Abstract
Since most dominant human mutations are single nucleotide substitutions1,2, we explored gene editing strategies to disrupt dominant mutations efficiently and selectively without affecting wild-type alleles. However, single nucleotide discrimination can be difficult to achieve3 because commonly used endonucleases, such as Streptococcus pyogenes Cas9 (SpCas9), can tolerate up to seven mismatches between guide RNA (gRNA) and target DNA. Furthermore, the protospacer-adjacent motif (PAM) in some Cas9 enzymes can tolerate mismatches with the target DNA3,4. To circumvent these limitations, we screened 14 Cas9/gRNA combinations for specific and efficient disruption of a nucleotide substitution that causes the dominant progressive hearing loss, DFNA36. As a model for DFNA36, we used Beethoven mice5, which harbor a point mutation in Tmc1, a gene required for hearing that encodes a pore-forming subunit of mechanosensory transduction channels in inner-ear hair cells6. We identified a PAM variant of Staphylococcus aureus Cas9 (SaCas9-KKH) that selectively and efficiently disrupted the mutant allele, but not the wild-type Tmc1/TMC1 allele, in Beethoven mice and in a DFNA36 human cell line. Adeno-associated virus (AAV)-mediated SaCas9-KKH delivery prevented deafness in Beethoven mice up to one year post injection. Analysis of current ClinVar entries revealed that ~21% of dominant human mutations could be targeted using a similar approach.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data accessibility
Raw sequencing files have been be uploaded to NCBI’s Sequence Read Archive (SRA) (https://www.ncbi.nlm.nih.gov/sra/PRJNA541170). A list of uploaded files including SRA IDs are listed in Supplementary Table 6. Detailed data analysis is available in the supplementary tables and supplementary data sets published with this manuscript. The pBG201 empty vector (pAAV-CMV-NLS(SV40)-SaCas9(E782K/N968K/R1015H)-NLS(nucleoplasmin)-3xHA-bGHpA0-U6-BsaI-sgRNA) is available upon completion of a standard Material Transfer Agreement with Harvard Medical School. The HAP1DFNA36 cell line is available upon request and completion of standard Material Transfer Agreement with Ecole Polytechnique Fédérale de Lausanne. Any other raw data including ABR traces and electrophysiological recordings that support the findings of this study are available from the corresponding author.
References
Landrum, M. J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D862–D868 (2016).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–198 (2015).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Vreugde, S. et al. Beethoven, a mouse model for dominant, progressive hearing loss DFNA36. Nat. Genet. 30, 257–258 (2002).
Pan, B. et al. TMC1 forms the pore of mechanosensory transduction channels in vertebrate inner ear hair cells. Neuron 99, 736–753 (2018).
Fu, Y., Sander, J. D., Reyon, D., Cascio, V. M. & Joung, J. K. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 32, 279–284 (2014).
Kleinstiver, B. P. et al. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Chen, J. S. et al. Enhanced proofreading governs CRISPR-Cas9 targeting accuracy. Nature 550, 407–410 (2017).
Slaymaker, I. M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science (80-) 351, 84–88 (2016).
Kleinstiver, B. P. et al. Broadening the targeting range of Staphylococcus aureus CRISPR-Cas9 by modifying PAM recognition. Nat. Biotechnol. 33, 1293–1298 (2015).
Kleinstiver, B. P. et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Christie, K. A. et al. Towards personalised allele-specific CRISPR gene editing to treat autosomal dominant disorders. Sci. Rep. 7, 16174 (2017).
Shin, J. W. et al. Permanent inactivation of Huntington’s disease mutation by personalized allele-specific CRISPR/Cas9. Hum. Mol. Genet. 25, 4566–4576 (2016).
Zhao, Y. et al. A novel DFNA36 mutation in TMC1 orthologous to the Beethoven (Bth) mouse associated with autosomal dominant hearing loss in a Chinese family. PLoS ONE 9, e97064 (2014).
Gao, X. et al. Treatment of autosomal dominant hearing loss by in vivo delivery of genome editing agents. Nature 553, 217–221 (2018).
Landegger, L. D. et al. A synthetic AAV vector enables safe and efficient gene transfer to the mammalian inner ear. Nat. Biotechnol. 35, 280–284 (2017).
Miller, D. G., Petek, L. M. & Russell, D. W. Adeno-associated virus vectors integrate at chromosome breakage sites. Nat. Genet. 36, 767–773 (2004).
Pan, B. et al. TMC1 and TMC2 are components of the mechanotransduction channel in hair cells of the mammalian inner ear. Neuron 79, 504–515 (2013).
Kawashima, Y. et al. Mechanotransduction in mouse inner ear hair cells requires transmembrane channel-like genes. J. Clin. Invest. 121, 4796–4809 (2011).
Askew, C. et al. Tmc gene therapy restores auditory function in deaf mice. Sci. Transl. Med. 7, 295ra108 (2015).
Shibata, S. B. et al. RNA interference prevents autosomal-dominant hearing loss. Am. J. Hum. Genet. 98, 1101–1113 (2016).
Yoshimura, H., Shibata, S. B., Ranum, P. T., Moteki, H. & Smith, R. J. H. Targeted allele suppression prevents progressive hearing loss in the mature murine model of human TMC1 deafness. Mol. Ther. 27, 681–690 (2019).
Narayan, D. S., Wood, J. P. M., Chidlow, G. & Casson, R. J. A review of the mechanisms of cone degeneration in retinitis pigmentosa. Acta Ophthalmol. 94, 748–754 (2016).
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Scheffer, D. I., Shen, J., Corey, D. P. & Chen, Z.-Y. Gene expression by mouse inner ear hair cells during development. J. Neurosci. 35, 6366–6380 (2015).
György, B. et al. CRISPR/Cas9 mediated disruption of the swedish APP allele as a therapeutic approach for early-onset Alzheimer’s disease. Mol. Ther. - Nucleic Acids 11, 429–440 (2018).
Zinn, E. et al. In silico reconstruction of the viral evolutionary lineage yields a potent gene therapy vector. Cell Rep. 12, 1056–1068 (2015).
Stauffer, E. A. & Holt, J. R. Sensory transduction and adaptation in inner and outer hair cells of the mouse auditory system. J. Neurophysiol. 98, 3360–3369 (2007).
Kotecki, M., Reddy, P. S. & Cochran, B. H. Isolation and characterization of a near-haploid human cell line. Exp. Cell Res. 252, 273–280 (1999).
Acknowledgements
We thank members of Holt/Géléoc and Corey laboratories for many helpful discussions and critical review of the manuscript. We thank the MGH DNA Core for sequencing services and the BCH Viral Core (BCH IDDRC, grant no. 1U54HD090255) for vector production. We also thank L. Pinello for assistance with CRISPResso analysis, V. Dillard for the help with TIDE analysis and A. A. Sousa for technical assistance with the GUIDE-Seq analysis. This work was supported by the Jeff and Kimberly Barber Fund and NIH grant nos. R01-DC013521 and R01-DC05439 (to J.R.H.), R01-DC000304 and R01-DC016932 (to D.P.C.), DC008853 (to G.S.G) and no. K99-CA218870, the Charles A. King Trust, a Banting Postdoctoral Fellowship (to B.P.K.), the Richard Floor Biorepository Fund, the Kosciuszko Foundation (to M.P.Z.) and by the Bertarelli Foundation. J.K.J. is supported by a Desmond and Ann Heathwood MGH Research Scholar award and B.G. was an Edward R. and Anne G. Lefler Center Postdoctoral Fellow.
Author information
Authors and Affiliations
Contributions
B.G. conceived the project, designed the experiments, analyzed the data, generated the figures and wrote the manuscript. C.N.-L. performed the hearing tests, confocal and SEM microscopy, and analyzed and generated the figures. B.P. performed in vivo injections and hair cell recordings, and analyzed and generated figures. Y.A. generated the viral vectors. K.D.K performed the functional tests for AAV-mediated CRISPR delivery to hair cells. B.P.K. assisted with genome editing and GUIDE-Seq experiments. S.P.G. performed the GUIDE-Seq off-target and the ClinVar dominant human mutation analyzes. M.P.Z. helped analyze the sequencing data. P.S., S.S. and B.S. designed and performed the HAP1 experiments and analyzed the data. J.K.J. designed the experiments and helped write the manuscript. G.S.G.G. performed the ABR and DPOAE recordings and analysis. D.P.C. designed the experiments and helped write the manuscript. J.R.H. conceived the project, designed the experiments, analyzed the data, generated the figures and wrote the manuscript. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.R.H. holds patents on TMC1 gene therapy and is an advisor to several biotech firms focused on inner-ear therapies. J.K.J. has financial interests in Beam Therapeutics, Editas Medicine, Pairwise Plants, Poseida Therapeutics, Transposagen Biopharmaceuticals and Verve Therapeutics. J.K.J.’s interests were reviewed and are managed by Massachusetts General Hospital and Partners HealthCare in accordance with their conflict of interest policies. J.K.J. holds equity in Excelsior Genomics, and is a member of the Board of Directors of the American Society of Gene and Cell Therapy. J.K.J. and B.P.K. are co-inventors on various patents and patent applications that describe gene editing and epigenetic editing technologies. The authors declare no other competing interests.
Additional information
Peer review information: Kate Gao and Brett Benedetti were the primary editors on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Targeting Tmc1Bth with SpCas9.
a, gRNA design for SpCas9. Mutation site is highlighted in red. PAM sites are marked by green nucleotides. Mismatching nucleotides are shown in blue. The numbers or letters (for example, 1.1) next to the PAM site represent gRNAs IDs. Our gRNA 1.1 is identical to the Tmc1-mut3 gRNA in the study of Gao et al.16. Plasmids encoding SpCas9-2A–GFP, along with the different gRNAs, were transfected into fibroblasts. Four days after transfection, GFP-positive cells were sorted by FACS. b, Sanger-sequencing traces from Tmc1Bth/WT or Tmc1WT/WT mouse fibroblasts transfected with SpCas9-2A–GFP with or without gRNA 1.1. GFP expressing cells were sorted by FACS 4 d after transfection. The mutation site is marked by a red arrow. Additional peaks appearing downstream (marked by black arrowheads) of the mutation site demonstrate sequence heterogeneity and thus indel formation. Similar results were obtained by all gRNAs from two technical replicates (forward and reverse sequencing). Genome editing is apparent both in Tmc1Bth/WT or Tmc1WT/WT cells with SpCas9 + gRNA 1.1. c, Sanger-sequencing data were analyzed by TIDE. The control sample (SpCas9–2A-GFP only, black) and the genome edited sample (SpCas9-2A–GFP + gRNA 1.1, green) are overlaid. Downstream of the expected cut site (blue dashed line) the percentage of aberrant sequences was quantified in the region for decomposition. d, Indel percentages (mean ± s.d.) in Tmc1Bth/WT or Tmc1WT/WT cells based on TIDE analysis. Cells were transfected in duplicates and two independent sequencing reactions (forward and reverse) were performed. No indel formation was observed in the case of 3.1, 3.2 and 3.3 gRNAs. gRNA 1.4 showed minimal, but specific genome editing on the Tmc1Bth/WT cells. All the other gRNAs mediated efficient indel formation both in Tmc1Bth/WT or Tmc1WT/WT cells. e, Targeted deep sequencing on control (SpCas9-2A–GFP only) cells, WT gRNA and the three most specific gRNAs (1.1, 2.1 and 2.4) in Tmc1Bth/WT (top) Tmc1WT/WT (bottom) cells. Indels were quantified after segregating Tmc1Bth and Tmc1WT reads by CRISPResso (only insertions and deletions were quantified, substitutions were ignored). None of the gRNAs are specific to the Tmc1Bth allele, and mediate efficient indel formation on the Tmc1WT allele as well (light blue). Sequencing was performed one time from pooled cells, transfected in triplicates. Numbers in pie charts represent the percentage of reads. Specificity was defined as the indel percentage towards the targeted allele among total indels. The gRNA with the highest selectivity towards the Tmc1Bth allele was gRNA 2.4. f, The most abundant reads in the SpCas9 + gRNA 2.4 treated cells, shown separately for Tmc1Bth (top) and Tmc1WT (bottom) reads. The CRISPR cut site is marked by a black dashed line. Dashes represent deleted nucleotides. Insertions are shown with nucleotides in red squares. Nucleotides in bold are substitutions; however, these were not quantified as CRISPR actions. Sequences were aligned to Bth allele; thus, in the bottom panel, WT reads appear as having a substitution (a T to A change). The mutation site is marked by red arrow. Indel formation is evident in both Tmc1Bth and Tmc1WT reads. g, Indel profiles from SpCas9 + gRNA 2.4 transfected Tmc1Bth/WT fibroblasts. Tmc1Bth and Tmc1WT reads are plotted separately. Minus numbers represent deletions, plus numbers represent insertions. Sequences without indels (value = 0) are omitted from the chart. The most common indel event is a single base deletion. h, Indels causing in-frame versus frame-shift mutations (percentages are shown) in the coding sequence after SpCas9 + gRNA 2.4 transfection. i, GUIDE-Seq analysis on SpCas9 + gRNA 2.4 transfected Tmc1Bth/WT fibroblasts. Genomic DNA was isolated from three biological replicates for sequencing on one occasion. Only one off-target site was identified. Numbers next to reads are read counts in the GUIDE-Seq assay.
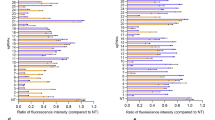
Extended Data Fig. 2 Number of indels based on targeted deep-sequencing data from Tmc1Bth/WT and Tmc1WT/WT cell lines treated with different Cas9 + gRNA combinations (from Fig. 1b).
Note that data points show non-normalized read counts. Cells were transfected on two different occasions (SpCas9 only and SpCas9 + gRNA 1.1 on four occasions) and genomic DNA from two independent biological samples on each transfection day were pooled for sequencing. Indels in the SaCas9-KKH treated Tmc1WT/WT lines are not different from the background (untreated samples). This method revealed high sensitivity, as the indel rates in CRISPR treated samples were 40–160-fold higher than the background indel rates observed in untreated samples.
Extended Data Fig. 3 Single nucleotide substitutions after Cas9 + gRNA treatment.
Cells were transfected on two different occasions (SpCas9 only and SpCas9 + gRNA 1.1 on four occasions) and genomic DNA from two independent biological samples on each transfection day were pooled for sequencing. Substitutions are given as percentages (that is normalized to total read counts) or non-normalized values (that is, the number of reads with substitutions). Analysis was performed on non-segregated .fastq files in Tmc1Bth/WT cells and in Tmc1WT/WT cells (top) and segregated.fastq files in Tmc1Bth/WT cells (bottom). Experimental conditions are the same as in Fig. 1b. Substitutions were not frequent (0.1–0.5% of reads) and there was no difference between untreated and treated samples in the percentage or number of reads with single nucleotide substitutions.
Extended Data Fig. 4 Background sequencing error versus indel formation with SaCas9-KKH (mean ± s.d.).
Background sequencing error rate (GFP only) and comparison indel events in SaCas9-KKH + gRNA 4.1/gRNA 4.2 and gRNA 4.3 transfected Tmc1WT/WT fibroblasts are plotted. The difference is not significant (two-tailed t-test, P = 0.3).
Extended Data Fig. 5 Effects of AAV–SaCas9-KKH–gRNA-4.2 on inner-ear function in WT and Bth mice.
a, Representative sensory transduction currents recorded at P14–P16 from IHCs and OHCs of WT mice injected with AAV–SaCas9-KKH–gRNA-4.2 at P1–P2. b, Mean ± s.e.m. maximal transduction current amplitudes for P14–P16 IHCs (left, n = 8) and OHCs (right, n = 6) from WT mice injected with AAV–SaCas9-KKH–gRNA-4.2 at P1–P2. c, Families of ABR waveforms recorded at 8 weeks from an uninjected Tmc1WT/WT mouse (left) and a Tmc1WT/WT mouse injected with AAV–SaCas9-KKH–gRNA-4.2 at P1 (right). Traces in bold indicate threshold. Scale bar applies to all traces. d, Mean ± s.d. ABR thresholds plotted as a function of stimulus frequency for six Tmc1WT/WT mice (black, n = 6) and three Tmc1WT/WT mice injected with AAV–SaCas9-KKH–gRNA-4.2 (gray) at 24 weeks of age (P = 0.9). e, Mean ± s.d. ABR thresholds plotted as a function of stimulus frequency for six Tmc1WT/WT mice (black), nine Tmc1Bth/WT and five Tmc1Bth/WT mice injected with AAV–SaCas9-KKH–gRNA-4.2 (green) at 24 weeks of age. 8 kHz thresholds for injected, 38 ± 11 dB (n = 9) and uninjected, 64 ± 19 dB (n = 8) were significantly different (P = 0.004, unpaired two-tailed t-test). ABR thresholds at higher frequencies (22 kHz) for injected (84 ± 15 dB, n = 9) and uninjected Tmc1Bth/WT mice (103 ± 5 dB, n = 8; P = 0.004, unpaired two-tailed t-test). f, Representative confocal images of 100-µm cochlear sections harvested at 24 weeks from the 8, 16 and 32 kHz regions from three uninjected Tmc1WT/WT mice (top) and three Tmc1WT/WT mice injected with AAV–SaCas9-KKH–gRNA-4.2 (bottom). The tissue was stained for MYO7A (red) and actin (green). g, Mean ± s.d., number of surviving IHCs (left) and OHCs (right) per 100-µm section (n = 3).
Extended Data Fig. 6 ABR amplitude, latencies and correlation of thresholds with surviving hair cells.
a,b, Peak 1 amplitudes (a) and peak 1 latencies (b) measured from 8 kHz ABR waveforms at the 8 week time point, from examples shown in Fig. 5a,c, for all Tmc1Bth/WT mice injected with AAV–SaCas9-KKH–gRNA-4.2 (green traces), and an example of an uninjected Tmc1Bth/WT (red trace). c, Mean ABR thresholds measured at 24 weeks, evoked by 8 and 16 kHz tone bursts (from Extended Data Fig. 5e) plotted as function of the mean percentage of surviving IHCs and OHCs from the 8 and 16 kHz regions (from Fig. 3h). The data were fit with a linear equation that had a slope of –2.1 dB per percentage point and a correlation coefficient of –0.82 (red line, Pearson’s r).
Extended Data Fig. 7 qPCR specific for ITR region in transfected plasmid (AAV–CMV–SaCas9-KKH–U6–gRNA) normalized to albumin gene in TMC1DFNA36 and TMC1WT HAP-1 cells.
Bars show mean ± s.d. Data points are from three independent biological replicates.
Extended Data Fig. 8 Number of indels based on targeted deep-sequencing data from TMC1DFNA36 and TMC1WT cells treated with different Cas9 + gRNA combinations (from Fig. 4c).
Note that data points show non-normalized read counts. Bars show mean ± s.d. Data points are from three independent biological replicates.
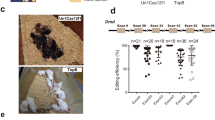
Extended Data Fig. 9 Targeted deep sequencing and CRISPResso analysis.
a, PCR and sequencing of the Tmc1 gene. b, Global indel analysis by CRISPResso. Reads were not segregated. c, Strategy to segregate Tmc1Bth reads and Tmc1WT reads. The region for splitting partly overlaps with indels and thus some reads cannot be assigned as mutant or WT. d, Allele-specific indel analysis by CRISPResso.
Supplementary information
Supplementary Information
Supplementary Notes 1–3
Supplementary Data Sets
Supplementary Data Sets 1–4
Supplementary Tables
Supplementary Tables 1–10
Rights and permissions
About this article
Cite this article
György, B., Nist-Lund, C., Pan, B. et al. Allele-specific gene editing prevents deafness in a model of dominant progressive hearing loss. Nat Med 25, 1123–1130 (2019). https://doi.org/10.1038/s41591-019-0500-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0500-9
This article is cited by
-
Genetic correction of induced pluripotent stem cells from a DFNA36 patient results in morphologic and functional recovery of derived hair cell-like cells
Stem Cell Research & Therapy (2024)
-
The applications of CRISPR/Cas-mediated genome editing in genetic hearing loss
Cell & Bioscience (2023)
-
Massively parallel evaluation and computational prediction of the activities and specificities of 17 small Cas9s
Nature Methods (2023)
-
Treatment of monogenic and digenic dominant genetic hearing loss by CRISPR-Cas9 ribonucleoprotein delivery in vivo
Nature Communications (2023)
-
Deafness: from genetic architecture to gene therapy
Nature Reviews Genetics (2023)