Abstract
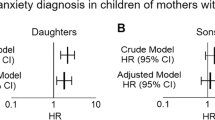
How obesity and elevated androgen levels in women with polycystic ovary syndrome (PCOS) affect their offspring is unclear. In a Swedish nationwide register-based cohort and a clinical case–control study from Chile, we found that daughters of mothers with PCOS were more likely to be diagnosed with PCOS. Furthermore, female mice (F0) with PCOS-like traits induced by late-gestation injection of dihydrotestosterone, with and without obesity, produced female F1–F3 offspring with PCOS-like reproductive and metabolic phenotypes. Sequencing of single metaphase II oocytes from F1–F3 offspring revealed common and unique altered gene expression across all generations. Notably, four genes were also differentially expressed in serum samples from daughters in the case–control study and unrelated women with PCOS. Our findings provide evidence of transgenerational effects in female offspring of mothers with PCOS and identify possible candidate genes for the prediction of a PCOS phenotype in future generations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw data of F1–F3 females and clinical characteristics of daughters are available through Dryad: https://doi.org/10.5061/dryad.jwstqjq4m. All raw and analyzed scRNA-seq data of mouse MII oocytes from F1–F3 females are available at the Gene Expression Omnibus database via accession number GSE133100.
For the Swedish register-based cohort, original data are held by the Swedish National Board of Health and Welfare and Statistics Sweden, and because of Swedish data privacy laws we cannot make the data publicly available. Any researcher can access the data by obtaining an ethical approval from a regional ethical review board and thereafter asking the Swedish National Board of Health and Welfare and Statistics Sweden for the original data. However, aggregated data used in the analysis of this study are available from the authors upon reasonable request and with approved data sharing and data processing agreements in line with the General Data Protection Regulation. Further use of these data must be authorized by the local ethics committee regarding the merit of the project involved. A detailed description of the unrelated case–control study, including global gene expression analyses in subcutaneous adipose tissue, has previously been published28. Source data for Figs. 1–4 and 6 and Extended Data Figs. 1–7 are presented with this paper.
References
March, W. A. et al. The prevalence of polycystic ovary syndrome in a community sample assessed under contrasting diagnostic criteria. Hum. Reprod. 25, 544–551 (2010).
Dumesic, D. A. et al. Scientific statement on the diagnostic criteria, epidemiology, pathophysiology, and molecular genetics of polycystic ovary syndrome. Endocr. Rev. 36, 487–525 (2015).
Azziz, R. PCOS in 2015: new insights into the genetics of polycystic ovary syndrome. Nat. Rev. Endocrinol. 12, 183 (2016).
Stener-Victorin, E. et al. Are there any sensitive and specific sex steroid markers for polycystic ovary syndrome? J. Clin. Endocrinol. Metab. 95, 810–819 (2010).
O’Reilly, M. W. et al. AKR1C3-mediated adipose androgen generation drives lipotoxicity in women with polycystic ovary syndrome. J. Clin. Endocrinol. Metab. 102, 3327–3339 (2017).
Luotola, K., Piltonen, T. T., Puurunen, J., Morin-Papunen, L. C. & Tapanainen, J. S. Testosterone is associated with insulin resistance index independently of adiposity in women with polycystic ovary syndrome. Gynecol. Endocrinol. 34, 40–44 (2018).
Barrett, E. S. et al. Anogenital distance in newborn daughters of women with polycystic ovary syndrome indicates fetal testosterone exposure. J. Dev. Orig. Health Dis. 9, 307–314 (2018).
Homburg, R., Gudi, A., Shah, A. & Layton, A, M. A novel method to demonstrate that pregnant women with polycystic ovary syndrome hyper-expose their fetus to androgens as a possible stepping stone for the developmental theory of PCOS. A pilot study. Reprod. Biol. Endocrinol. 15, 61 (2017).
Hanson, M. A. & Gluckman, P. D. Early developmental conditioning of later health and disease: physiology or pathophysiology? Physiol. Rev. 94, 1027–1076 (2014).
Roland, A. V., Nunemaker, C. S., Keller, S. R. & Moenter, S. M. Prenatal androgen exposure programs metabolic dysfunction in female mice. J. Endocrinol. 207, 213–223 (2010).
Hu, M. et al. Maternal testosterone exposure increases anxiety-like behavior and impacts the limbic system in the offspring. Proc. Natl Acad. Sci. USA 112, 14348–14353 (2015).
Guerrero-Bosagna, C. & Skinner, M. K. Environmentally induced epigenetic transgenerational inheritance of phenotype and disease. Mol. Cell Endocrinol. 354, 3–8 (2012).
Guerrero-Bosagna, C. et al. Epigenetic transgenerational inheritance of vinclozolin induced mouse adult onset disease and associated sperm epigenome biomarkers. Reprod. Toxicol. 34, 694–707 (2012).
Skinner, M. K. et al. Ancestral dichlorodiphenyltrichloroethane (DDT) exposure promotes epigenetic transgenerational inheritance of obesity. BMC Med. 11, 228 (2013).
Manikkam, M., Tracey, R., Guerrero-Bosagna, C. & Skinner, M. K. Dioxin (TCDD) induces epigenetic transgenerational inheritance of adult onset disease and sperm epimutations. PLoS One 7, e46249 (2012).
Fornes, R., et al. Mice exposed to maternal androgen excess and diet-induced obesity have altered phosphorylation of catechol-O-methyltransferase in the placenta and fetal liver. Int. J. Obes. https://doi.org/10.1038/s41366-018-0314-8 (2019).
Saben, J. L. et al. Maternal metabolic syndrome programs mitochondrial dysfunction via germline changes across three generations. Cell Rep. 16, 1–8 (2016).
Cesta, C. E., et al. Maternal polycystic ovary syndrome and risk of neuropsychiatric disorders in offspring: prenatal androgen exposure or genetic confounding? Psychol. Med. https://doi.org/10.1017/S0033291719000424 (2019).
Witham, E. A., Meadows, J. D., Shojaei, S., Kauffman, A. S. & Mellon, P. L. Prenatal exposure to low levels of androgen accelerates female puberty onset and reproductive senescence in mice. Endocrinology 153, 4522–4532 (2012).
Fornes, R. et al. The effect of androgen excess on maternal metabolism, placental function and fetal growth in obese dams. Sci. Rep. 7, 8066 (2017).
Chen, D. et al. The TFAP2C-regulated OCT4 naive enhancer is involved in human germline formation. Cell Rep.25, 3591–3602 (2018).
Kaiser, S. et al. Reduced gene dosage of Tfap2c impairs trophoblast lineage differentiation and alters maternal blood spaces in the mouse placenta. Biol. Reprod. 93, 1–13 (2015).
Sharma, N. et al. Tpbpa–Cre-mediated deletion of TFAP2C leads to deregulation of Cdkn1a, Akt1 and the ERK pathway, causing placental growth arrest. Development 143, 787–798 (2016).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Kim, D. A. & Suh, E. K. Defying DNA double-strand break-induced death during prophase I meiosis by temporal TAp63ɑ phosphorylation regulation in developing mouse oocytes. Mol. Cell Biol. 34, 1460–1473 (2014).
Ribeiro, J. R., Lovasco, L. A., Vanderhyden, B. C. & Freiman, R. N. Targeting TBP-associated factors in ovarian cancer. Front. Oncol. 4, 45 (2014).
Sasaki, D., Kotoh, J., Watadani, R. & Matsumoto, K. New animal models reveal that coenzyme Q2 (Coq2) and placenta-specific 8 (Plac8) are candidate genes for the onset of type 2 diabetes associated with obesity in rats. Mamm. Genome 26, 619–629 (2015).
Kokosar, M. et al. Epigenetic and transcriptional alterations in human adipose tissue of polycystic ovary syndrome. Sci. Rep. 6, 22883 (2016).
Zhang, Y. et al. Transcriptome landscape of human folliculogenesis reveals oocyte and granulosa cell interactions. Mol. Cell 72, 1021–1034 (2018).
Gudmundsdottir, B. et al. POGZ is required for silencing mouse embryonic β-like hemoglobin and human fetal hemoglobin expression. Cell Rep. 23, 3236–3248 (2018).
Weissenrieder, J. S., Reilly, J. E., Neighbors, J. D. & Hohl, R. J. Inhibiting geranylgeranyl diphosphate synthesis reduces nuclear androgen receptor signaling and neuroendocrine differentiation in prostate cancer cell models. Prostate 79, 21–30 (2019).
Li, C. et al. Transmembrane protein 214 (TMEM214) mediates endoplasmic reticulum stress-induced caspase 4 enzyme activation and apoptosis. J. Biol. Chem. 288, 17908–17917 (2013).
Tao, W. et al. Lipid-induced muscle insulin resistance is mediated by GGPPS via modulation of the RhoA/Rho kinase signaling pathway. J. Biol. Chem. 290, 20086–20097 (2015).
Gu, M. et al. RhoA phosphorylation mediated by Rho/RhoA-associated kinase pathway improves the anti-freezing potentiality of murine hatched and diapaused blastocysts. Sci. Rep. 7, 6705 (2017).
Beck, A. R., Miller, I. J., Anderson, P. & Streuli, M. RNA-binding protein TIAR is essential for primordial germ cell development. Proc. Natl Acad. Sci. USA 95, 2331–2336 (1998).
Jagannath, A. et al. The CRTC1–SIK1 pathway regulates entrainment of the circadian clock. Cell 154, 1100–1111 (2013).
Doherty, D. A., Newnham, J. P., Bower, C. & Hart, R. Implications of polycystic ovary syndrome for pregnancy and for the health of offspring. Obstet. Gynecol. 125, 1397–1406 (2015).
Roos, N. et al. Risk of adverse pregnancy outcomes in women with polycystic ovary syndrome: population based cohort study. BMJ 343, d6309 (2011).
Crisosto, N. et al. Higher luteinizing hormone levels associated with antimullerian hormone in postmenarchal daughters of women with polycystic ovary syndrome. Fertil. Steril. 111, 381–388 (2018).
Torchen, L. C., Legro, R. S. & Dunaif, A. Distinctive reproductive phenotypes in peripubertal girls at risk for polycystic ovary syndrome. J. Clin. Endocrinol. Metab. 104, 3355–3361 (2019).
Palomba, S. et al. Pervasive developmental disorders in children of hyperandrogenic women with polycystic ovary syndrome: a longitudinal case–control study. Clin. Endocrinol. 77, 898–904 (2012).
Manneras-Holm, L. et al. Adipose tissue has aberrant morphology and function in PCOS: enlarged adipocytes and low serum adiponectin, but not circulating sex steroids, are strongly associated with insulin resistance. J. Clin. Endocrinol. Metab. 96, E304–E311 (2011).
Dumesic, D. A., et al. Adipose insulin resistance in normal-weight polycystic ovary syndrome women. J. Clin. Endocrinol. Metab. 104, 2171–2183 (2019).
Petta, S. et al. Insulin resistance and hyperandrogenism drive steatosis and fibrosis risk in young females with PCOS. PLoS One 12, e0186136 (2017).
Aiken, C. E. & Ozanne, S. E. Transgenerational developmental programming. Hum. Reprod. Update 20, 63–75 (2014).
Crisosto, N. et al. Anti-Mullerian hormone levels in peripubertal daughters of women with polycystic ovary syndrome. J. Clin. Endocrinol. Metab. 92, 2739–2743 (2007).
Sir-Petermann, T. et al. Metabolic and reproductive features before and during puberty in daughters of women with polycystic ovary syndrome. J. Clin. Endocrinol. Metab. 94, 1923–1930 (2009).
Crisosto, N. et al. Improvement of hyperandrogenism and hyperinsulinemia during pregnancy in women with polycystic ovary syndrome: possible effect in the ovarian follicular mass of their daughters. Fertil. Steril. 97, 218–224 (2012).
Palomba, S. et al. Macroscopic and microscopic findings of the placenta in women with polycystic ovary syndrome. Hum. Reprod. 28, 2838–2847 (2013).
Maliqueo, M. et al. Placental STAT3 signaling is activated in women with polycystic ovary syndrome. Hum. Reprod. 30, 692–700 (2015).
Cavalli, G. & Heard, E. Advances in epigenetics link genetics to the environment and disease. Nature 571, 489–499 (2019).
Xavier, M. J., Roman, S. D., Aitken, R. J. & Nixon, B. Transgenerational inheritance: how impacts to the epigenetic and genetic information of parents affect offspring health. Hum. Reprod. Update 25, 518–540 (2019).
Hentze, M. W., Castello, A., Schwarzl, T. & Preiss, T. A brave new world of RNA-binding proteins. Nat. Rev. Mol. Cell Biol.19, 327–341 (2018).
Meyer, C. et al. The TIA1 RNA-binding protein family regulates EIF2AK2-mediated stress response and cell cycle progression. Mol. Cell 69, 622–635 (2018).
Dapas, M., et al. Family-based quantitative trait meta-analysis implicates rare noncoding variants in DENND1A in polycystic ovary syndrome. J. Clin. Endocrinol. Metab. https://doi.org/10.1210/jc.2018-02496 (2019).
Tee, M. K. et al. Alternative splicing of DENND1A, a PCOS candidate gene, generates variant 2. Mol. Cell. Endocrinol. 434, 25–35 (2016).
McAllister, J. M. et al. Overexpression of a DENND1A isoform produces a polycystic ovary syndrome theca phenotype. Proc. Natl Acad. Sci. USA 111, E1519–E1527 (2014).
Wang, F. et al. Alternative splicing of the androgen receptor in polycystic ovary syndrome. Proc. Natl Acad. Sci. USA 112, 4743–4748 (2015).
Furuhashi, M. et al. Serum FABP5 concentration is a potential biomarker for residual risk of atherosclerosis in relation to cholesterol efflux from macrophages. Sci. Rep. 7, 217 (2017).
Miyamoto, K., Uechi, A. & Saito, K. The zinc finger domain of RING finger protein 141 reveals a unique RING fold. Protein Sci. 26, 1681–1686 (2017).
Swanson, M., Sauerbrei, E. E. & Cooperberg, P. L. Medical implications of ultrasonically detected polycystic ovaries. J. Clin. Ultrasound 9, 219–222 (1981).
Manti, M., et al. Maternal androgen excess induces cardiac hypertrophy and left ventricular dysfunction in female mice offspring. Cardiovasc. Res. https://doi.org/10.1093/cvr/cvz180 (2019).
Manti, M. et al. Maternal androgen excess and obesity induce sexually dimorphic anxiety-like behavior in the offspring. FASEB J. 32, 4158–4171 (2018).
Benrick, A. et al. Adiponectin protects against development of metabolic disturbances in a PCOS mouse model. Proc. Natl Acad. Sci. USA 114, E7187–E7196 (2017).
Nilsson, M. E. et al. Measurement of a comprehensive sex steroid profile in rodent serum by high-sensitive gas chromatography–tandem mass spectrometry. Endocrinology 156, 2492–2502 (2015).
Mehlem, A., Hagberg, C. E., Muhl, L., Eriksson, U. & Falkevall, A. Imaging of neutral lipids by Oil Red O for analyzing the metabolic status in health and disease. Nat. Protoc. 8, 1149–1154 (2013).
Picelli, S. E. A. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Love, M. I. et al. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Yu, G. et al. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Acknowledgements
We thank S. Pilström, J. Victorin and S. Edström for technical assistance during animal work; TSE Systems and the Metabolic Phenotyping Centre at the Strategic Research Program in Diabetes at the Karolinska Institutet; and the electron microscopy unit EMil at Huddinge University Hospital at the Karolinska Institutet. This work was funded by the Swedish Medical Research Council (project no. 2014-2775, 2018-02435 ESV; 2014-2870 QD; 2018-02119 MR), the Novo Nordisk Foundation (NNF17OC0026724, NNF18OC0033992 and NNF19OC0056647 ESV) and the Strategic Research Program in Diabetes at the Karolinska Institutet to E.S.-V.; the Adlerbertska Research Foundation to E.S.-V.; Karolinska Institutet KID funding to E.S.-V. and Q.D.; the Swedish Association of Medical Research and the Åke Wiberg Foundation to Q.D.; the Regional Agreement on Medical Training and Clinical Research between the Stockholm County Council and the Karolinska Institutet to E.S.-V.; the Royal Swedish Academy of Sciences (KVA, BS2015-0012) to S.R.; the National Fund for Scientific and Technological Development (FONDECYT) project no. 1071007 and 1151531 to T.S.-P.; and FONDECYT project no. 1181798 to M.M. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
S.R. designed the study, performed the transgenerational mouse data collection, performed molecular analyses, analyzed the data, prepared the figures and assisted in writing the manuscript. Y.P. performed and analyzed the scRNA-seq data, validated the sequencing data, performed the placenta and embryonic data analyses, prepared the figures and was involved in manuscript preparation. H.L. and H.-P.P. designed and performed the second mouse experiments with placental and embryonic data collection and analyses. M.M., R.F., Z.Z. and J.M. performed the transgenerational phenotyping of mice and analyzed the data. C.O. performed the serum sex steroid analyses with gas chromatography–mass spectrometry. E.L. performed the mouse and human molecular data analyses. N.C., M.M., B.E., A.L.d.G. and T.S.-P. provided clinical information for the case–control study, provided biological serum samples and assisted in manuscript preparation. H.L., M.A.R. and C.E.C. provided data from the Swedish nationwide register-based cohort study, analyzed these data and assisted in manuscript preparation. A.B. was involved in the study design, interpretation of the results and preparation of the manuscript. E.S.-V. and Q.D. designed the study, analyzed the data, prepared the figures and wrote the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
H.L. has served as a speaker for Evolan Pharma and Shire and has received research grants from Shire, all outside the scope of the submitted work. All other authors have no conflicts of interest to declare.
Additional information
Peer review information Jennifer Sargent was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Effect of diet-induced obesity prior mating.
a-c, Body weight development (n = 80 per group) and food and calorie intake per day of F0-dams during the 6-weeks on the diet prior to mating. Food and calorie intake were calculated as food consumption/mouse/cage/day. d-e, Body composition (fat and lean mass) normalized to body weight (gram) (n = 40 per group). f, Blood glucose levels at different time point during the OGTT and (g) glucose area under the curve (AUC) at 0 to 90 minutes in F0-dams (n = 25 per group). h, Serum insulin levels at 0 and 15 minutes before and after the oral glucose challenge (For 0/15 minutes, F0-CD = 23/15, F0-HFHS = 13/17). Comparisons between groups was performed by two-tailed unpaired Student t-test. All data are presented as mean ± s.e.m. Number of mice are stated in bars of each group. Veh = vehicle; DHT = dihydrotestosterone; CD = control diet; HFHS = high-fat high-sucrose.
Extended Data Fig. 2 Diet-induced maternal obesity and prenatal androgen exposure effects on litter size.
a, Pups per litter size in F1-F3 adult offspring. For litter size, there was a main effect in the androgenized + obese lineage in F1 female offspring (two-way ANOVA, Tukey’s post hoc analysis), and in the obese [F 1, 117 = 6.688, P = 0.01], and in the androgenized lineage, [F 1, 117 = 15.51, P = 0.0001]. In F2 offspring, there was a main effect in the obese lineage (two-way ANOVA, Tukey’s post hoc analysis) [F 1, 32 = 10.12, P = 0.0033]. The number of mice/group are specified in Supplementary Table 7. b, Representative images showing absorbed embryos (yellow arrows) at E12.5 in the control and obese + androgenized lineage. c, mRNA expression of germ cell markers developmental pluripotency associated 3 (Dppa3) and mouse vasa homologue (Mvh) in the gonads of F2 fetuses at E12.5 in the control (n = 14/8), androgenized (n = 3/4), obese (n = 7/5), and obese + androgenized (n = 4/3) lineages, and (two-way ANOVA, Tukey’s post hoc analysis) d, mRNA expression of germ cell marker Dppa3 and Mvh in the gonads of F2 fetus at E18.5 in the control (n = 9/10), androgenized (n = 8/10), and obese (n = 8/10) lineages (one-way ANOVA, Dunnett’s post hoc analysis). All data are presented as mean ± s.e.m. Veh = vehicle; DHT = dihydrotestosterone; CD = control diet; HFHS = high-fat high-sucrose; Mat = maternal; GMat = grand-maternal; GGMat = great-grand maternal.
Extended Data Fig. 3 Diet-induced maternal obesity and prenatal androgen exposure effects on offspring estrus cycle changes.
a, Quantitative analysis of estrous cyclicity in F1 to F3 adult female offspring. (F1: MatCD + Veh, n = 18, MatCD + DHT, n = 15, MatHFHS + Veh, n = 14, MatHFHS + DHT, n = 12. F2: GMatCD + Veh, n = 13, GMatCD + DHT, n = 11, GMatHFHS + Veh, n = 16. F3: GGMatCD + Veh, n = 11, GGMatCD + DHT, n = 8, GGMatHFHS + Veh, n = 9). M/D, metestrus and diestrus; E, estrus; P, proestrus. For proestrus, there was a main effect in the obese lineage of F1 female offspring (two-way ANOVA, Tukey’s post hoc analysis) [F 1, 54 = 16.31, P < 0.001]. For estrus, there was a main effect in the androgenized lineage of F1 female offspring (two-way ANOVA, Tukey’s post hoc analysis) [F 1, 54 = 40.396, P < 0.001]. For metestrus/diestrus, there was a main effect in the androgenized lineage of F1 female offspring (two-way ANOVA, Tukey’s post hoc analysis) [F 1, 54 = 28.953, P < 0.001] and in the obese lineage [F 1, 54 = 6.808, P < 0.01]. There were disrupted estrus cycles in the androgenized lineage of F2 offspring (one-way ANOVA, Dunnett’s post hoc analysis) [F2, 49 = 14.19; P < 0.0001]. b, Representative estrous cyclicity assessment in 12-week old female mice for 10 consecutive days by vaginal cytology in F1-F3 adult offspring. The data are present in violin plot showing the frequency distribution curves. The median and quartiles values are shown in dotted and dashed lines respectively. Veh = vehicle; DHT = dihydrotestosterone; CD = control diet; HFHS = high-fat high-sucrose; Mat = maternal; GMat = grand-maternal; GGMat = great-grand maternal.
Extended Data Fig. 4 Diet-induced maternal obesity and prenatal androgen exposure effects on circulating sex steroids in F1-F3 adult female offspring.
Serum (a) testosterone, (b) androstenedione, and (c) Dihydrotestosterone levels measured by gas-chromatography mass spectrometry (GC-MS/MS) in F1, F2 and F3 adult offspring. d, Serum anti-Müllerian hormone (AMH) levels measured by ELISA in F1, F2, and F3 adult offspring (n = number of animals) Comparisons between groups in F1 was performed using two-way ANOVA followed by Tukey’s post hoc analysis and by using one-way ANOVA followed by Dunnett’s post hoc analysis in F2 and F3 generations. All data are presented as mean ± s.e.m. Number of mice are stated in bars of each group. Veh = vehicle; DHT = dihydrotestosterone; CD = control diet; HFHS = high-fat high-sucrose; Mat = maternal; GMat = grand-maternal; GGMat = great-grand maternal.
Extended Data Fig. 5 Diet-induced maternal obesity and prenatal androgen exposure effects on body weight and adipose tissue and liver gene expression.
a-b, Body weight development and area under the curve (AUC) of body weight in F1, F2 and F3 adult female offspring from 3 to15 weeks of age. c, Percent lean mass normalized with body weight (grams) in F1 to F3 adult female offspring at 18-week of age. d, Subcutaneous adipose tissue mRNA expression (2-ΔΔCT) of genes involved in adipogenesis; Bmp4, Zfp423, Cebpa, Cebpb, Ppard, and Pparg of F1-F3 female offspring. (n = animals per group. F1: MatCD + Veh, n = 6, MatCD + DHT, n = 5, MatHFHS + Veh, n = 5, MatHFHS + DHT, n = 5. F2: GMatCD + Veh, n = 6, GMatCD + DHT, n = 5, GMatHFHS + Veh, n = 5, GMatHFHS + DHT, n = 1. F3: GGMatCD + Veh, n = 4, GGMatCD + DHT, n = 3, GGMatHFHS + Veh, n = 5). e, Gene expression (2-ΔΔCT) of lipid biosynthesis and free fatty acid oxidation pathways; Acaca, Fasn, Scd1, Pparg, Fitm1, Apoa1, Lxra, and Srebf1 in liver of F1-F3 female offspring. (n = animals per group. F1: MatCD + Veh, n = 9, MatCD + DHT, n = 10, MatHFHS + Veh, n = 11, MatHFHS + DHT, n = 8. F2: GMatCD + Veh, n = 10, GMatCD + DHT, n = 7, GMatHFHS + Veh, n = 10, GMatHFHS + DHT, n = 1. F3: GGMatCD + Veh, n = 9, GGMatCD + DHT, n = 8, GGMatHFHS + Veh, n = 8). F1: two-way ANOVA, Tukey’s post hoc analysis; F2 and F3: one-way ANOVA, Dunnett’s post hoc analysis. All data are presented as mean ± s.e.m. Number of mice are stated in bars of each group or in the text. Veh = vehicle; DHT = dihydrotestosterone; CD = control diet; HFHS = high-fat high-sucrose; Mat = maternal; GMat = grand-maternal; GGMat = great-grand maternal.
Extended Data Fig. 6 Diet-induced maternal obesity and prenatal androgen exposure effects on glucose homeostasis.
a-b, Blood glucose levels at different time point during oral glucose tolerance test (OGTT) and glucose area under the curve (AUC) at 0 to 90 minutes in F1 to F3 adult female offspring. (F1: MatCD + Veh, n = 16, MatCD + DHT, n = 15, MatHFHS + Veh, n = 11, MatHFHS + DHT, n = 12. F2: GMatCD + Veh, n = 13, GMatCD + DHT, n = 9, GMatHFHS + Veh, n = 12, GMatHFHS + DHT, n = 1. F3: GGMatCD + Veh, n = 11, GGMatCD + DHT, n = 8, GGMatHFHS + Veh, n = 9). (n = number of animals per group). c, Serum insulin levels at 0 and 15 minutes before and after oral glucose challenge. (F1: MatCD + Veh, n = 16, MatCD + DHT, n = 15, MatHFHS + Veh, n = 13, MatHFHS + DHT, n = 12. F2: GMatCD + Veh, n = 13, GMatCD + DHT, n = 10, GMatHFHS + Veh, n = 12, GMatHFHS + DHT, n = 1. F3: GGMatCD + Veh, n = 11, GGMatCD + DHT, n = 8, GGMatHFHS + Veh, n = 8). F1: two-way ANOVA, Tukey’s post hoc analysis; F2 and F3: one-way ANOVA, Dunnett’s post hoc analysis. All data are presented as mean ± s.e.m. Number of mice are stated in bars of each group. Veh = vehicle; DHT = dihydrotestosterone; CD = control diet; HFHS = high-fat high-sucrose; Mat = maternal; GMat = grand-maternal; GGMat = great-grand maternal.
Extended Data Fig. 7 Diet-induced maternal obesity and prenatal androgen exposure alters mitochondrial morphology in MII oocytes of F1, F2 and F3.
a, e, and i Representative transmission electron microscopy (TEM) images of F1–F3 MII oocytes (n = 5 MII oocytes per mice per group from three adult offspring). Blue arrows denote abnormal mitochondria shape and vacuoles. Yellow arrow denotes normal mitochondrial shape. b, f, and j, Average number of mitochondria per MII oocytes (n = number of animals per group). For average mitochondrial number, there was a main effect in the obese lineage in F1 female offspring (two-way ANOVA) [F 1, 210 = 12.10, P = 0.0006] and in in F3 offspring (one-way ANOVA) [F2, 115 = 3.746; P = 0.026]. c, g, and k, Mitochondrial DNA content per MII oocytes, expressed in 2-ΔCT value (n = number of animals per group). For mitochondrial DNA content, there was a main effect in the androgenized lineage in F1 female offspring (two-way ANOVA) [F 1, 52 = 8.335, P = 0.0057]. d, h, and l, Quantitative analysis of lipid droplets in MII oocytes in the four different groups (n = number of animals per group). For average lipid droplets, there was a main effect in the androgenized lineage in F1 female offspring (two-way ANOVA) [F 1, 16 = 19.98, P = 0.0004]. Comparisons between groups F1 and 2 generations were performed using two-way ANOVA followed by Tukey’s post hoc analysis and F3 generation one-way ANOVA followed by Dunnett’s post hoc analysis. Data are presented as mean ± s.e.m. Veh = vehicle; DHT = dihydrotestosterone; CD = control diet; HFHS = high-fat high-sucrose; Mat = maternal; GMat = grand-maternal; GGMat = great-grand maternal.
Extended Data Fig. 8 Single cell RNA sequencing of MII oocytes.
a, Principal component analysis (PCA) plot of all MII oocytes clustered according to generation indicating batch effects due to technical variability. b, PCA plot of MII oocytes of F1-F3 generations in the control, androgenized, and obese lineages after batch effect correction (c) PCA plot of MII oocytes of F1-F3 generations in control, androgenized, and obese lineages based on 641 differentially expressed genes (DEGs). Generation: F1 = first generation; F2 = second generation and F3 = third generation. Condition: CD + Veh = control lineage; CD + DHT = androgenized lineage: HFHS + Veh = obese lineage. F1 = first generation; F2 = second generation and F3 = third generation. Veh = vehicle; DHT = dihydrotestosterone; CD = control diet; HFHS = high-fat high-sucrose.
Extended Data Fig. 9 Venn diagram of orthologues genes.
Venn diagram demonstrating overlap of orthologous genes in human and mouse MII oocytes, respectively. The human data are from previous published reports: GSE107746.
Supplementary information
Supplementary Information
Supplementary Tables 1–8
Supplementary Dataset 1
Source data for Supplementary Table 1
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 3b
Imaging Source Data
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 6
Statistical Source Data,
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 4
Statistical Source Data
Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 7b,d,f,h,j,l
Unprocessed Western Blots
Rights and permissions
About this article
Cite this article
Risal, S., Pei, Y., Lu, H. et al. Prenatal androgen exposure and transgenerational susceptibility to polycystic ovary syndrome. Nat Med 25, 1894–1904 (2019). https://doi.org/10.1038/s41591-019-0666-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0666-1
This article is cited by
-
Association between single nucleotide polymorphisms, TGF-β1 promoter methylation, and polycystic ovary syndrome
BMC Pregnancy and Childbirth (2024)
-
Elevated circulating adiponectin levels do not prevent anxiety-like behavior in a PCOS-like mouse model
Scientific Reports (2024)
-
Polycystic ovary syndrome
Nature Reviews Disease Primers (2024)
-
Appropriate glycemic management protects the germline but not the uterine environment in hyperglycemia
EMBO Reports (2024)
-
Gestational testosterone excess early to mid-pregnancy disrupts maternal lipid homeostasis and activates biosynthesis of phosphoinositides and phosphatidylethanolamines in sheep
Scientific Reports (2024)