Abstract
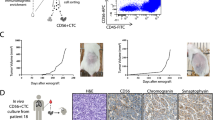
Approximately 50% of patients with early-stage non-small-cell lung cancer (NSCLC) who undergo surgery with curative intent will relapse within 5 years1,2. Detection of circulating tumor cells (CTCs) at the time of surgery may represent a tool to identify patients at higher risk of recurrence for whom more frequent monitoring is advised. Here we asked whether CellSearch-detected pulmonary venous CTCs (PV-CTCs) at surgical resection of early-stage NSCLC represent subclones responsible for subsequent disease relapse. PV-CTCs were detected in 48% of 100 patients enrolled into the TRACERx study3, were associated with lung-cancer-specific relapse and remained an independent predictor of relapse in multivariate analysis adjusted for tumor stage. In a case study, genomic profiling of single PV-CTCs collected at surgery revealed higher mutation overlap with metastasis detected 10 months later (91%) than with the primary tumor (79%), suggesting that early-disseminating PV-CTCs were responsible for disease relapse. Together, PV-CTC enumeration and genomic profiling highlight the potential of PV-CTCs as early predictors of NSCLC recurrence after surgery. However, the limited sensitivity of PV-CTCs in predicting relapse suggests that further studies using a larger, independent cohort are warranted to confirm and better define the potential clinical utility of PV-CTCs in early-stage NSCLC.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Most data generated or analyzed during this study are included in this published article. The sequencing data are available through the Cancer Research UK & University College London Cancer Trials Centre for non-commercial research purposes. Access will be granted upon review of a project proposal that will be evaluated by a TRACERx data access committee, and an appropriate data access agreement will be entered into, subject to any applicable ethical approvals.
Change history
03 June 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Uramoto, H. & Tanaka, F. Recurrence after surgery in patients with NSCLC. Transl. Lung Cancer Res. 3, 242–249 (2014).
Taylor, M. D. et al. Tumor recurrence after complete resection for non-small cell lung cancer. Ann. Thorac. Surg. 93, 1813–1820 (2012).
Jamal-Hanjani, M. et al. Tracking the evolution of non-small-cell lung cancer. N Engl J Med. 376, 2109–2121 (2017).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer Statistics, 2017. CA Cancer J. Clin. 67, 7–30 (2017).
Aceto, N., Toner, M., Maheswaran, S. & Haber, D. A. En route to metastasis: circulating tumor cell clusters and epithelial-to-mesenchymal transition. Trends Cancer 1, 44–52 (2015).
Hodgkinson, C. et al. Circulating tumour cells from small cell lung cancer patients are tumourigenic. Lung Cancer 87, S1 (2015).
Morrow, C. J. et al. Tumourigenic non-small-cell lung cancer mesenchymal circulating tumour cells: a clinical case study. Ann. Oncol. 27, 1155–1160 (2016).
Baccelli, I. et al. Identification of a population of blood circulating tumor cells from breast cancer patients that initiates metastasis in a xenograft assay. Nat. Biotechnol. 31, 539–544 (2013).
Girotti, M. R. et al. Application of sequencing, liquid biopsies, and patient-derived xenografts for personalized medicine in melanoma. Cancer Discov. 6, 286–299 (2016).
Mohan, S., Chemi, F. & Brady, G. Challenges and unanswered questions for the next decade of circulating tumour cell research in lung cancer. Transl. Lung Cancer Res. 6, 454–472 (2017).
Crosbie, P. A. et al. Circulating tumor cells detected in the tumor-draining pulmonary vein are associated with disease recurrence after surgical resection of NSCLC. J. Thorac. Oncol. 11, 1793–1797 (2016).
Jamal-Hanjani, M. et al. Tracking genomic cancer evolution for precision medicine: the lung TRACERx study. PLoS Biol. 12, e1001906 (2014).
Abbosh, C. et al. Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution. Nature 545, 446 (2017).
Romero-Palacios, P. J. et al. Liquid biopsy beyond of cancer: circulating pulmonary cells as biomarkers of COPD aggressivity. Crit. Rev. Oncol. Hematol. 136, 31–36 (2019).
Deleye, L., Vander Plaetsen, A. S., Weymaere, J., Deforce, D. & Van Nieuwerburgh, F. Short tandem repeat analysis after whole genome amplification of single B-lymphoblastoid cells. Sci. Rep. 8, 1255 (2018).
Lin, C. W. et al. MicroRNA-135b promotes lung cancer metastasis by regulating multiple targets in the Hippo pathway and LZTS1. Nat. Commun. 4, 1877 (2013).
Heitzer, E. et al. Complex tumor genomes inferred from single circulating tumor cells by array-CGH and next-generation sequencing. Cancer Res. 73, 2965 (2013).
Lohr, J. G. et al. Whole-exome sequencing of circulating tumor cells provides a window into metastatic prostate cancer. Nat. Biotechnol. 32, 479–484 (2014).
Gill, R. & Schumacher, M. A simple test of the proportional hazards assumption. Biometrika 74, 289–300 (1987).
Therneau, T. M., Grambsch, P. M. & Fleming, T. R. Martingale-based residuals for survival models. Biometrika 77, 147–160 (1990).
Altman, D. G., McShane, L. M., Sauerbrei, W. & Taube, S. E. Reporting Recommendations for Tumor Marker Prognostic Studies (REMARK): explanation and elaboration. PLoS Med. 9, e1001216 (2012).
R Code Team. R: A Language and Environment for Statistical Computing. (R Foundation for Statistical Computing, 2013).
Therneau, T. A package for survival analysis in S v 2.38 (2015).
Kassambara, A., Kosinski, M. & Biecek, P. survminer: Drawing Survival Curves using’ggplot2’. R package version 0.3 1 (2017).
Heagerty, P. J. & Zheng, Y. Survival model predictive accuracy and ROC curves. Biometrics 61, 92–105 (2005).
Mesquita, B. et al. Molecular analysis of single circulating tumour cells following long-term storage of clinical samples. Mol. Oncol. 11, 1687–1697 (2017).
Rothwell, D. G. et al. Genetic profiling of tumours using both circulating free DNA and circulating tumour cells isolated from the same preserved whole blood sample. Mol. Oncol. 10, 566–574 (2016).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Andrews, S. FastQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2010).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol. 31, 213–219 (2013).
Ha, G. et al. Integrative analysis of genome-wide loss of heterozygosity and monoallelic expression at nucleotide resolution reveals disrupted pathways in triple-negative breast cancer. Genome Res. 22, 1995–2007 (2012).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Tamborero, D. et al. Cancer Genome Interpreter annotates the biological and clinical relevance of tumor alterations. Genome Med. 10, 25 (2018).
Acknowledgements
We sincerely thank the patients and their families for donating blood samples for research. We thank E. Aidaros-Talbot for administrative assistance with the manuscript. We also thank J. Shaw for kindly providing plasma (relapse time point) of patient CRUK0242. TRACERx is funded by Cancer Research UK (grant C11496/A17786). This research was supported by Cancer Research UK–Core funding to the CRUK Manchester Institute (C5759/A27412) Centre, funding to the CRUK Manchester Centre (C5759/A25254) and funding of the CRUK Lung Cancer Centre of Excellence. Support was also received from the Manchester Experimental Cancer Medicine Centre and Manchester NIHR Biomedical Research Centre. F.C. is funded by the CANCER-ID Consortium (115749-Cancer-ID). B.M. is funded by Menarini Biomarkers Singapore PTE Ltd. C.S.K. is funded by The Manchester MRC Single Cell Research Centre (MR/M008908/1). C.S. is Royal Society Napier Research Professor. This work was supported by the Francis Crick Institute, which receives its core funding from Cancer Research UK (FC001169, FC001202), the UK Medical Research Council (FC001169, FC001202) and the Wellcome Trust (FC001169, FC001202). C.S. is funded by Cancer Research UK (TRACERx and CRUK Cancer Immunotherapy Catalyst Network), the CRUK Lung Cancer Centre of Excellence, Stand Up 2 Cancer (SU2C), the Rosetrees Trust, the Butterfield and Stoneygate Trusts, NovoNordisk Foundation (ID16584), the Prostate Cancer Foundation and the Breast Cancer Research Foundation (BCRF). The research leading to these results has received funding from the European Research Council (ERC) under the European Union’s Seventh Framework Programme (FP7/2007-2013) Consolidator Grant (FP7-THESEUS-617844), European Commission ITN (FP7-PloidyNet 607722), an ERC Advanced Grant (PROTEUS) from the European Research Council under the European Union’s Horizon 2020 research and innovation programme (grant agreement 835297). Support was also provided to C.S. by the National Institute for Health Research, the University College London Hospitals Biomedical Research Centre and the Cancer Research UK University College London Experimental Cancer Medicine Centre.
Author information
Authors and Affiliations
Consortia
Contributions
C.S., C.D., P.C., D.G.R. and G.B. developed the clinical study, directed research and co-wrote the manuscript. F.C. designed and conducted experiments, analyzed data and drafted the manuscript with the assistance of D.G.R. and N.M.G. S.G., S.P.P., G.W., N.B., N.M.G., C.S.K., S.F., C.M. and M.D. provided bioinformatic support for the study. C.A. provided support for the clinical interpretation of the data. C.Z. performed statistical analysis. C.A. and D.M. performed central pathology review. D.B., D.S.T. and B.M. provided support for single-cell isolation. M.J.-H., J.P., F.G., R.S., M.A.B., C.H., S.V., Y.S., P.C., S.W., D.B., J.T., F.B. and A.H. supported patient recruitment and sample management and provided clinical support for the study.
Corresponding authors
Ethics declarations
Competing interests
C.D. receives research grants/support from AstraZeneca, Astex Pharmaceuticals, Bioven, Amgen, Carrick Therapeutics, Merck, Taiho Oncology, GSK, Bayer, Boehringer Ingelheim, Roche, BMS, Novartis, Celgene and Epigene Therapeutics, Menarini, Angle PLC and Clearbridge Biomedics, all of which are outside the scope of this paper. C.D. has received honoraria/consultancy fees from Biocartis, Merck and AstraZeneca and Illumina, again outside the scope of this work. C.S. receives grant support from Pfizer, AstraZeneca, BMS and Roche-Ventana, Boehringer-Ingelheim. C.S. has consulted for Pfizer, Novartis, GlaxoSmithKline, MSD, BMS, Celgene, AstraZeneca, Illumina, Genentech, Roche-Ventana, GRAIL, Medicxi and the Sarah Cannon Research Institute and is an adviser for Dynamo Therapeutics. C.S. is a shareholder of Apogen Biotechnologies, Epic Bioscience and GRAIL, and has stock options in and is cofounder of Achilles Therapeutics.
Additional information
Peer review information Joao Monteiro was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 PV-CTCs are associated with lung-cancer-specific relapse.
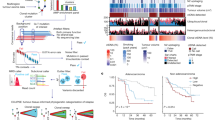
a, tdROC curves showing true-positive and false-positive rates for the 65th, 75th and 85th PV-CTC quantiles (≥3, ≥7, and ≥39 PV-CTCs per 7.5 ml blood, respectively) alongside the previously published threshold from our pilot study (≥18 PV-CTCs per 7.5 ml blood)11. All predictions were made at 720 d. Sensitivity and specificity of each category are shown along with area under ROC value. b, Kaplan–Meier curve showing lung-cancer-specific relapse-free survival for 98 patients stratified as PV-CTC high or low according to the 75th quantile (≥7 PV-CTCs per 7.5 ml blood). The number of patients at risk for each time point is indicated below the time point and color coded according to the high or low group. P values, HRs and relative 95% CIs (two-sided log-rank test) are indicated. c, Forest plot showing the results of multivariate regression analysis for patients with PV-CTC high or low status (≥7 PV-CTCs per 7.5 ml blood). The x-axis represents the HR with the reference line (dashed), and significance was calculated using a Cox proportional hazards model. The estimated HRs and their 95% CIs are presented as error bars. The log-rank test used was two sided.
Extended Data Fig. 2 Technical details of single-cell analysis of patient CRUK0242.
a, Consort diagram describing samples used for downstream analysis. Only patients with ≥5 PV-CTCs (n = 29) were processed through single-cell isolation (DEPArray). Single cells were not isolated from 6 of the 29 samples owing to failures during sample loading into the DEPArray machine. From the remaining 23 samples, 7 patients whose single CTCs isolated did not meet morphology criteria (Methods) were excluded. Sixteen samples were processed for WGA, and two patients whose CTCs did not show good quality GII in quality control after WGA were removed (Methods). b, Table showing cases of relapse in the patients with single PV-CTCs isolated. c, Agarose gel showing results of a quality-control PCR assay used to determine the genome integrity of each sample (n = 1). 0–4 bands determine the overall DNA integrity of each sample. DEPArray images of corresponding PV-CTC (green, cytokeratin; blue, CD45; purple, DAPI) are shown above. d, Examples of copy number profiles detected in single PV-CTCs, CECs, and WBC controls. Blue and red indicate regions of copy number loss and gain, respectively.
Extended Data Fig. 3 Genetic relationship between PV-CTCs, primary tumor and metastatic disease.
a, Venn diagram showing the overlap of somatic mutations detected among single PV-CTCs, primary tumor and metastatic tumor. b, Venn diagram showing the overlap of somatic mutations detected among single PV-CTCs, metastatic tumor and cfDNA isolated at the time of relapse.
Extended Data Fig. 4 Summary of all mutations detected in patient CRUK0242.
Heat map showing the comparison of SNVs detected in primary tumor regions, metastasis, PV-CTCs, CECs, WBCs and cfDNA samples (cfDNA before surgery was isolated from peripheral blood; cfDNA surgery was isolated from the pulmonary vein; and cfDNA relapse was isolated at the time of relapse). Mutations are ordered according to their clonality established by primary tumor analysis.
Supplementary information
Supplementary Tables 1–12
Supplementary Tables 1-12
Rights and permissions
About this article
Cite this article
Chemi, F., Rothwell, D.G., McGranahan, N. et al. Pulmonary venous circulating tumor cell dissemination before tumor resection and disease relapse. Nat Med 25, 1534–1539 (2019). https://doi.org/10.1038/s41591-019-0593-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0593-1
This article is cited by
-
Single-cell low-pass whole genome sequencing accurately detects circulating tumor cells for liquid biopsy-based multi-cancer diagnosis
npj Precision Oncology (2024)
-
Resistance to BRAF inhibition explored through single circulating tumour cell molecular profiling in BRAF-mutant non-small-cell lung cancer
British Journal of Cancer (2024)
-
Tumor-derived cell-free DNA and circulating tumor cells: partners or rivals in metastasis formation?
Clinical and Experimental Medicine (2024)
-
A clinically feasible circulating tumor cell sorting system for monitoring the progression of advanced hepatocellular carcinoma
Journal of Nanobiotechnology (2023)
-
The presence of circulating genetically abnormal cells in blood predicts risk of lung cancer in individuals with indeterminate pulmonary nodules
BMC Pulmonary Medicine (2023)