Abstract
Mucociliary clearance, the physiological process by which mammalian conducting airways expel pathogens and unwanted surface materials from the respiratory tract, depends on the coordinated function of multiple specialized cell types, including basal stem cells, mucus-secreting goblet cells, motile ciliated cells, cystic fibrosis transmembrane conductance regulator (CFTR)-rich ionocytes, and immune cells1,2. Bronchiectasis, a syndrome of pathological airway dilation associated with impaired mucociliary clearance, may occur sporadically or as a consequence of Mendelian inheritance, for example in cystic fibrosis, primary ciliary dyskinesia (PCD), and select immunodeficiencies3. Previous studies have identified mutations that affect ciliary structure and nucleation in PCD4, but the regulation of mucociliary transport remains incompletely understood, and therapeutic targets for its modulation are lacking. Here we identify a bronchiectasis syndrome caused by mutations that inactivate NIMA-related kinase 10 (NEK10), a protein kinase with previously unknown in vivo functions in mammals. Genetically modified primary human airway cultures establish NEK10 as a ciliated-cell-specific kinase whose activity regulates the motile ciliary proteome to promote ciliary length and mucociliary transport but which is dispensable for normal ciliary number, radial structure, and beat frequency. Together, these data identify a novel and likely targetable signaling axis that controls motile ciliary function in humans and has potential implications for other respiratory disorders that are characterized by impaired mucociliary clearance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Sequence data that support the findings of this study have been deposited in NCBI GenBank under accession numbers MK806425 and MK806426. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium through the PRIDE partner repository with the dataset identifier PXD01660061. Plasmids pLRC1-NEK10p:NEK10-3XFLAG and pLRC1-FOXJ1p:NEK10-3XFLAG are available for review and distribution through Addgene (plasmid numbers 137030 and 137031). All other data and computer code are provided within the paper or in the Supplementary information. Raw data for statistical tests (.xlsx files) and uncropped immunoblots that correspond to to Figs. 1–4 and Extended Data Figs. 2,3,5 (.pdf files) are provided.
Change history
29 January 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41591-020-0773-z
References
Tilley, A. E., Walters, M. S., Shaykhiev, R. & Crystal, R. G. Cilia dysfunction in lung disease. Annu. Rev. Physiol. 77, 379–406 (2015).
Montoro, D. T. et al. A revised airway epithelial hierarchy includes CFTR-expressing ionocytes. Nature 560, 319–324 (2018).
Gould, C. M., Freeman, A. F. & Olivier, K. N. Genetic causes of bronchiectasis. Clin. Chest Med. 33, 249–263 (2012).
Zariwala, M. A., Knowles, M. R. & Omran, H. Genetic defects in ciliary structure and function. Annu. Rev. Physiol. 69, 423–450 (2007).
Online Mendelian Inheritance in Man, OMIM (McKusick–Nathans Institute of Genetic Medicine, Johns Hopkins University, November 2019); https://www.omim.org/
Moniz, L., Dutt, P., Haider, N. & Stambolic, V. Nek family of kinases in cell cycle, checkpoint control and cancer. Cell Div. 6, 18 (2011).
Thiel, C. et al. NEK1 mutations cause short-rib polydactyly syndrome type Majewski. Am. J. Hum. Genet. 88, 106–114 (2011).
Smith, L. A. et al. Development of polycystic kidney disease in juvenile cystic kidney mice: insights into pathogenesis, ciliary abnormalities, and common features with human disease. J. Am. Soc. Nephrol. 17, 2821–2831 (2006).
Moniz, L. S. & Stambolic, V. Nek10 mediates G2/M cell cycle arrest and MEK autoactivation in response to UV irradiation. Mol. Cell. Biol. 31, 30–42 (2011).
Porpora, M. et al. Counterregulation of cAMP-directed kinase activities controls ciliogenesis. Nat. Commun. 9, 1224 (2018).
Fulcher, M. L., Gabriel, S., Burns, K. A., Yankaskas, J. R. & Randell, S. H. Well-differentiated human airway epithelial cell cultures. Methods Mol. Med. 107, 183–206 (2005).
Knowles, M. R., Zariwala, M. & Leigh, M. Primary ciliary dyskinesia. Clin. Chest Med. 37, 449–461 (2016).
Karczewski, K. J. et al. Variation across 141,456 human exomes and genomes reveals the spectrum of loss-of-function intolerance across human protein-coding genes. bioRxivorg 49, 531210 (2019).
Ostrowski, L. E., Hutchins, J. R., Zakel, K. & O’Neal, W. K. Targeting expression of a transgene to the airway surface epithelium using a ciliated cell-specific promoter. Mol. Ther. 8, 637–645 (2003).
Liu, L. et al. Method for quantitative study of airway functional microanatomy using micro-optical coherence tomography. PLoS ONE 8, e54473 (2013).
Knowles, M. R., Daniels, L. A., Davis, S. D., Zariwala, M. A. & Leigh, M. W. Primary ciliary dyskinesia. Recent advances in diagnostics, genetics, and characterization of clinical disease. Am. J. Respir. Crit. Care Med. 188, 913–922 (2013).
He, Y., Zeng, M. Y., Yang, D., Motro, B. & Nuñez, G. NEK7 is an essential mediator of NLRP3 activation downstream of potassium efflux. Nature 530, 354–357 (2016).
Carrera, A. C., Alexandrov, K. & Roberts, T. M. The conserved lysine of the catalytic domain of protein kinases is actively involved in the phosphotransfer reaction and not required for anchoring ATP. Proc. Natl Acad. Sci. USA 90, 442–446 (1993).
Moniz, L. Characterization of NimA-related Kinase 10 (NEK10): A Role in Checkpoint Control. PhD thesis, Univ. of Toronto (2010).
Richards, M. W. et al. An autoinhibitory tyrosine motif in the cell-cycle-regulated Nek7 kinase is released through binding of Nek9. Mol. Cell 36, 560–570 (2009).
Doan, M. et al. Diagnostic potential of imaging flow cytometry. Trends Biotechnol. 36, 649–652 (2018).
Wallmeier, J. et al. Mutations in CCNO result in congenital mucociliary clearance disorder with reduced generation of multiple motile cilia. Nat. Genet. 46, 646–651 (2014).
Boon, M. et al. MCIDAS mutations result in a mucociliary clearance disorder with reduced generation of multiple motile cilia. Nat. Commun. 5, 4418 (2014).
Vladar, E. K., Nayak, J. V., Milla, C. E. & Axelrod, J. D. Airway epithelial homeostasis and planar cell polarity signaling depend on multiciliated cell differentiation. JCI Insight 1, 183 (2016).
Ostrowski, L. E. in Cell Biology 3rd edn, Vol. 2 (ed., Celis, J. E.) Ch. 14 (Elsevier, 2005).
Leopold, P. L., O’Mahony, M. J., Lian, X. J., Tilley, A. E., Harvey, B.-G. & Crystal, R. G. Smoking is associated with shortened airway cilia. PLoS ONE 4, e8157 (2009).
Oltean, A., Schaffer, A. J., Bayly, P. V. & Brody, S. L. Quantifying ciliary dynamics during assembly reveals stepwise waveform maturation in airway cells. Am. J. Respir. Cell Mol. Biol. 59, 511–522 (2018).
Bottier, M., Thomas, K. A., Dutcher, S. K. & Bayly, P. V. How does cilium length affect beating? Biophys. J. 116, 1292–1304 (2019).
Block, H. et al. Immobilized-metal affinity chromatography (IMAC): a review. Meth. Enzymol. 463, 439–473 (2009).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
The Gene Ontology Consortium. The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res. 47, D330–D338 (2019).
Ostrowski, L. E. et al. A proteomic analysis of human cilia identification of novel components. Mol. Cell Proteomics 1, 451–465 (2002).
Vieira Braga, F. A. et al. A cellular census of human lungs identifies novel cell states in health and in asthma. Nat. Med. 25, 1153–1163 (2019).
Tabula Muris Consortium. et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Wloga, D. et al. Members of the NIMA-related kinase family promote disassembly of cilia by multiple mechanisms. Mol. Biol. Cell 17, 2799–2810 (2006).
Bradley, B. A. & Quarmby, L. M. A NIMA-related kinase, Cnk2p, regulates both flagellar length and cell size in Chlamydomonas. J. Cell Sci. 118, 3317–3326 (2005).
Hilton, L. K., Gunawardane, K., Kim, J. W., Schwarz, M. C. & Quarmby, L. M. The kinases LF4 and CNK2 control ciliary length by feedback regulation of assembly and disassembly rates. Curr. Biol. 23, 2208–2214 (2013).
Lin, H. et al. A NIMA-related kinase suppresses the flagellar instability associated with the loss of multiple axonemal structures. PLoS Genet. 11, e1005508 (2015).
Hessel, J. et al. Intraflagellar transport gene expression associated with short cilia in smoking and COPD. PLoS ONE 9, e85453 (2014).
Chen, Z.-G. et al. Aberrant epithelial remodeling with impairment of cilia architecture in non-cystic fibrosis bronchiectasis. J. Thorac. Dis. 10, 1753–1764 (2018).
Neuberger, T., Burton, B., Clark, H. & Van Goor, F. Use of primary cultures of human bronchial epithelial cells isolated from cystic fibrosis patients for the pre-clinical testing of CFTR modulators. Methods Mol. Biol. 741, 39–54 (2011).
Carr, I. M. et al. Interactive visual analysis of SNP data for rapid autozygosity mapping in consanguineous families. Hum. Mutat. 27, 1041–1046 (2006).
Hoffmann, K. & Lindner, T. H. easyLINKAGE-Plus—automated linkage analyses using large-scale SNP data. Bioinformatics 21, 3565–3567 (2005).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015).
Liu, L. et al. Imaging the subcellular structure of human coronary atherosclerosis using micro-optical coherence tomography. Nat. Med. 17, 1010–1014 (2011).
Sbalzarini, I. F. & Koumoutsakos, P. Feature point tracking and trajectory analysis for video imaging in cell biology. J. Struct. Biol. 151, 182–195 (2005).
Eng, J. K., McCormack, A. L. & Yates, J. R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Reiter, J. F. & Leroux, M. R. Genes and molecular pathways underpinning ciliopathies. Nat. Rev. Mol. Cell Biol. 18, 533–547 (2017).
Ishikawa, H. & Marshall, W. F. Ciliogenesis: building the cell’s antenna. Nat. Rev. Mol. Cell Biol. 12, 222–234 (2011).
Teves, M. E., Nagarkatti-Gude, D. R., Zhang, Z. & Strauss, J. F. Mammalian axoneme central pair complex proteins: broader roles revealed by gene knockout phenotypes. Cytoskeleton 73, 3–22 (2016).
Osinka, A. et al. Ciliary proteins: filling the gaps. Recent advances in deciphering the protein composition of motile ciliary complexes. Cells 8, 730 (2019).
Zhao, L., Hou, Y., Picariello, T., Craige, B. & Witman, G. B. Proteome of the central apparatus of a ciliary axoneme. J. Cell Biol. 218, 2051–2070 (2019).
Satish Tammana, T. V., Tammana, D., Diener, D. R. & Rosenbaum, J. Centrosomal protein CEP104 (Chlamydomonas FAP256) moves to the ciliary tip during ciliary assembly. J. Cell Sci. 126, 5018–5029 (2013).
Niwa, S. et al. KIF19A is a microtubule-depolymerizing kinesin for ciliary length control. Dev. Cell 23, 1167–1175 (2012).
Lai, C. K. et al. Functional characterization of putative cilia genes by high-content analysis. Mol. Biol. Cell 22, 1104–1119 (2011).
Vasudevan, K. K. et al. Kinesin-13 regulates the quantity and quality of tubulin inside cilia. Mol. Biol. Cell 26, 478–494 (2015).
Piao, T. et al. A microtubule depolymerizing kinesin functions during both flagellar disassembly and flagellar assembly in Chlamydomonas. Proc. Natl Acad. Sci. USA 106, 4713–4718 (2009).
Wang, L. et al. Flagellar regeneration requires cytoplasmic microtubule depolymerization and kinesin-13. J. Cell Sci. 126, 1531–1540 (2013).
Broekhuis, J. R., Verhey, K. J. & Jansen, G. Regulation of cilium length and intraflagellar transport by the RCK-kinases ICK and MOK in renal epithelial cells. PLoS ONE 9, e108470 (2014).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Acknowledgements
We are grateful to the patients and family members who participated in this research. We thank J. Rajagopal for advice and reagents, B. Stripp (Cedars-Sinai) and I. Cheeseman (Whitehead Institute for Biomedical Research) for antibodies, the Peterson (1957) Nanotechnology Materials Core Facility at the Koch Institute for electron microscopy services, S. Mordecai for advice and assistance with IFC, H. Zheng for assistance with statistical methods, M. Mino-Kenudson for pulmonary pathology assistance, KFSHRC genotyping and sequencing core facilities for technical help, M. Manion and the Primary Ciliary Dyskinesia Foundation for support, L. Ostrowski for assistance with ciliary motility studies, K. Sullivan and N. Capps for patient coordination and specimen collection, V. Madden, J. Stonebraker, R. Pace, and K. Burns for technical assistance, H. Dang and H. Namkoong for bioinformatics assistance, and A. Lee and members of the Sabatini laboratory for critical reading of the manuscript. This work was supported by grants from the National Institutes of Health: T32HL116275 (R.R.C.), T32CA009216 (M.S.T.), U54HL096458 and UL1TR000083 (M.R.K., M.A.Z., M.L.D., and P.R.S.), R01HL071798 (M.R.K., M.L.D., and M.A.Z.), R01HL117836-01 (M.R.K., M.L.D., and M.A.Z.), R01CA129105 and R37AI047389 (D.M.S.). Additional funding support was provided by the Massachusetts General Hospital Department of Medicine Pathways program (R.R.C.) and by the King Salman Center for Disability Research and the Saudi Human Genome Program (F.S.A.). The Amnis ImageStreamX Mk II was purchased using a National Institutes of Health Shared Instrumentation Grant 1S10OD012027-01A1 to Massachusetts General Hospital. D.M.S. is an investigator at the Howard Hughes Medical Institute and an American Cancer Society Research Professor.
Author information
Authors and Affiliations
Contributions
R.R.C. initiated the project, phenotyped the index proband, designed and performed all the experiments except as noted, analyzed the data, prepared the figures, and wrote the manuscript. D.T.M. assisted with the design and performance of the cell culture, IFM, and FACS experiments, analyzed the data, and edited the manuscript. H.M.L. performed the μOCT experiments, analyzed the data, and prepared the figures. J.Y. assisted with molecular cloning, site-directed mutagenesis, and cell culture, and edited the manuscript. H.E.S. performed the whole exome sequencing and linkage analysis on kindreds 1–3. M.S.T. acquired the clinical histopathology images and prepared the figures. G.W.D. performed the sequencing, molecular biology, and high-speed video microscopy (HSVM) analysis on kindred 4. M.A.Z. led the molecular analysis of kindred 5. J.C. performed and interpreted the kindred 5 ciliary electron microscopy and HSVM. M.L.D. identified the kindred 5 patients and provided clinical data. P.R.S. performed the HSVM. K.E.B. and L.P.H. assisted with the acquisition of proband 1 clinical samples. I.A. identified bronchiectasis kindreds 2 and 3. E.M.F. assisted with the analysis of the phosphoproteomics data. V.V. assisted with the IFM experiments, analyzed the data, and edited the manuscript. H.O. supervised and led the kindred 4 analyses. M.R.K. supervised and led the kindred 5 molecular analysis and clinical phenotyping. G.J.T. supervised the μOCT experiments and analyzed the data. F.S.A. supervised the whole exome sequencing and linkage analysis of kindreds 1–3, analyzed the genetics data, and edited the manuscript. D.M.S. supervised the project, designed the experiments, and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Massachusetts General Hospital, the Whitehead Institute for Biomedical Research, and the Massachusetts Institute of Technology are in the process of filing a provisional patent application covering the therapeutic augmentation of NEK10 signaling in disorders of mucociliary clearance (R.R.C. and D.M.S., inventors). All other authors have no competing interests.
Additional information
Peer review information Kate Gao was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Recurrent NEK10 mutations in familial bronchiectasis.
a, Pedigree indicating affected siblings (filled), proband (‘p’), and subjects from whom genomic DNA was available for analysis (asterisks). b, Chest CT of siblings ‘a’ and ‘b’ from panel (a), with arrows indicating regions of bronchiectatic lung. c, RefSeq-annotated NEK10 variants annotated with transcription start sites, transcript sizes, predicted protein molecular weights, and exon–exon junctions assayed by qRT–PCR in Fig. 1d. d, Immunoblotting against indicated NEK10 epitopes, representative of three experiments. HBEC bands are non-specific. Full-length 133 kDa NEK10 protein is indicated with a dashed box. e, Pedigree of kindred 2. Asterisks denote family members from whom genomic DNA was available, the dashed line indicates consanguinity by shared tribal ancestry, and the Sanger sequencing trace confirms c.1869dupT. f, Chest radiograph of proband 2, with arrow highlighting bronchiectasis. g, Pedigree of kindred 3. The dashed line indicates consanguinity by shared geographical ancestry, and the Sanger sequencing trace confirms c.2243C>T. h, CT from proband 3 demonstrating cystic (green arrow) and cylindrical (red arrow) bronchiectasis. i, Pedigree of kindred 4. The Sanger sequencing trace confirms c.1371+1G>T. j, CT from proband 4 indicating right middle lobe (red arrow) and left lower lobe (green arrow) bronchiectasis. k, Proband 4 nasal biopsy TEM demonstrating normal radial ciliary ultrastructure. Scale bar, 200 nm. l, Pedigree of kindred 5. The dashed line indicates consanguinity by shared tribal ancestry, and the Sanger sequencing trace confirms c.2317C>T. m, CTs of affected siblings in (l), demonstrating bronchiectasis. n–o, Nasal biopsy TEM of affected siblings in (l). Scale bars, 1 μm (n) and 200 nm (o).
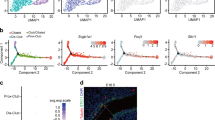
Extended Data Fig. 2 NEK10 loss does not detectably alter airway epithelial differentiation.
a, 18S rRNA-normalized relative NEK10 expression during ALI differentiation. n = 1 ALI culture per time point. b–d, 18S rRNA-normalized relative expression of ciliated cell markers FOXJ1 and DNAH5 (b), secretory cell marker SCGB1A1 (c), and basal cell marker KRT5 (d). n = 1 ALI culture per time point. e,f, Whole-mount immunofluorescence microscopy against SCGB1A1 (e, upper panel), goblet cell marker MUC5AC (e, lower panel), KRT5 (f, upper panel), and ciliated cell marker Ac-α-tubulin (f, lower panel). Scale bars, 100 μm. Bar graphs indicate the fraction of the surface epithelium occupied by marker-positive cells. n = 4 per condition, representative of 6 ALI differentiations. Mean ± s.d. g, Schematic depiction of bioinformatic NEK10 promoter (red) identification using the indicated UCSC genome browser hg19 tracks: CpG islands, H3K27-Ac, DNAse I hypersensitivity clusters, and transcription factor chromatin immunoprecipitation sequencing (ChIP-seq). h, Live GFP imaging of ALI cultures of the indicated genotypes and maturity, representative of three independent ALI differentiations. Scale bars, 200 μm. i, Gating strategy for FACS sorting of GFP-labeled ALI cultures. Numbers indicate the percentage of gated cells per population.
Extended Data Fig. 3 Functional consequences of NEK10 activity manipulation.
a, Quantification of analysis in Fig. 2c. Mean ± s.d. b, Kymographs of μOCT-based particle tracking from mature ALI cultures, representative of three independent ALI differentiations. c, CBF (μOCT) of mature ALI cultures of the indicated genotypes. n = 27 for NEK10WT and n = 22 for NEK10G>C, pooled from three independent ALI differentiations. Mean ± s.e.m. d, Immunoblotting of mature ALI lysates after CRISPR–Cas9-mediated gene editing with the indicated sgRNAs, representative of two experiments. Short (S) versus long (L) exposures are indicated. e, Quantification of analysis in Fig. 2f. Mean ± s.d. f, CBF of mature ALI cultures edited with the indicated sgRNAs, n = 8 per condition pooled from three independent ALI differentiations. Mean ± s.e.m. g, Immunoblotting of mature ALI lysates transduced with the indicated cDNAs, representative of 2 experiments. Short (S) versus long (L) exposures are indicated. h, Quantification of analysis in Fig. 2i. Mean ± s.d. i, Pseudocolored video microscopy of mature ALI cultures transduced with the indicated cDNAs. Representative fields from three independent ALI differentiations are shown. Scale bars, 50 μm. j, CBF of mature ALI cultures transduced with the indicated cDNAs. n = 4 per condition, pooled from three independent ALI differentiations. Mean ± s.e.m. *P ≤ 0.05, **P ≤ 0.01, ****P ≤ 0.0001.
Extended Data Fig. 4 Experimental manipulation of NEK10 activity alters ciliated cell morphology.
a, Gating strategy for IFC analysis of MCCs. b, Representative images and masking data of cells in (a), demonstrating the ability to generate single NEK10:eGFP+ ciliated cells for analysis. c, Confocal MIPs of mature ALI cultures edited with the indicated sgRNAs after IFM against Ac-α-tubulin, representative of two independent ALI differentiations. Scale bars, 25 μm. d, Confocal MIPs of mature ALI cultures transduced with the indicated cDNAs after IFM against Ac-α-tubulin, representative of two independent ALI differentiations. Scale bars, 25 μm. e, H&E stained mature ALI samples of the indicated genotypes after sectioning orthogonal to the epithelial surface, representative of three independent ALI differentiations.
Extended Data Fig. 5 Structural and proteomic abnormalities in NEK10-deficient airway epithelium.
a, SEMs of mature ALI cultures edited with the indicated sgRNAs, representative of two independent ALI differentiations. Scale bars, 100 μm (upper panels) and 1 μm (lower panels). b, Immunoblotting against the indicated proteins from lysates generated from purified cilia (lanes 2 and 4) or remaining de-ciliated mature ALI cultures (lanes 1 and 3), representative of two experiments. c, Cumulative distribution of phosphopeptides by log2[fold change] between indicated conditions. The solid (sgNEK10b) and dashed (sgNEK10c) red lines illustrate the population of depleted phosphopeptides upon NEK10 deletion. d, Table of GO classes enriched among genes (n = 395) whose peptides are depleted >1.5 fold log2[fold change] after targeting with sgNEK10b. The enrichment levels, P values, and false discovery rates are indicated. e, Cumulative distribution of phosphopeptides by log2[fold change]. Previously validated PCD proteins are in red and all other detected proteins are in black, as in Fig. 4e. f, Cumulative distribution of phosphopeptides by log2[fold change]. Previously validated non-PCD ciliopathy proteins are in red and all other detected proteins are in black, as in Fig. 4e.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2.
Supplementary Video 1
Impaired ciliary motility in NEK10-deficient airway epithelium. Mature ALI cultures of the indicated genotype imaged by ×40 phase contrast microscopy at 30 frames per s, documenting the motion amplitude in NEK10WT and NEK10G>C epithelia.
Supplementary Video 2
Impaired mucociliary transport in NEK10-deficient airway epithelium. Mature ALI cultures of the indicated genotype imaged by μOCT after the addition of apical polystyrene beads, documenting particle transport in NEK10WT and NEK10G>C epithelia.
Source data
Source Data Fig. 1
Unprocessed western blots
Source Data Fig. 1
Statistical source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 2
Statistical source data
Source Data Extended Data Fig. 3
Unprocessed western blots
Source Data Extended Data Fig. 3
Statistical source data
Source Data Extended Data Fig. 5
Unprocessed western blots
Source Data Extended Data Fig. 5
Statistical source data
Rights and permissions
About this article
Cite this article
Chivukula, R.R., Montoro, D.T., Leung, H.M. et al. A human ciliopathy reveals essential functions for NEK10 in airway mucociliary clearance. Nat Med 26, 244–251 (2020). https://doi.org/10.1038/s41591-019-0730-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0730-x
This article is cited by
-
Dual-color live imaging unveils stepwise organization of multiple basal body arrays by cytoskeletons
EMBO Reports (2024)
-
Dnah9 mutant mice and organoid models recapitulate the clinical features of patients with PCD and provide an excellent platform for drug screening
Cell Death & Disease (2022)
-
Imaging flow cytometry
Nature Reviews Methods Primers (2022)
-
Combining RSPH9 founder mutation screening and next-generation sequencing analysis is efficient for primary ciliary dyskinesia diagnosis in Saudi patients
Journal of Human Genetics (2022)
-
Diverse monogenic subforms of human spermatogenic failure
Nature Communications (2022)