Abstract
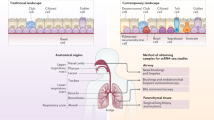
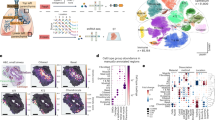
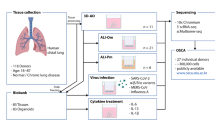
Human lungs enable efficient gas exchange and form an interface with the environment, which depends on mucosal immunity for protection against infectious agents. Tightly controlled interactions between structural and immune cells are required to maintain lung homeostasis. Here, we use single-cell transcriptomics to chart the cellular landscape of upper and lower airways and lung parenchyma in healthy lungs, and lower airways in asthmatic lungs. We report location-dependent airway epithelial cell states and a novel subset of tissue-resident memory T cells. In the lower airways of patients with asthma, mucous cell hyperplasia is shown to stem from a novel mucous ciliated cell state, as well as goblet cell hyperplasia. We report the presence of pathogenic effector type 2 helper T cells (TH2) in asthmatic lungs and find evidence for type 2 cytokines in maintaining the altered epithelial cell states. Unbiased analysis of cell–cell interactions identifies a shift from airway structural cell communication in healthy lungs to a TH2-dominated interactome in asthmatic lungs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data requests for raw and analyzed data and materials will fall under two categories. Datasets from healthy live volunteers and live volunteers with asthma will be promptly reviewed by the University of Groningen. Any data and materials that can be shared will be released via a Material Transfer Agreement. These datasets can be found on European Genome–phenome Archive (https://www.ebi.ac.uk/ega/home) EGAS00001001755. Datasets generated from lung resection samples using Drop-seq can be accessed in GSE130148. Datasets generated from deceased donors fall under Open Access Policies of the Human Cell Atlas (https://www.humancellatlas.org for details). This data can be accessed at European Genome–phenome Archive (https://www.ebi.ac.uk/ega/home) EGAS00001002649. Interactive exploration tool: www.lungcellatlas.org.
References
Tata, P. R. & Rajagopal, J. Plasticity in the lung: making and breaking cell identity. Development 144, 755–766 (2017).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Montoro, D. T. et al. A revised airway epithelial hierarchy includes CFTR-expressing ionocytes. Nature 560, 319–324 (2018).
Plasschaert, L. W. et al. A single-cell atlas of the airway epithelium reveals the CFTR-rich pulmonary ionocyte. Nature 560, 377–381 (2018).
Bisset, L. R. & Schmid-Grendelmeier, P. Chemokines and their receptors in the pathogenesis of allergic asthma: progress and perspective. Curr. Opin. Pulm. Med. 11, 35–42 (2005).
Colvin, R. A. et al. Synaptotagmin-mediated vesicle fusion regulates cell migration. Nat. Immunol. 11, 495–502 (2010).
Urawa, M. et al. Protein S is protective in pulmonary fibrosis. J. Thromb. Haemost. 14, 1588–1599 (2016).
Wujak, A. et al. FXYD1 negatively regulates Na+/K+-ATPase activity in lung alveolar epithelial cells. Respir. Physiol. Neurobiol. 220, 54–61 (2016).
Krotova, K. et al. Alpha-1 antitrypsin-deficient macrophages have increased matriptase-mediated proteolytic activity. Am. J. Respir. Cell Mol. Biol. 57, 238–247 (2017).
Vogl, T. et al. S100A12 is expressed exclusively by granulocytes and acts independently from MRP8 and MRP14. J. Biol. Chem. 274, 25291–25296 (1999).
Mitchell, A. et al. LILRA5 is expressed by synovial tissue macrophages in rheumatoid arthritis, selectively induces pro-inflammatory cytokines and IL-10 and is regulated by TNF-α, IL-10 and IFN-γ. Eur. J. Immunol. 38, 3459–3473 (2008).
Condon, T. V., Sawyer, R. T., Fenton, M. J. & Riches, D. W. H. Lung dendritic cells at the innate-adaptive immune interface. J. Leukoc. Biol. 90, 883–895 (2011).
Baumann, U., Routes, J. M., Soler-Palacín, P. & Jolles, S. The lung in primary immunodeficiencies: new concepts in infection and inflammation. Front. Immunol. 9, 1837 (2018).
Holgate, S. T. et al. Asthma. Nat. Rev. Dis. Primers 1, 15025 (2015).
Lopez-Guisa, J. M. et al. Airway epithelial cells from asthmatic children differentially express proremodeling factors. J. Allergy Clin. Immunol. 129, 990–997.e6 (2012).
Alcala, S. E. et al. Mitotic asynchrony induces transforming growth factor-β1 secretion from airway epithelium. Am. J. Respir. Cell Mol. Biol. 51, 363–369 (2014).
Harkness, L. M., Ashton, A. W. & Burgess, J. K. Asthma is not only an airway disease, but also a vascular disease. Pharmacol. Ther. 148, 17–33 (2015).
Balzar, S. et al. Mast cell phenotype, location, and activation in severe asthma. Data from the Severe Asthma Research Program. Am. J. Respir. Crit. Care Med. 183, 299–309 (2011).
Truyen, E. et al. Evaluation of airway inflammation by quantitative Th1/Th2 cytokine mRNA measurement in sputum of asthma patients. Thorax 61, 202–208 (2006).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Erle, D. J. & Sheppard, D. The cell biology of asthma. J. Cell Biol. 205, 621–631 (2014).
Danahay, H. et al. Notch2 is required for inflammatory cytokine-driven goblet cell metaplasia in the lung. Cell Rep. 10, 239–252 (2015).
Gomi, K., Arbelaez, V., Crystal, R. G. & Walters, M. S. Activation of NOTCH1 or NOTCH3 signaling skews human airway basal cell differentiation toward a secretory pathway. PLoS ONE 10, e0116507 (2015).
Ordovas-Montanes, J. et al. Allergic inflammatory memory in human respiratory epithelial progenitor cells. Nature 560, 649–654 (2018).
Luo, W. et al. Airway epithelial expression quantitative trait loci reveal genes underlying asthma and other airway diseases. Am. J. Respir. Cell Mol. Biol. 54, 177–187 (2016).
Wu, C. A. et al. Bronchial epithelial cells produce IL-5: implications for local immune responses in the airways. Cell. Immunol. 264, 32–41 (2010).
Laitinen, L. A., Laitinen, A. & Haahtela, T. Airway mucosal inflammation even in patients with newly diagnosed asthma. Am. Rev. Respir. Dis. 147, 697–704 (1993).
Arima, M. & Fukuda, T. Prostaglandin D2 and TH2 inflammation in the pathogenesis of bronchial asthma. Korean J. Intern. Med. 26, 8–18 (2011).
Xue, L. et al. Prostaglandin D2 activates group 2 innate lymphoid cells through chemoattractant receptor-homologous molecule expressed on TH2 cells. J. Allergy Clin. Immunol. 133, 1184–1194 (2014).
Dougherty, R. H. et al. Accumulation of intraepithelial mast cells with a unique protease phenotype in TH2-high asthma. J. Allergy Clin. Immunol. 125, 1046–1053.e8 (2010).
Hol, B. E., van de Graaf, E. A., Out, T. A., Hische, E. A. & Jansen, H. M. IgM in the airways of asthma patients. Int. Arch. Allergy Appl. Immunol. 96, 12–18 (1991).
Muehling, L. M., Lawrence, M. G. & Woodfolk, J. A. Pathogenic CD4+ T cells in patients with asthma. J. Allergy Clin. Immunol. 140, 1523–1540 (2017).
Oja, A. E. et al. Trigger-happy resident memory CD4+ T cells inhabit the human lungs. Mucosal Immunol. 11, 654–667 (2018).
Mitson-Salazar, A. et al. Hematopoietic prostaglandin D synthase defines a proeosinophilic pathogenic effector human TH2 cell subpopulation with enhanced function. J. Allergy Clin. Immunol. 137, 907–918.e9 (2016).
Wambre, E. et al. A phenotypically and functionally distinct human T H2 cell subpopulation is associated with allergic disorders. Sci. Transl. Med. 9, eaam9171 (2017).
Lam, E. P. S. et al. IL-25/IL-33-responsive TH2 cells characterize nasal polyps with a default TH17 signature in nasal mucosa. J. Allergy Clin. Immunol. 137, 1514–1524 (2016).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal–fetal interface in humans. Nature 563, 347–353 (2018).
Weckmann, M., Kopp, M. V., Heinzmann, A. & Mattes, J. Haplotypes covering the TNFSF10 gene are associated with bronchial asthma. Pediatr. Allergy Immunol. 22, 25–30 (2011).
Harada, M. et al. Thymic stromal lymphopoietin gene promoter polymorphisms are associated with susceptibility to bronchial asthma. Am. J. Respir. Cell Mol. Biol. 44, 787–793 (2011).
Grotenboer, N. S., Ketelaar, M. E., Koppelman, G. H. & Nawijn, M. C. Decoding asthma: translating genetic variation in IL33 and IL1RL1 into disease pathophysiology. J. Allergy Clin. Immunol. 131, 856–865 (2013).
Holgate, S. T. et al. Epithelial-mesenchymal communication in the pathogenesis of chronic asthma. Proc. Am. Thorac. Soc. 1, 93–98 (2004).
Heijink, I. H. et al. Down-regulation of E-cadherin in human bronchial epithelial cells leads to epidermal growth factor receptor-dependent Th2 cell-promoting activity. J. Immunol. 178, 7678–7685 (2007).
Song, J. et al. Aberrant DNA methylation and expression of SPDEF and FOXA2 in airway epithelium of patients with COPD. Clin. Epigenetics 9, 42 (2017).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
van den Brink, S. C. et al. Single-cell sequencing reveals dissociation-induced gene expression in tissue subpopulations. Nat. Methods 14, 935–936 (2017).
Young, M. D. & Behjati, S. SoupX removes ambient RNA contamination from droplet based single cell RNA sequencing data. Preprint at https://doi.org/10.1101/303727 (2018).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, P281–291.E9 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Mereu, E. et al. matchSCore: matching single-cell phenotypes across tools and experiments. Preprint at https://doi.org/10.1101/314831 (2018).
Acknowledgements
We thank J. Eliasova (scientific illustrator) for support with design of figures, the Sanger Single Cell Genomics Core Facility for support with the SmartSeq2 protocol, E. Rawlins for feedback and critical reading of the manuscript, L. Mamanova for the technical support, as well as all the members of the Teichmann lab for scientific input. We are grateful to the Cambridge Biorepository for Translational Medicine (CBTM) for the provision of tissue from deceased organ donors, to all tissue donors, and to K. Sjollema and the UMCG Imaging and Microscopy Center. We gratefully acknowledge the provision of human biomaterial and clinical data from the CPC-M bioArchive and its partners at the Asklepios Biobank Gauting, the Klinikum der Universität München, and the Ludwig-Maximilians-Universität München. This project was funded by and part of the Open Targets collaboration (https://www.opentargets.org), a GlaxoSmithKline collaborative agreement with University Medical Center Groningen, Wellcome (WT206194), the Lung Foundation Netherlands (projects no. 5.1.14.020 and 4.1.18.226), Health-Holland, Top Sector Life Sciences and Health, and Human Cell Atlas Wellcome Stragetic Science Support (211276/Z/18/Z). R.V.-T. was supported by EMBO and HFSP Long Term fellowships. T.G. by the Marie Curie ENLIGHT-TEN training network. L.M.S. acknowledges funding from the European Union’s Horizon 2020 research and innovation program under the Marie Sklodowska-Curie grant agreement no. 753039. H.B.S. acknowledges funding by the Helmholtz Association and the German Center for Lung Research (DZL). F.J.T. acknowledges financial support by the German Research Foundation (DFG) within the Collaborative Research Centre 1243. F.J.T acknowledges subproject A17, by the Helmholtz Association (Incubator grant sparse2big, grant number ZT-I-0007) and by the Chan Zuckerberg Initiative D.A.F. (advised fund of Silicon Valley Community Foundation), grant number 182835. A.C. and P.M.S. acknowledge European Research Council (project 677501 – ZF_Blood). This publication is part of the Human Cell Atlas (www.humancellatlas.org/publications).
Author information
Authors and Affiliations
Contributions
S.A.T., M.C.N., M.v.d.B., K.A., A.J.v.O. and H.B.S. designed the project. F.A.V.B., M.C.N. and S.A.T. wrote the paper. F.A.V.B., O.A.C., S.B., L.H., J.J., E.S.F., P.M.S., K.T.M., I.A., M.R.J. and M.S. generated the data. F.A.V.B., G.K., M.B., L.M.S., T.G., J.J., M.E., S.P., K.P., H.V.F. and F.J.T. analyzed the data. F.A.V.B., G.K., O.A.C., L.M.S., T.G., J.J., R.V.-T., K.A., S.P., A.C., K.S.-P., W.T., G.H.K., A.J.v.O., C.T.-L., H.B.S., M.v.d.B, F.J.T., M.B. and K.B.M. interpreted the data. All authors read the manuscript, offered feedback, and approved it before submission.
Corresponding authors
Ethics declarations
Competing interests
K.A. and A.J.v.O. are employees of GlaxoSmithKline. M.B., O.A.C., S.B., M.v.d.B. and M.C.N. received project funding from GlaxoSmithKline.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Unbiased clustering of upper airway, lower airway, and parenchymal lung tissue cells and strategy to divide the dataset used in Figs. 1 and 2.
a, t-SNE plot depicting unbiased cluster assignment of combined dataset of parenchyma and upper and lower airways, highlighting EPCAM-high cell clusters (epithelial cells) in blue and other cell types in green. b, t-SNE as in a, highlighting expression of cell lineage markers. c, t-SNE plot of only epithelial cells, as defined in a. d, t-SNE as in c, showing expression of individual genes used for cell-type assignment of different epithelial subsets. e, Table depicting cell-type assignment of clusters identified in c. Alveolar type 1 cells were not identified in an individual cluster in this analysis and they were selected by the high expression of the type 1 marker AGER. Panels a–e, n = 12 individuals. Complete sample distribution in Fig. 1b.
Extended Data Fig. 2 Resection lung material analysis via Drop-seq.
a, t-SNE showing single cells obtained from lung resection samples analyzed using Drop-seq. b, Heat map depicting top differentially expressed genes by log fold change among the clusters present in a. c, matchSCore analysis comparing the clusters present in the Drop-seq analysis of lung resection material to the clusters identified in Figs. 1 and 2. Panels a and b, n = 4 individuals. Panel c, n = 16 individuals.
Extended Data Fig. 3 Ionocyte cell identity and molecular profile.
a, Immunohistochemistry staining for FOXI1, using hematoxylin as counterstain. b, Fluorescent staining for ionocyte markers: mouse anti-FOXI1 and rabbit anti-CFTR. c–f, Fluorescent staining for FOXI1 (ionocyte-specific) and other epithelial markers: neuroendocrine cell marker (rabbit anti-synaptophysin) (c), ciliated cell marker (mouse anti-α-tubulin) (d), basal cell marker (mouse anti-α-KRT5) (e), and secretory cell marker (mouse anti-MUC5AC) (f). DAPI (blue) stains nuclei. Representative picture out of ten individuals analyzed.
Extended Data Fig. 4 Marker gene analysis of specific goblet, ciliated, and neuroendocrine cells in human lungs.
a, Heat map depicting the expression of marker genes identified by differential expression analysis comparing goblet 1 versus goblet 2 cell types. b, Heat map depicting the expression of marker genes identified by differential expression analysis. Ciliated 1 genes and shared nasal signature genes generated by comparing ciliated 1 versus ciliated 2. Ciliated 2 genes generated by comparing ciliated 2 versus the combination of ciliated 1, goblet 1, and goblet 2, to subtract the nasal signature. c, t-SNE plot depicting the expression of the neuroendocrine marker CHGA. d, Heat map depicting the expression of neuroendocrine markers in neuroendocrine cells as well as in the other clusters identified in our analysis. Neuroendocrine cells identified based on CHGA expression. Neuroendocrine markers obtained from ref. 24. Panels a–d, n = 12 individuals. Complete sample distribution in Fig. 1b.
Extended Data Fig. 5 Cell-type assignment strategy for the assignment of non-epithelial cells discussed in Fig. 2.
a, t-SNE colored by unbiased clustering of the non-epithelial dataset described in Fig. 2. b, t-SNE depicting single cells colored by their respective sample origin. c, t-SNE as in a, showing the expression of lineage markers used for cell-type assignment. d, Table with the strategy used for cluster cell assignment of the cell types present in Fig. 2. e, Cluster distribution in the two donors from which we collected paired nasal brushes, airway brushes, and airway biopsies. f, Violin plots depicting the expression of immunoglobulin genes in the B cell cluster divided by tissue of origin. Representative genes of IgG, IgA, IgE, and IgM producing B cells depicted. Panels a–f, n = 12 individuals. Complete sample distribution in Fig. 2a.
Extended Data Fig. 6 Unbiased clustering of lower airway biopsy samples of healthy volunteers and volunteers with asthma and the strategy used to divide the dataset depicted in Fig. 3.
a, t-SNE plot depicting unbiased cluster assignment of combined dataset of healthy control and asthma lower airway biopsies, highlighting EPCAM-high cell clusters (epithelial cells) in blue and other cell types in green. b, t-SNE as in a, highlighting expression of cell lineage markers. c, t-SNE plot of only epithelial cells, as defined in a. d, t-SNE as in c, showing expression of individual genes used for cell-type assignment of different epithelial subsets. e, Table depicting cell-type assignment of clusters identified in c. f, Fluorescence in situ hybridization image of lung airway biopsy of one asthma patient. Green marks FOXJ1 (ciliated marker), red marks MUC5AC (secretory marker), and blue marks DAPI (nuclei). White arrow points to areas in which both MUC5AC and FOXJ1 are co-expressed in the exact same region (yellow color) or are contained in the vicinities of one nucleus, suggesting co-expression of the transcripts in the same cell. n = 6 healthy volunteers and 6 volunteers with asthma. Full sample distribution in Fig. 3b. Panels a–e, n = 12 individuals. Panel f depicts the analysis performed in n = 1 asthma donor.
Extended Data Fig. 7 IL-4/IL-13- and Notch-driven gene transcription signatures in goblet cell metaplasia in asthma.
a, Violin plot depicting IL-4/IL-13 genes signature expression in each cluster of biopsy epithelial cells (signature obtained from ref. 24). b, Violin plot depicting the expression of the IL-4/IL-13 signature (as in a) in healthy control and asthma cells of selected clusters. c, Violin plot depicting Notch signature genes (HES4, HES5, HEY1, HEY2, HEYL, and NRARP) expression in each cluster of biopsy epithelial cells. d, Violin plot depicting the expression of Notch signature genes (as in c) in healthy control and asthma cells of selected clusters. e–g, Expression of secretory specific (HES4 and SPDEF) and ciliated specific (FOXJ1) transcription factors in club (e), goblet (f), ciliated (g), and mucous ciliated (h) cells. Panels a–h, n = 12 individuals. Complete sample distribution in Fig. 3b.
Extended Data Fig. 8 Clustering and cell-type assignment of non-epithelial cells in the airways of healthy control patients and patients with asthma and their expression of prostaglandin enzymes.
a, t-SNE depicting unbiased clustering of the non-epithelial dataset described in Fig. 4. b, t-SNE as in a, showing the expression of lineage markers used for cell-type assignment. c, Table with the strategy used for cluster cell assignment of the cell types present in Fig. 4. d, Cartoon illustrating the eicosanoids pathway, its receptors and in which cells each gene is expressed in the lungs, with special notation for asthma alterations. Panels a–c, n = 12 individuals. Complete sample distribution in Fig. 4a. Panel d was generated based on n = 15 individuals. Complete sample distribution in Figs. 3b, 4a, and 5b.
Extended Data Fig. 9 Comparative analysis of bulk transcriptomes versus single-cell RNA-seq data on matched airway biopsies.
a, UMAP displaying bulk transcriptomes of biopsies before digestion (blue), of the single-cell suspension obtained from digestion and used to load the 10x microfluidics device (orange) and pseudo-bulk samples generated by collapsing all the data obtained from single-cell transcriptomes (green). b, Heat map depicting expression of several epithelial and non-epithelial cell markers, as shown in Extended Data Figs. 6 and 8. Gene markers of rare cells (neuroendocrine, eosinophils, and tuft) not identified in our single-cell clusters depicted in specific colors. Panels a–c, n = 8 individuals.
Extended Data Fig. 10 Canonical T cell markers expression in TH CD4 clusters.
a, Heat map depicting genes differentially expressed in the nine clusters of CD4 T cells using an initial unbiased clustering method, followed by manual selection of TH1, TH2, and TH17 clusters based on canonical cytokines. a–f, Violin plots depicting canonical markers of tissue-resident memory cells (b) and TH1 (c), TH2 (d), TH17 (e), and Treg (f) CD4 T cells. Depicted markers based on literature search of well-established T cell subsets. Panels a–f, n = 15 individuals. Complete sample distribution in Fig. 5b.
Supplementary information
Supplementary Information
Supplementary Tables 1–10
Source data
Source Data Fig. 1
Cell numbers per cluster used to design Fig. 1d,e
Source Data Fig. 2
Cell numbers per cluster used to design Fig. 2c,d
Source Data Fig. 3
Cell numbers per cluster used to design Fig. 3c
Source Data Fig. 4
Cell numbers per cluster used to design Fig. 4c
Source Data Fig. 5
Cell numbers per cluster used to design Fig. 5d,e,h
Source Data Fig. 6
P values calculated from cellPhone DB used to design Fig. 6a–d
Rights and permissions
About this article
Cite this article
Vieira Braga, F.A., Kar, G., Berg, M. et al. A cellular census of human lungs identifies novel cell states in health and in asthma. Nat Med 25, 1153–1163 (2019). https://doi.org/10.1038/s41591-019-0468-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0468-5
This article is cited by
-
Single-cell analysis reveals alterations in cellular composition and cell-cell communication associated with airway inflammation and remodeling in asthma
Respiratory Research (2024)
-
A single-cell atlas of lung homeostasis reveals dynamic changes during development and aging
Communications Biology (2024)
-
Semi-supervised integration of single-cell transcriptomics data
Nature Communications (2024)
-
Airway hillocks are injury-resistant reservoirs of unique plastic stem cells
Nature (2024)
-
A high-resolution view of the heterogeneous aging endothelium
Angiogenesis (2024)