Abstract
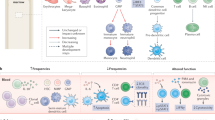
Understanding of the factors governing immune responses in cancer remains incomplete, limiting patient benefit. In this study, we used mass cytometry to define the systemic immune landscape in response to tumor development across five tissues in eight mouse tumor models. Systemic immunity was dramatically altered across models and time, with consistent findings in the peripheral blood of patients with breast cancer. Changes in peripheral tissues differed from those in the tumor microenvironment. Mice with tumor-experienced immune systems mounted dampened responses to orthogonal challenges, including reduced T cell activation during viral or bacterial infection. Antigen-presenting cells (APCs) mounted weaker responses in this context, whereas promoting APC activation rescued T cell activity. Systemic immune changes were reversed with surgical tumor resection, and many were prevented by interleukin-1 or granulocyte colony-stimulating factor blockade, revealing remarkable plasticity in the systemic immune state. These results demonstrate that tumor development dynamically reshapes the composition and function of the immune macroenvironment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All mass cytometry data are publicly available at https://premium.cytobank.org/cytobank/projects/2433/ or by request to the senior author without restrictions.
Code availability
The updated Statistical Scaffold package is available at https://github.com/SpitzerLab/statisticalScaffold.
Change history
02 April 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41591-024-02947-2
References
Philip, M. et al. Chromatin states define tumour-specific T cell dysfunction and reprogramming. Nature 545, 452–456 (2017).
Spitzer, M. H. et al. Systemic immunity is required for effective cancer immunotherapy. Cell 168, 487–502 (2017).
Fransen, M. F. et al. Tumor-draining lymph nodes are pivotal in PD-1/PD-L1 checkpoint therapy. JCI Insight 3, 1–7 (2018).
Tang, H. et al. PD-L1 on host cells is essential for PD-L1 blockade-mediated tumor regression. J. Clin. Invest. 128, 580–588 (2018).
Chamoto, K. et al. Mitochondrial activation chemicals synergize with surface receptor PD-1 blockade for T cell-dependent antitumor activity. PNAS 114, E761–E770 (2017).
Mathios, D. et al. Anti-PD-1 antitumor immunity is enhanced by local and abrogated by systemic chemotherapy in GBM. Sci. Transl. Med. 8, 370ra180 (2016).
Lin, H. et al. Host expression of PD-L1 determines efficacy of PD-L1 pathway blockade-mediated tumor regression. J. Clin. Invest. 128, 805–815 (2018).
Curiel, T. J. et al. Blockade of B7-H1 improves myeloid dendritic cell-mediated antitumor immunity. Nat. Med. 9, 562–567 (2003).
Yost, K. E. et al. Clonal replacement of tumor-specific T cells following PD-1 blockade. Nat. Med. 25, 1251–1259 (2019).
McAllister, S. S. & Weinberg, R. A. The tumour-induced systemic environment as a critical regulator of cancer progression and metastasis. Nat. Cell Biol. 16, 717–727 (2014).
Zhang, S. et al. The role of myeloid-derived suppressor cells in patients with solid tumors: a meta-analysis. PLoS ONE 11, e0164514 (2016).
Casbon, A.-J. et al. Invasive breast cancer reprograms early myeloid differentiation in the bone marrow to generate immunosuppressive neutrophils. Proc. Natl Acad. Sci. USA 112, E566–E575 (2015).
Meyer, M. A. et al. Breast and pancreatic cancer interrupt IRF8-dependent dendritic cell development to overcome immune surveillance. Nat. Commun. 9, 1–19 (2018).
Barnstorf, I. et al. Chronic virus infection compromises memory bystander T cell function in an IL-6/ STAT1-dependent manner. J. Exp. Med. 216, 571–586 (2019).
Snell, L. M. et al. CD8+ T cell priming in established chronic viral infection preferentially directs differentiation of memory-like cells for sustained immunity. Immunity 49, 678–694 (2018).
Osborne, L. C. et al. Virus-helminth coinfection reveals a microbiota-independent mechanism of immunomodulation. Science 345, 578–582 (2014).
Danna, E. A. et al. Surgical removal of primary tumor reverses tumor-induced immunosuppression despite the presence of metastatic disease. Cancer Res. 64, 2205–2211 (2004).
Ghochikyan, A. et al. Primary 4T1 tumor resection provides critical ‘window of opportunity’ for immunotherapy. Clin. Exp. Metastasis 31, 185–198 (2014).
Mosely, S. I. S. et al. Rational selection of syngeneic preclinical tumor models for immunotherapeutic drug discovery. Cancer Immunol. Res. 5, 29–41 (2017).
Westcott, P. M. K. et al. The mutational landscapes of genetic and chemical models of Kras-driven lung cancer. Nature 517, 489–492 (2015).
Zeitouni, B. et al. Abstract 1840: Whole-exome somatic mutation analysis of mouse cancer models and implications for preclinical immunomodulatory drug development. In Proceedings of the 107th Annual Meeting of the American Association for Cancer Research https://doi.org/10.1158/1538-7445.AM2017-1840 (AACR, 2017).
Heinzel, F. P., Sadick, M. D., Holaday, B. J., Coffman, R. L. & Locksley, R. M. Reciprocal expression of interferon γ or interleukin 4 during the resolution or progression of murine leishmaniasis. Evidence for expansion of distinct helper T cell subsets. J. Exp. Med. 169, 59–72 (1989).
Kather, J. N. et al. Topography of cancer-associated immune cells in human solid tumors. eLife 7, e36967 (2018).
Spitzer, M. H. et al. An interactive reference framework for modeling a dynamic immune system. Science 349, 1259425 (2015).
Anz, D. et al. CD103 is a hallmark of tumor-infiltrating regulatory T cells. Int. J. Cancer 129, 2417–2426 (2011).
Ross, E. A. et al. CD31 is required on CD4+ T cells to promote T cell survival during Salmonella infection. J. Immunol. 187, 1553–1565 (2011).
Hänninen, A., Maksimow, M., Alam, C., Morgan, D. J. & Jalkanen, S. Ly6C supports preferential homing of central memory CD8+ T cells into lymph nodes. Eur. J. Immunol. 41, 634–644 (2011).
Fourcade, J. et al. Upregulation of Tim-3 and PD-1 expression is associated with tumor antigen-specific CD8+ T cell dysfunction in melanoma patients. J. Exp. Med. 207, 2175–2186 (2010).
Mita, Y. et al. Crucial role of CD69 in anti-tumor immunity through regulating the exhaustion of tumor-infiltrating T cells. Int. Immunol. 30, 559–567 (2018).
Sun, C., Mezzadra, R. & Schumacher, T. N. Regulation and function of the PD-L1 checkpoint. Immunity 48, 434–452 (2018).
Bianchini, M. et al. PD-L1 expression on nonclassical monocytes reveals their origin and immunoregulatory function. Sci. Immunol. 4, eaar3054 (2019).
Busch, D. H., Pilip, I. M., Vijh, S. & Pamer, E. G. Coordinate regulation of complex T cell populations responding to bacterial infection. Immunity 8, 353–362 (1998).
Kaech, S. M. & Ahmed, R. Memory CD8+ T cell differentiation: initial antigen encounter triggers a developmental program in naïve cells. Nat. Immunol. 2, 415–422 (2001).
Herndler-Brandstetter, D. et al. KLRG1+ effector CD8+ T cells lose KLRG1, differentiate into all memory T cell lineages, and convey enhanced protective immunity. Immunity 48, 716–729 (2018).
Jung, S. et al. In vivo depletion of CD11c+ dendritic cells abrogates priming of CD8+ T cells by exogenous cell-associated antigens. Immunity 17, 211–220 (2002).
Gabrilovich, D. I., Corak, J., Ciernik, I. F., Kavanaugh, D. & Carbone, D. P. Decreased antigen presentation by dendritic cells in patients with breast cancer. Clin. Cancer Res. 3, 483–490 (1997).
Coffelt, S. B. et al. IL-17-producing γδ T cells and neutrophils conspire to promote breast cancer metastasis. Nature 522, 345–348 (2015).
Wu, W.-C. et al. Circulating hematopoietic stem and progenitor cells are myeloid-biased in cancer patients. Proc. Natl Acad. Sci. USA 111, 4221–4226 (2014).
Apte, R. N. et al. Effects of micro-environment- and malignant cell-derived interleukin-1 in carcinogenesis, tumour invasiveness and tumour-host interactions. Eur. J. Cancer 42, 751–759 (2006).
Wu, T. C. et al. IL1 receptor antagonist controls transcriptional signature of inflammation in patients with metastatic breast cancer. Cancer Res. 78, 5243–5258 (2018).
Singer, C. F. et al. Interleukin-1ɑ protein secretion in breast cancer is associated with poor differentiation and estrogen receptor ɑ negativity. Int. J. Gynecol. Cancer 16, 556–559 (2006).
Pickup, M., Novitskiy, S. & Moses, H. L. The roles of TGFβ in the tumour microenvironment. Nat. Rev. Cancer 13, 788–799 (2013).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Suzuki, A. et al. IL-1 production as a regulator of G-CSF and IL-6 production in CSF-producing cell lines. Br. J. Cancer 65, 515–518 (1992).
Mittal, R., Wagener, M., Breed, E. R., Liang, Z. & Yoseph, B. P. Phenotypic T cell exhaustion in a murine model of bacterial infection in the setting of pre-existing malignancy. PLoS ONE 9, 93523 (2014).
Xie, J. et al. Pre-existing malignancy results in increased prevalence of distinct populations of CD4+ T cells during sepsis. PLoS ONE 13, e0191065 (2018).
Russ, A. J. et al. Melanoma-induced suppression of tumor antigen-specific T cell expansion is comparable to suppression of global T cell expansion. Cell. Immunol. 271, 104–109 (2011).
Klastersky, J. & Aoun, M. Opportunistic infections in patients with cancer. Ann. Oncol. 15, iv329–iv335 (2004).
Baluch, A. & Pasikhova, Y. Influenza vaccination in oncology patients. Curr. Infect. Dis. Rep. 15, 486–490 (2013).
O’Hara, M. H. et al. Abstract CT004: A Phase Ib study of CD40 agonistic monoclonal antibody APX005M together with gemcitabine (Gem) and nab-paclitaxel (NP) with or without nivolumab (Nivo) in untreated metastatic ductal pancreatic adenocarcinoma (PDAC) patients. Clin. Trials https://doi.org/10.1158/1538-7445.am2019-ct004 (2019).
Zuckerman, N. S. et al. Altered local and systemic immune profiles underlie lymph node metastasis in breast cancer patients. Int. J. Cancer 132, 2537–2547 (2012).
Wang, L. et al. Connecting blood and intratumoral Treg cell activity in predicting future relapse in breast cancer. Nat. Immunol. 20, 1220–1230 (2019).
Kosaka, A., Ohkuri, T., Program, B. T. & Okada, H. Combination of an agonistic anti-CD40 monoclonal antibody and the COX-2 inhibitor celecoxib induces anti-glioma effects by promotion of type-1 immunity in myeloid cells and T-cells. Cancer Immunol. Immunother. 63, 847–857 (2014).
Tseng, W. W. et al. Development of an orthotopic model of invasive pancreatic cancer in an immunocompetent murine host. Clin. Cancer Res. 16, 3684–3695 (2010).
Kathryn, E. et al. Cutting edge: CD4 and CD8 T cells are intrinsically different in their proliferative responses. J. Immunol. 168, 1528–1532 (2002).
Zunder, E. R. et al. Palladium-based mass-tag cell barcoding with a doublet-filtering scheme and single cell deconvolution algorithm. Nat. Protoc. 10, 316–333 (2015).
Finck, R. et al. Normalization of mass cytometry data with bead standards. Cytometry A 83, 483–494 (2013).
Bair, E. & Tibshirani, R. Semi-supervised methods to predict patient survival from gene expression data. PLoS Biol. 2, 0511–0522 (2004).
Dumeaux, V. et al. Interactions between the tumor and the blood systemic response of breast cancer patients. PLoS Comput. Biol. 13, e1005680 (2017).
Aran, D., Hu, Z. & Butte, A. J. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 18, 1–14 (2017).
Acknowledgements
We thank the UCSF Flow Cytometry Core and S. Tamaki for CyTOF maintenance and M.H. Barcellos-Hoff, R. Levine, H. Okada, E. Engleman and J. Bluestone for cell lines, transgenic mice and reagents. We thank L. Lanier, Z. Werb, M.H. Barcellos-Hoff and L. Fong for insightful feedback. This work was supported by National Institutes of Health (NIH) grants DP5OD023056 and P50CA097257 (UCSF Brain Tumor SPORE Developmental Research Program), funds from the UCSF Program for Breakthrough Biomedical Research and investigator funding from the Parker Institute for Cancer Immunotherapy to M.H.S. and by NIH grant S10OD018040, which enabled procurement of the mass cytometer used in this study. This study makes use of data generated by the Norwegian Women and Cancer Study. A full list of investigators who contributed to the generation of the data is available at http://site.uit.no/nowac/. Funding for the project was provided by European Research Council grant ERC-2008-AdG 232997. The Norwegian Women and Cancer Study group is not responsible for the analysis or interpretation of the data presented.
Author information
Authors and Affiliations
Contributions
Conceptualization: B.M.A, K.J.H., Y.C. and M.H.S.; experimental methodology: B.M.A., K.J.H., C.E.B., A.V., R.D., I.T., D.M.M., N.W.C., Y.C. and M.H.S.; computational methodology: B.M.A. and M.H.S.; investigation: all authors; writing, original draft: B.M.A.; writing, review and editing: all authors; funding acquisition: M.H.S.; supervision: M.H.S.
Corresponding author
Ethics declarations
Competing interests
M.H.S. receives research funding from Roche/Genentech, Bristol-Myers Squibb and Valitor and has been a paid consultant for Five Prime Therapeutics, Ono Pharmaceutical and January Inc.
Additional information
Peer review information Saheli Sadanand was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Main Mass Cytometry Gating Scheme.
a, Main gating strategy for identifying major immune cell populations from mass cytometry datasets.
Extended Data Fig. 2 Systemic immunity is distinctly remodeled across tumor models.
a, Relative abundance of total leukocytes infiltrating the TME across eight tumor models. b-f, Scaffold maps of spleen cell frequencies across five distinct tumor models, SB28 glioblastoma (b), MC38 colorectal (c), LMP pancreatic (d), B16 melanoma (e), and Braf/PTEN melanoma (f), comparing late stage tumor burden to their respective health littermate controls. g, Heatmaps of the log2 adjusted fold change in bulk immune cell frequencies across all five tissues, where relevant, across all models. h, Heatmaps of the log2 adjusted fold change in bulk immune cell frequencies comparing the parental LLC and engineered LLC expressing reporters GFP and Luciferase, with cell labels in g. Lower inset shows Pearson’s correlation between these systemic immune features.
Extended Data Fig. 3 Systemic immunity is distinctly remodeled over tumor development.
a, Pearson’s correlation between MMTV-PyMT primary tumor size and change in systemic immune composition, measured as Aitchison distance. b, Degree of systemic immune change by Aitchison distance over tumor growth (left) and after removing the contribution of primary tumor size by linear regression (right). c, Percent of PyMT expressing metastatic cancer cells in the lung (green) and primary draining lymph node (blue). d, Pearson’s correlation between lung or lymph node metastasis and the residual changes in systemic immune composition after regressing out primary tumor burden. e, Heatmap of the log2 adjusted fold change in bulk spleen immune cell frequencies for each 400mm2 tumor-bearing mouse, ranging from 0 to high metastatic disease. f, Pearson’s correlation between tumor mass and absolute number of infiltrating leukocytes in 4T1 breast tumors. g, Spleen immune absolute cell counts, adjusted absolute cell counts per mg of tissue, and unadjusted immune frequencies at each time point for neutrophils, B cells and T cells of the 4T1 breast tumor model. h, PCA of relative immune cell frequencies from each major immune tissue over time in the MMTV-PyMT breast tumor model. Vectors designate progression from control (first point) to 25 mm2, 50mm2, 125mm2, and 400mm2 (last point, arrowhead). i, Scaffold maps of immune cell frequencies in the spleen at each time point of 4T1 tumor burden, colored by log2 fold change in frequency compared to the previous time point.
Extended Data Fig. 4 Immunity is distinctly remodeled by compartment over tumor development.
a-d, Scaffold maps of immune cell frequencies over 4T1 tumor progression in the tumor draining lymph node (a) blood (b), bone marrow (c), and tumor (d), colored by fold change from the previous time point.
Extended Data Fig. 5 Tumor growth shifts the systemic T cell composition across models.
a-b, PCA of T cell cluster frequencies across lymphoid tissues over tumor development for the 4T1 (a) and MMTV-PyMT (b) breast tumor models. Vectors designate directional progression from control (first point) to late stage disease (last point, arrowhead). In a, tumor time points include day 7, 14, 21, and 35 after 4T1 cancer cell transplant. In b, tumor time points include tumor sizes of 25 mm2, 50 mm2, 125 mm2, and 400 mm2. c-e, CD3 + CD11b- leukocytes from all tissues clustered together from healthy and MMTV-PyMT tumor-burdened animals at progressive tumor sizes. c, Heatmap of each T cell cluster frequency, by row, in each site and across the individual 2-3 animals per time point. d, Stacked bar plot of the log2 fold change in cluster frequency between early (25 mm2) and late (400 mm2) disease time points, colored by tissue. e, Heatmap of the protein expression defining each T cell cluster, column normalized to each protein’s maximum positive expression. f-h, Representative scatter plots of key proteins that define T cell clusters changing in frequency in the designated site between early and late disease stage for CD8 T cells (f), Tregs (g), and CD4 T Cells (h).
Extended Data Fig. 6 Tumor growth shifts the systemic mononuclear phagocyte composition.
a, CD3- CD19- leukocytes from all tissues clustered together from healthy and 4T1 tumor-burdened animals at progressive time points. Left, stacked bar plot of the log2 fold change in cluster frequency between early (day 7) and late (day 35) times points, colored by tissue. Right, heatmap of the protein expression defining each cluster, column normalized to each protein’s maximum positive expression. b, Curves of the mean cell frequencies over time in the 4T1 breast tumor model from designated mononuclear phagocyte cell types, colored by tissue. c, PCA of the mononuclear phagocyte cell frequencies from each tissue over time in the 4T1 breast tumor model. Vectors designate progression from control (first point) to day 7, 14, 21, and 35 (last point, arrowhead). Coloring of tissues for a-c corresponds to labels in c.
Extended Data Fig. 7 PD-1 and PD-L1 expression is dynamic over tumor growth.
a, Distribution of PD-1 and PD-L1 signal intensities on tumor infiltrating leukocytes over time in the 4T1 or LLC tumor models. Coloring of time points for a-d corresponds to legend in a. b, Percent of total infiltrating leukocytes (left of dashed line) or CD45−, non-endothelial cells (right of dashed line) with high PD-1 or PD-L1 expression in the 4T1 or LLC tumor models. c, Percent of leukocytes with high PD-1 or PD-L1 expression over time and across tissues, 4T1 model. d, Pearson’s correlation between median PD-L1 signal intensity on blood versus tumor infiltrating leukocytes, 4T1 model. e, Percent of each major immune cell subset expressing high PD-1 or PD-L1 in the tumor, blood, and spleen, identified manually. Cell subsets below 0.2% of total leukocytes were not included, X. Bars ordered by time point, beginning at healthy control. Double positive PD-1/PD-L1 expression was rare and not illustrated. p*< 0.05, One-Way ANOVA, with Tukey correction versus control tissue or healthy mammary fat pad (blue in b-c, fill corresponding to bar color in e), or versus day 7 (green in b-c).
Extended Data Fig. 8 Tumor burden induces tissue-specific changes in immune cell cycling.
a-b, Log2 fold change in bulk Ki67 expressing leukocytes in each tissue tissues for 4T1, LLC and MMTV tumors (a), and over 4T1 tumor progression (b). p*< 0.05, One-Way ANOVA, with Tukey correction versus control. c-d, Statistical Scaffold maps of Ki67 expression in immune cells of the tumor draining lymph node comparing control to day 21 (c) and the Spleen over time (d) in 4T1 tumor burdened animals. e, Percent of increasing clusters (red, total of 56), decreasing clusters (blue, total of 90), or unchanged cluster that have corresponding changes in cell cycle markers Ki67 and cleaved Caspase-3.
Extended Data Fig. 9 Tumor driven deficits in T cell responses are cell-extrinsic.
a, Quantification of bulk CD8+ T cell populations in the spleen of healthy or LLC tumor-burdened mice after 7 days of Lm infection, Two-Way ANOVA with Bonferroni correction. b, Expression of inflammatory cytokines, INFy, IL-2, and TNFa in splenic CD8 T Cells isolated from control or LLC tumor-burdened mice after in vitro differentiation with CD3, CD28 and IL-2, and re-stimulation with BrefeldinA and PMA Ionomycin. c, Scatter plots of CD11b and Ly6G showing expected neutrophilia in OT-I TCR transgenic mice with LLC tumor burden. d, Histograms of CD80, CD86, and CD83 signal intensity on cDCs from healthy or LLC tumor-burdened mice at day 2 of Lm-OVA infection. e, Median signal intensity of CD80, PD-L1 and CD54 activation markers on splenic cDCs from healthy or LLC tumor-burdened mice compared to IL-12p70 or CD40 treatment at day 7 of Lm-OVA infection. F, Median signal intensity of PD-L1 on splenic cDCs from untreated or CD40 treated LLC tumor-burdened (day 21) mice. G, Quantification of splenic CD8 + T cell proliferation in healthy, untreated or CTLA-4 treated LLC tumor-burdened animals in response to 7 days of Lm-OVA infection. p*<0.05, two-tailed t-test.
Extended Data Fig. 10 Tumor resection resets systemic immune organization and function.
a-c, Statistical scaffold maps of spleen immune cell frequencies (a) and proliferation by Ki67 expression (b) in 4T1 resected mice, and of spleen immune cell frequencies in MC38 resected mice (c) compared to healthy control. Insets show resected mice compared to tumor-burdened mice. d-e, Heatmap of the log2 fold changes in splenic immune cell frequencies for local or lung recurrences from control mice (d), and for IL-1, G-CSF, or TGFβ blockade from untreated LLC tumor-burdened mice (e). f-g, Heatmaps of T cell cluster expression profiles and log2 fold change from control for LLC (f) and 4T1 (g) for the spleen and draining lymph node. h, Median signal intensity of CD86 and PD-L1 on splenic cDCs from healthy, LLC tumor-burdened, resected, or resected mice with local recurrence at day 7 of Lm-OVA infection. i, Concentration of circulating cytokines, IL-1α and G-CSF from healthy, LLC tumor-burdened, resected, or resected mice with local recurrence. j, Concentration of cytokines, IL-1α, IL-1β and G-CSF from in vitro cell culture media conditioned with LLC cancer cells. k, Concentration of circulating G-CSF from control or LLC tumor-bearing mice, or LLC tumor-bearing mice treated with either IL-1 or G-CSF blocking antibodies. p*<0.05, two-tailed t-test.
Supplementary information
Supplementary Information
Supplementary Tables 1–3
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Allen, B.M., Hiam, K.J., Burnett, C.E. et al. Systemic dysfunction and plasticity of the immune macroenvironment in cancer models. Nat Med 26, 1125–1134 (2020). https://doi.org/10.1038/s41591-020-0892-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-020-0892-6
This article is cited by
-
Harnessing 3D in vitro systems to model immune responses to solid tumours: a step towards improving and creating personalized immunotherapies
Nature Reviews Immunology (2024)
-
New immune cell engagers for cancer immunotherapy
Nature Reviews Immunology (2024)
-
The CUL5 E3 ligase complex negatively regulates central signaling pathways in CD8+ T cells
Nature Communications (2024)
-
Combinational delivery of TLR4 and TLR7/8 agonist enhanced the therapeutic efficacy of immune checkpoint inhibitors to colon tumor
Molecular and Cellular Biochemistry (2024)
-
Targeting the epigenome to reinvigorate T cells for cancer immunotherapy
Military Medical Research (2023)