a specificity of 59.2% (Fig. 3b). Similarly to recent findings in AI-based breast cancer screening3, our results indicate that AI-based skin cancer screening could triage high-risk cases and extend the intervals between face-to-face visits in low-risk cases. The optimal operating points to balance the potential benefits of AI-based triage with the risk of filtering out patients with skin cancer remain to be determined.
A possible explanation for the reasonably accurate performance of the CNN as a tool for triage in telemedicine, despite the inclusion of non-pigmented skin lesions, is that pigmented and non-pigmented variants of keratinocyte cancer share common criteria. However, this cannot be guaranteed in other settings; the results of the International Skin Imaging Collaboration (ISIC) 2019 challenge, for example, demonstrated that AI does not work reliably on out-of-distribution images28. Furthermore, we show that, within the domain of pigmented skin lesions, AI-based support helps less experienced raters to improve to the expert level in telemedicine (Fig. 3c). Limitations of the telemedicine setting are that the sample did not include melanomas and the number of malignant cases was relatively small.
In another scenario, we asked dermatologists to rethink their face-to-face decisions in suspicious cases after providing them with AI-based multiclass probabilities, but without making them aware that they had previously managed the patient. As shown in Fig. 3d, with AI-based support, dermatologists switched from ‘excision’ to ‘monitor’ in 15.5% (7 of 45) of decisions for benign lesions, without increasing the number of malignant lesions that switched contrariwise. This result illustrates how human–computer collaboration could decrease the number of unwarranted interventions and costs. AI-based support in this setting increased the frequency of correct specific diagnoses from 55.6% to 75.0% (P = 0.029, two-sided paired Wilcoxon signed-rank test; n = 11 raters).
Finally, we demonstrate that explanations for AI-based predictions can be translated into a human-understandable visual concept. In a previous study, we showed that misclassification of pigmented actinic keratoses by humans is one reason for the superiority of AI over human experts9. By analyzing gradient-weighted class activation mapping (Grad-CAM29), we show that attention of the CNN outside the object is higher for the prediction of actinic keratoses than for other categories (Extended Data Fig. 1). Background attention30,31 is not necessarily a Clever Hans predictor32,33 but can be part of a valid general concept. Chronic ultraviolet light damage causes actinic keratoses and is always present in the surrounding skin of actinic keratoses but not necessarily in other disease categories. We hypothesize that, due to visual entrenchment, humans focus on the lesion and not on the background and frequently miss this clue. Here we show that teaching medical students to pay attention to chronic sun damage in the background improved the frequency of correct diagnoses of pigmented actinic keratoses from 32.5% (95% CI 30.0% to 35.0%) to 47.3% (95% CI 43.9% to 50.8%; P = 3.6 × 10−13, two-sided paired t-test; n = 189 raters). The overall frequency of correct diagnoses in all categories combined increased from 55.2% to 59.1% (mean difference of 3.7%, 95% CI 2.4% to 5.3%; P = 3.4 × 10−6, two-sided paired t-test, t = 5.2, d.f. = 188; n = 189 raters; Fig. 3e).
This study examines human–computer collaboration from multiple angles and under varying conditions. We used the domain of skin cancer recognition for simplicity, but our study could serve as a framework for similar research in image-based diagnostic medicine. In contrast to the current narrative, our findings suggest that the primary focus should shift from human–computer competition to human–computer collaboration. From a regulatory perspective, the performance of AI-based systems should be tested under real-world conditions in the hands of the intended users and not as stand-alone devices. Only then can we expect to rationally adopt and improve AI-based decision support and to accelerate its evolution.
Methods
Network training
We fine-tuned a CNN for classification of seven different categories of the HAM10000 dataset17. We performed training on NVIDIA graphics processing units (GPUs) using the Pytorch34 framework and chose a ResNet34 (ref. 35) architecture, with weights initiated by pretraining on ImageNet36 data. Cross-entropy served as the loss function, with weighting dependent on the frequency of classes within the dataset. The learning rate was initialized at 0.0001, with a tenfold reduction in case of no validation loss improvement for more than three epochs, but a minimum of 1 × 10−9. We used adaptive moment estimation (Adam37) as the optimizer and performed a maximum of 100 training epochs with early stopping. Images were presented in batches of 32, randomly cropped and resized to 224 × 224 pixels without normalization of a mean pixel, randomly rotated by 90 degrees and flipped with minor jitter in color, contrast, saturation and hue.
The publicly available HAM10000 dataset, which corresponds to the training set of the ISIC 2018 challenge18, was the source of images used for training and fivefold cross-validation. We selected the single best performing network on the hold-out validation set for further interaction with raters. For inference, images were cropped to 80% and resized to 224 × 224 pixels, with minor test-time augmentation consisting of horizontal flipping and rotation by 0 or 90 degrees. For the telemedicine dataset, we also applied color normalization via Shades of Gray38 with a Minkowski norm of 6. The multiclass probabilities presented to the raters were obtained by applying a softmax function to contain all class probabilities between 0–100%. To find similar images, we used the same CNN to extract the feature vector of the target image and compared it to feature vectors of images in the HAM10000 dataset via cosine similarity20. We stored the four closest images of each class and presented them in the AI-based CBIR decision support.
Interaction platform and raters
Online interaction platform
The web-based platform DermaChallenge, which was developed at the Medical University of Vienna, served as the interface through which the performance of human raters and AI for the diagnostic task was evaluated and quantified. The platform is split into a back end and a front end, and both are deployed on a stack of well-known web technologies (Linux, Apache, MySQL and PHP). Please refer to the Nature Research Reporting Summary for details of the specific software versions used. The back end offers a representational state transfer interface to load and persist data, as well as JavaScript Object Notation web tokens to authenticate participants. The transport layer security and secure sockets layer protocol are used to encrypt all communications. The front end is optimized for mobile devices (mobile phones and tablets) but can also be used on any other platform via a JavaScript-enabled web browser. Before public deployment, five users tested the platform.
Recruitment and characteristics of raters
We used mailing lists and social media posts of the International Society of Dermoscopy to recruit online raters. To participate in the study, raters had to register with a username, valid email address and password. In addition, we asked raters for details on their age (age groups spanning 10 years), gender, country, profession and years of experience in dermatoscopy ((1) less than 1 year, (2) opportunistic use for more than 1 year, (3) regular use for 1 to 5 years, (4) regular use for more than 5 years or (5) more than 10 years of experience). Each rater had to perform multiple screening tests to ensure that the self-reported experience matched actual skills. Screening tests consisted of simple domain-specific tasks, for example, to assign one of the seven possible diagnoses to ten cases, to separate melanomas from non-melanomas and to separate seborrheic keratoses from other lesions. We recruited 302 raters for the first interaction study that screened different forms of AI-based support, and 155 raters were recruited for the extended interaction study (inclusion of images with faulty AI-based support) and the telemedicine study (Supplementary Table 4). The distribution of raters according to task is presented in Supplementary Table 4. Second-opinion raters consisted of eight board-certified dermatologists and three dermatology residents, who were recruited because they diagnosed and managed more than two suspicious skin lesions on a face-to-face basis between April and September 2019. For the knowledge transfer study, we invited fourth-year medical students to participate; of the 650 medical students invited, 200 agreed to participate and 189 answered more than 50% of the test questions.
Characteristics of images and patients
The benchmark test set of the ISIC 2018 challenge served as the sample for the interaction studies9. Of the 1,511 dermoscopic images in this set, 928 images were collected in the Department of Dermatology at the Medical University of Vienna, 267 images were collected in the skin cancer practice of Cliff Rosendahl in Queensland and the remaining 316 images were collected in other centers in Turkey (n = 117), New Zealand (n = 87), Sweden (n = 92) and Argentina (n = 20), to ensure diversity of skin types. The mean age of patients was 50.8 years (s.d. 17.4 years), and 46.2% of patients were female. The Austrian image set consists of lesions from patients referred to a tertiary European center specializing in the early detection of melanoma in high-risk groups. This group of patients is mainly of European ancestry and have a large number of nevi and skin types I–III. The Australian image set includes lesions from patients of a primary-care facility in an area with a high incidence of skin cancer. Patients are typified by Celtic complexion, skin type I or II and chronic sun damage. Routine pathology evaluation (n = 786), biology (that is, >1.5 years of sequential dermoscopic imaging without changes; n = 458), expert consensus in common, straightforward, non-melanocytic cases that were not excised (n = 260) and in vivo confocal images (n = 7) served as the ground truth. Controversial cases with ambiguous histopathologic reports were excluded. Due to random sampling, only 1,412 of 1,511 images were finally used and evaluated by the raters. The 1,412 used cases consisted of 43 AKIECs, 93 BCCs, 217 BKLs, 44 DFs, 171 MELs, 809 NVs and 35 VASCs.
For the telemedicine study, we included 93 of 98 participants (mean age 41.1 years (s.d. 12.2 years); 71% female) from the intervention arm of a recently conducted prospective randomized study27 on mobile teledermoscopy for skin self-examinations. All 93 patients permitted reuse of their images. The participants had at least two skin cancer risk factors (light skin complexion and fair hair; skin that never or rarely tans and always or mostly burns; a family history of melanoma or a personal history of skin cancer, or many nevi; and residing in Queensland) as self-reported in the eligibility survey. A teledermoscopic evaluation was performed for all lesions. Face-to-face examination by an experienced board-certified dermatologist (H.P.S.) or the histopathologic report, in cases where the lesion was removed, served as the ground truth. The set of lesions consisted of 1,521 images of 596 lesions, including 29 AKIECs, 6 BCCs, 102 BKLs, 410 NVs, 2 squamous cell carcinomas (SCCs) and 9 VASCs. For calculation of diagnostic values, ground-truth data were mapped to classes of the HAM10000 dataset, if possible. We excluded nonspecific categories (n = 38 lesions) such as ‘other’, ‘no lesion’ or ‘previously removed’, because they could be mapped to neither the ‘benign’ nor ‘malignant’ category. The sample also included images that were not represented in the training data (non-pigmented variants of keratinocyte cancers, mucosal lesions and low-quality images), which were excluded from the telemedicine support study but not from the triage study, to better simulate a realistic scenario.
For the second-opinion study, we searched the database of the Department of Dermatology at the Medical University of Vienna for dermoscopy images taken between April and September 2019. We included images if the lesion was excised and had a definite histopathologic diagnosis and if lesions were examined by a physician who was responsible for the face-to-face diagnosis of at least two other cases in this time period. The final sample set (n = 79) included 3 AKIECs, 23 BCCs, 13 BKLs, 2 DFs, 15 MELs, 21 NVs, 1 ‘other’ (scar) and 1 SCC. The mean age of patients was 64.6 years (s.d. 19.8 years), and 34.5% of patients were female. Patients were mainly of European ancestry and had skin type II (41.7%), III (57.1%) or IV (1.2%). As in the telemedicine scenario, we did not exclude images of categories that were not present in the training data or images of low quality.
For the knowledge transfer study, the sample cases (n = 25) were randomly selected from the ISIC 2018 test set and stratified by diagnosis (6 AKIECs, 3 BCCs, 3 BKLs, 3 DFs, 3 MELs, 4 NVs and 3 VASCs).
Design of diagnostic studies
To test the interaction of raters with different forms of AI-based decision support, we generated batches of 28 images. Each batch contained four randomly selected examples of every class. The raters’ task was to diagnose the 28 unknown test images, first without and then with one type of decision support. We created a stratified randomization procedure to ensure a balanced distribution of the four types of decision support over all disease categories. The interaction study was online from 29 May 2019 to 15 January 2020. We excluded tests if the number of correct answers was lower than expected by chance to avoid noisy random data. We included only the first five tests for each rater to avoid biasing the results toward raters with high repetitions.
The extended interaction study was open for participation between 15 January 2020 and 18 February 2020, presenting only multiclass probabilities as decision support. It included one image for every diagnosis from the ISIC 2018 test set with unaltered AI-based multiclass probabilities, two images with shuffled (resulting as incorrect) AI-based multiclass probabilities and eight images from the telemedicine study (see ‘Characteristics of images and patients’). The image sources were not disclosed to the raters.
The second-opinion study was performed on a local web interface. Physicians who examined the patient face to face in real life were asked to reconsider their diagnosis and decisions with AI-based support. The case presentations included metadata (age, gender and localization), overview and close-up images (if available) and dermoscopic images. Physicians were not made aware that they had treated the patient before or of their previous decision on the case. Physicians were asked to provide their best diagnosis out of the seven predefined disease categories, as well as an extra category termed ‘other’, followed by their management decision (‘no intervention’, ‘monitor’ or ‘excise’). No time constraints were set for this task.
For the knowledge transfer study, we first examined the gradient-weighted class-activation maps29, which were created for all images of the training set. We observed that the background attention of the CNN was significantly higher for predictions of the ‘pigmented actinic keratosis’ class than for other classes (P = 4.6 × 10−12, two-sided unpaired t-test; Extended Data Fig. 2). We interpreted this finding as a diagnostic clue that points to the severely sun-damaged skin in the background of actinic keratoses, which is usually absent or not as severe in other disease categories. To test the hypothesis that teaching this clue to humans will improve their diagnostic skills, fourth-year medical students without previous knowledge of skin cancer detection received a 30-min introductory lecture about dermoscopy, and immediately thereafter students had to diagnose 25 test images (single best diagnosis). Answers were collected with a wireless audience response and voting system. Next, the lecturer presented an additional clue of ‘sun-damaged skin in the surrounding skin of actinic keratoses’ and the students repeated the test.
Statistics
To simulate collective ratings of realistically small human groups (Fig. 3a), we confined the dataset to images with at least three distinct ratings (resulting range of ratings per image: 3–69). For each image, we created 30 bootstraps of three to five randomly selected ratings, whichever was the maximum available without replacement, and determined the most common rating as the prediction of the collective (first past the post). Ties were broken randomly. Next, we calculated the proportion of correct bootstrapped predictions to obtain the mean accuracy for each image as published previously39. To combine human collectives with CNN-based predictions, we took the arithmetic mean of the sum of the human multiclass probabilities, which were derived from the frequencies of bootstrapped human ratings, and the corresponding CNN-based multiclass probabilities. For analyses of diagnoses, we averaged the results for each image before comparisons; for analyses of raters with and without decision support, we calculated the arithmetic mean for each user before comparisons. The mean answering time for each user in every interaction modality served as a surrogate marker for confidence; answers that were faster or slower than the individual mean were regarded as ‘confident’ or ‘non-confident’, respectively.
For the filtering procedure in the telemedicine study, we used a predefined cutoff of ≥0.17 to indicate malignancy, because this cutoff was selected by human raters in the interaction study (Extended Data Fig. 2). If patients photographed a lesion more than once, a single image above the cutoff was sufficient to label the lesion as ‘probably malignant’ and likewise on the patient level. We used a one-sample t-test to distinguish whether continuous data with normal distributions deviated from zero. Comparisons of continuous data between groups were performed with paired or unpaired t-tests or Wilcoxon signed-rank test, as appropriate. A chi-squared test was used to compare proportions. All reported P values were corrected for multiple testing (Holm–Bonferroni40), and a two-sided P value < 0.05 was regarded as statistically significant. All analyses were performed using R v3.6.2 (ref. 41), and plots were created with ggplot v3.2.1 (ref. 42) and ggalluvial v0.11.1.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The origin of training set images is reported in the dataset publication of HAM10000 (https://doi.org/10.1038/sdata.2018.161)17. Training set and test set images with reader data are available from the Harvard Dataverse (https://doi.org/10.7910/DVN/DBW86T), where lesion segmentation data will be available upon publication of this work. Test-set images with ground truth are available upon request from the ISIC archive (challenge@isic-archive.com).
Code availability
Code for the CNN is available upon request from the corresponding author for academic use.
References
Webster, P. Virtual health care in the era of COVID-19. Lancet 395, 1180–1181 (2020).
He, J. et al. The practical implementation of artificial intelligence technologies in medicine. Nat. Med. 25, 30–36 (2019).
McKinney, S. M. et al. International evaluation of an AI system for breast cancer screening. Nature 577, 89–94 (2020).
Gulshan, V. et al. Development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. JAMA 316, 2402–2410 (2016).
Esteva, A. et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542, 115–118 (2017).
Haenssle, H. A. et al. Man against machine: diagnostic performance of a deep learning convolutional neural network for dermoscopic melanoma recognition in comparison to 58 dermatologists. Ann. Oncol. 29, 1836–1842 (2018).
Han, S. S. et al. Classification of the clinical images for benign and malignant cutaneous tumors using a deep learning algorithm. J. Invest. Dermatol. 138, 1529–1538 (2018).
Marchetti, M. A. et al. Results of the 2016 International Skin Imaging Collaboration International Symposium on Biomedical Imaging challenge: comparison of the accuracy of computer algorithms to dermatologists for the diagnosis of melanoma from dermoscopic images. J. Am. Acad. Dermatol. 78, 270–277 (2018).
Tschandl, P. et al. Comparison of the accuracy of human readers versus machine-learning algorithms for pigmented skin lesion classification: an open, web-based, international, diagnostic study. Lancet Oncol. 20, 938–947 (2019).
Garg, A. X. et al. Effects of computerized clinical decision support systems on practitioner performance and patient outcomes: a systematic review. JAMA 293, 1223–1238 (2005).
Codella, N. C. F. et al. Collaborative human–AI (CHAI): evidence-based interpretable melanoma classification in dermoscopic images. In Understanding and Interpreting Machine Learning in Medical Image Computing Applications (eds., Kenji Suzuki, Mauricio Reyes, Tanveer Syeda-Mahmood, ETH Zurich, Ben Glocker, Roland Wiest, Yaniv Gur, Hayit Greenspan, Anant Madabhushi) 97–105 (Springer International Publishing, 2018).
Bien, N. et al. Deep-learning-assisted diagnosis for knee magnetic resonance imaging: development and retrospective validation of MRNet. PLoS Med. 15, e1002699 (2018).
Mobiny, A., Singh, A. & Van Nguyen, H. Risk-aware machine learning classifier for skin lesion diagnosis. J. Clin. Med. 8, 1241 (2019).
Han, S. S. et al. Augment intelligence dermatology: deep neural networks empower medical professionals in diagnosing skin cancer and predicting treatment options for 134 skin disorders. J. Invest. Dermatol. https://doi.org/10.1016/j.jid.2020.01.019 (2020).
Hekler, A. et al. Superior skin cancer classification by the combination of human and artificial intelligence. Eur. J. Cancer 120, 114–121 (2019).
Lakhani, P. & Sundaram, B. Deep learning at chest radiography: automated classification of pulmonary tuberculosis by using convolutional neural networks. Radiology 284, 574–582 (2017).
Tschandl, P., Rosendahl, C. & Kittler, H. The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci. Data 5, 180161 (2018).
Codella, N. et al. Skin lesion analysis toward melanoma detection 2018: a challenge hosted by the International Skin Imaging Collaboration (ISIC). Preprint at https://arxiv.org/abs/1902.03368 (2019).
Sadeghi, M., Chilana, P. K. & Atkins, M. S. How users perceive content-based image retrieval for identifying skin images. In Understanding and Interpreting Machine Learning in Medical Image Computing Applications (eds., Kenji Suzuki, Mauricio Reyes, Tanveer Syeda-Mahmood, ETH Zurich, Ben Glocker, Roland Wiest, Yaniv Gur, Hayit Greenspan, Anant Madabhushi) 141–148 (Springer International Publishing, 2018).
Tschandl, P., Argenziano, G., Razmara, M. & Yap, J. Diagnostic accuracy of content-based dermatoscopic image retrieval with deep classification features. Br. J. Dermatol. 181, 155–165 (2019).
Cai, C. J. et al. Human-centered tools for coping with imperfect algorithms during medical decision-making. In Proc. 2019 CHI Conference on Human Factors in Computing Systems 1–14 (Association for Computing Machinery, 2019).
Wang, M. & Deng, W. Deep visual domain adaptation: a survey. Neurocomputing 312, 135–153 (2018).
Finlayson, S.G. et al. Adversarial attacks on medical machine learning. Science 363, 1287–1289 (2019).
Navarrete-Dechent, C. et al. Automated dermatological diagnosis: hype or reality? J. Invest. Dermatol. 138, 2277–2279 (2018).
Winkler, J. K. et al. Association between surgical skin markings in dermoscopic images and diagnostic performance of a deep learning convolutional neural network for melanoma recognition. JAMA Dermatol. 155, 1135–1141 (2019).
Cai, C. J., Winter, S., Steiner, D., Wilcox, L. & Terry, M. ‘Hello AI’: uncovering the onboarding needs of medical practitioners for human–AI collaborative decision-making. In Proc. ACM on Human–Computer Interaction (Association for Computing Machinery, 2019).
Janda, M. et al. Accuracy of mobile digital teledermoscopy for skin self-examinations in adults at high risk of skin cancer: an open-label, randomised controlled trial. Lancet Digit. Health 2, e129–e137 (2020).
Gessert, N., Nielsen, M., Shaikh, M., Werner, R. & Schlaefer, A. Skin lesion classification using ensembles of multi-resolution EfficientNets with meta data. MethodsX 7, 100864 (2020).
Selvaraju, R. R. et al. Grad-CAM: visual explanations from deep networks via gradient-based localization. Int. J. Comput. Vis. 128, 336–359 (2020).
Li, X., Wu, J., Chen, E. Z. & Jiang, H. From deep learning towards finding skin lesion biomarkers. Conf. Proc. IEEE Eng. Med. Biol. Soc. 2019, 2797–2800 (2019).
Bissoto, A., Fornaciali, M., Valle, E. & Avila, S. (De)constructing bias on skin lesion datasets. Preprint at https://arxiv.org/abs/1904.08818 (2019).
Lapuschkin, S. et al. Unmasking Clever Hans predictors and assessing what machines really learn. Nat. Commun. 10, 1096 (2019).
Samek, W., Montavon, G., Vedaldi, A., Hansen, L. K. & Müller, K.-R. Explainable AI: Interpreting, Explaining and Visualizing Deep Learning (Springer Nature, 2019).
Paszke, A. et al. PyTorch: an imperative style, high-performance deep learning library. In Advances in Neural Information Processing Systems 32 (eds. Wallach, H. et al.) 8026–8037 (Curran Associates, 2019).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proc. IEEE Conference on Computer Vision and Pattern Recognition 770–778 (2016).
Russakovsky, O. et al. ImageNet large scale visual recognition challenge. Int. J. Comput. Vis. 115, 211–252 (2015).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. In Proc. 3rd International Conference for Learning Representations (eds., Bengio, Y., LeCun, Y.) (2015).
Barata, C., Celebi, M. E. & Marques, J. S. Improving dermoscopy image classification using color constancy. IEEE J. Biomed. Health Inform. 19, 1146–1152 (2015).
Rinner, C., Kittler, H., Rosendahl, C. & Tschandl, P. Analysis of collective human intelligence for diagnosis of pigmented skin lesions harnessed by gamification via a web-based training platform: simulation reader study. J. Med. Internet Res. 22, e15597 (2020).
Holm, S. A simple sequentially rejective multiple test procedure. Scand. J. Statist. 6, 65–70 (1979).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2019).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag, 2016).
Acknowledgements
We thank all dermachallenge.com users for their participation. We gratefully acknowledge the support of NVIDIA Corporation with the donation of the Titan Xp GPU used for this research.
Author information
Authors and Affiliations
Contributions
P.T., C.R. and H.K. conceived and designed experiments. P.T. trained the CNN and produced image predictions. C.R. and P.T. developed the web-based reader platforms together with H.K., and M.J. and H.P.S. provided data for the telemedicine study. H.K. and P.T. conducted statistical analyses. H.K., P.T., C.R., Z.A., G.A., N.C., A.H., M.J., A.L., C.L., J.M., J.P., S.P., C.R., H.P.S. and I.Z. helped collect rater data and interpreted findings. H.K., P.T., C.R., N.C. and A.H. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interests: P.T. received fees from Silverchair and an unrestricted 1-year postdoc grant from MetaOptima Technology. N.C. is an IBM employee and owns diverse investments across technology and health-care companies. A.H. is a consultant to Canfield Scientific and an advisory board member of Scibase. M.J. is funded by a National Health and Medical Research Council (NHMRC) TRIP Fellowship (APP1151021). H.P.S. is a shareholder of MoleMap and e-derm-consult and undertakes regular teledermatological reporting for both companies, is a medical consultant for Canfield Scientific and Revenio Research Oy and a medical advisor for First Derm and MetaOptima Technology, and has a Medical Advisory Board Appointment with MoleMap. H.P.S. holds an Australian NHMRC Practitioner Fellowship (APP1137127). All other authors report no conflict of interest in the topic of this manuscript. This project was conducted after ethical committee review at the Medical University of Vienna under protocol numbers 1804/2017, 1503/2018 and 2308/2019. All participants of the study platform agreed to academic research of usage data upon registration and have continuous ability to withdraw that consent.
Additional information
Peer review information Javier Carmona was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The neural network puts more relative attention to the non-lesion background in actinic keratoses.
Gradient-weighted Class Activation Maps (Grad-CAM, right column) for the top-1 prediction class of the CNN were created for all HAM10000 images, a sample for every ground-truth class is shown in the left column. The mean activation value per pixel of background- and lesion-area were estimated using manual segmentation masks (middle column). The quotient of background over lesion activation showed higher background activation for the predictions of the class AKIEC class versus all other classes (mean .48 vs. .32, p = 4.6 × 10−12, two-sided unpaired t-test). Thick central lines denote the median, lower and upper box limits the first and third quartiles, whiskers extend from the box to the most extreme value not further than 1.5 times the IQR.
Extended Data Fig. 2 Raters choose an asymmetric decision cutoff for malignancy.
a, When changing answers from benign to malignant (dark blue) or malignant to benign (light blue) diagnoses, the average cutoff for the AI-provided malignancy-probability was not 50% but <25% (yellow dotted line). b, On the ROC-curve for detecting malignant cases of the underlying AI (black line), this cutoff chosen inherently by the users (yellow dot), that is without instructions or prior knowledge about the AI accuracy, had a higher sensitivity and was closer to the ideal cutoff (blue dot), as measured by Youden’s index, than the ‘symmetric’ 50% cutoff (black dot).
Supplementary information
Supplementary Information
Supplementary Tables 1–4.
Rights and permissions
About this article
Cite this article
Tschandl, P., Rinner, C., Apalla, Z. et al. Human–computer collaboration for skin cancer recognition. Nat Med 26, 1229–1234 (2020). https://doi.org/10.1038/s41591-020-0942-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-020-0942-0
This article is cited by
-
Human-AI interaction in skin cancer diagnosis: a systematic review and meta-analysis
npj Digital Medicine (2024)
-
Diagnostic decisions of specialist optometrists exposed to ambiguous deep-learning outputs
Scientific Reports (2024)
-
Deep learning-aided decision support for diagnosis of skin disease across skin tones
Nature Medicine (2024)
-
Heterogeneity and predictors of the effects of AI assistance on radiologists
Nature Medicine (2024)
-
Artificial intelligence assisted food science and nutrition perspective for smart nutrition research and healthcare
Systems Microbiology and Biomanufacturing (2024)