Abstract
Malformations of the human cortex represent a major cause of disability1. Mouse models with mutations in known causal genes only partially recapitulate the phenotypes and are therefore not unlimitedly suited for understanding the molecular and cellular mechanisms responsible for these conditions2. Here we study periventricular heterotopia (PH) by analyzing cerebral organoids derived from induced pluripotent stem cells (iPSCs) of patients with mutations in the cadherin receptor–ligand pair DCHS1 and FAT4 or from isogenic knockout (KO) lines1,3. Our results show that human cerebral organoids reproduce the cortical heterotopia associated with PH. Mutations in DCHS1 and FAT4 or knockdown of their expression causes changes in the morphology of neural progenitor cells and result in defective neuronal migration dynamics only in a subset of neurons. Single-cell RNA-sequencing (scRNA-seq) data reveal a subpopulation of mutant neurons with dysregulated genes involved in axon guidance, neuronal migration and patterning. We suggest that defective neural progenitor cell (NPC) morphology and an altered navigation system in a subset of neurons underlie this form of PH.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The scRNA-seq data used in this study have been deposited in the Gene Expression Omnibus under accession number GSE124031. All relevant accession codes are provided. Further details on the methods can be found in the Life Sciences Reporting Summary. Additional data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Cappello, S. et al. Mutations in genes encoding the cadherin receptor-ligand pair DCHS1 and FAT4 disrupt cerebral cortical development. Nat. Genet. 45, 1300–1308 (2013).
Romero, D. M., Bahi-Buisson, N. & Francis, F. Genetics and mechanisms leading to human cortical malformations. Semin. Cell. Dev. Biol. 76, 33–75 (2018).
Mansour, S. et al. Van Maldergem syndrome: further characterisation and evidence for neuronal migration abnormalities and autosomal recessive inheritance. Eur. J. Hum. Genet. 20, 1024–1031 (2012).
Liu, J. S. Molecular genetics of neuronal migration disorders. Curr. Neurol. Neurosci. Rep. 11, 171–178 (2011).
Cardoso, C. et al. Periventricular heterotopia, mental retardation, and epilepsy associated with 5q14.3-q15 deletion. Neurology 72, 784–792 (2009).
Dubeau, F. et al. Periventricular and subcortical nodular heterotopia. A study of 33 patients. Brain 118(Pt 5), 1273–1287 (1995).
Tassi, L. et al. Electroclinical, MRI and neuropathological study of 10 patients with nodular heterotopia, with surgical outcomes. Brain 128, 321–337 (2004).
Aghakhani, Y. et al. The role of periventricular nodular heterotopia in epileptogenesis. Brain 128, 641–651 (2005).
Heinzen, E. L. et al. De novo and inherited private variants in MAP1B in periventricular nodular heterotopia. PLoS Genet. 14, e1007281 (2018).
O’Neill, A. C. et al. A primate-specific isoform of PLEKHG6 regulates neurogenesis and neuronal migration. Cell Rep. 25, 2729–2741.e6 (2018).
Lancaster, M. A. & Knoblich, J. A. Generation of cerebral organoids from human pluripotent stem cells. Nat. Protoc. 9, 2329–2340 (2014).
Ishiuchi, T., Misaki, K., Yonemura, S., Takeichi, M. & Tanoue, T. Mammalian fat and dachsous cadherins regulate apical membrane organization in the embryonic cerebral cortex. J. Cell Biol. 185, 959–967 (2009).
Cappello, S. et al. The Rho-GTPase cdc42 regulates neural progenitor fate at the apical surface. Nat. Neurosci. 9, 1099–1107 (2006).
Pacary, E. et al. Proneural transcription factors regulate different steps of cortical neuron migration through Rnd-mediated inhibition of RhoA signaling. Neuron 69, 1069–1084 (2011).
Camp, J. G. et al. Human cerebral organoids recapitulate gene expression programs of fetal neocortex development. Proc. Natl Acad. Sci. USA 112, 15672–15677 (2015).
Topol, A., Tran, N. N. & Brennand, K. J. A guide to generating and using hiPSC derived NPCs for the study of neurological diseases. J. Vis. Exp. https://doi.org/10.3791/52495 (2015).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell developmental trajectories.. Nat. Methods 14, 979–982 (2017).
Wang, J. et al. Epilepsy-associated genes. Seizure 44, 11–20 (2017).
Riesenberg, S. & Maricic, T. Targeting repair pathways with small molecules increases precise genome editing in pluripotent stem cells. Nat. Commun. 9, 2164 (2018).
Meyer, M. & Kircher, M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harb. Protoc. 2010, pdb.prot5448 (2010).
Renaud, G., Stenzel, U. & Kelso, J. leeHom: adaptor trimming and merging for Illumina sequencing reads. Nucleic Acids Res. 42, e141 (2014).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Boyer, L. F., Campbell, B., Larkin, S., Mu, Y. & Gage, F. H. Dopaminergic differentiation of human pluripotent cells. Curr. Protoc. Stem Cell Biol. Chapter 1, Unit1H.6 (2012).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Pilz, G.-A. et al. Amplification of progenitors in the mammalian telencephalon includes a new radial glial cell type. Nat. Commun. 4, 2125 (2013).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Treutlein, B. et al. Reconstructing lineage hierarchies of the distal lung epithelium using single-cell RNA-seq. Nature 509, 371–375 (2014).
Renaud, G., Kircher, M., Stenzel, U. & Kelso, J. freeIbis: an efficient basecaller with calibrated quality scores for Illumina sequencers. Bioinformatics 29, 1208–1209 (2013).
Renaud, G., Stenzel, U., Maricic, T., Wiebe, V. & Kelso, J. deML: robust demultiplexing of Illumina sequences using a likelihood-based approach. Bioinformatics 31, 770–772 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2008).
Fietz, S. A. et al. Transcriptomes of germinal zones of human and mouse fetal neocortex suggest a role of extracellular matrix in progenitor self-renewal. Proc. Natl Acad. Sci. 109, 11836–11841 (2012).
Kang, H. M. et al. Multiplexed droplet single-cell RNA-sequencing using natural genetic variation. Nat. Biotechnol. 36, 89–94 (2018).
Acknowledgements
We thank the families participating in this study for their involvement. We thank Y. Lu for help generating the microRNAs, M. Karow and I. Buchsbaum for helping with experiments and fruitful discussions in the lab, T. Öztürk for excellent technical support, A. Weigert for organoid culture, J. Kageyama for helping with data processing, R. Snabel for helping with Smart-seq2 libraries, the Core Unit Flow Cytometry at the Zentrum für Infektionsmedizin (veterinary faculty of the University of Leipzig) and the Core Unit Qualitätsmanagement/Technologieplattform at the Sächsischer Inkubator für Klinische Translation (SIKT) in Leipzig for karyotyping. This work was supported by funding from the DFG CA1205/2-1 (S.C.), ForIPS (M.G.), by the Max Planck Society (S.C., B.T.), by the Boehringer Ingelheim Fonds (S.K.), by the Health Research Council of NZ and Curekids (S.P.R.) and by an ERC Starting Grant (B.T.).
Author information
Authors and Affiliations
Contributions
S.C. conceived and designed the research project, J.K., S.K., C.K., A.C.A.-M., R.D.G., S.R., A.C.O., C.T., M. Santel. and E.R. performed experiments and collected data, J.K., C.K., S.K., J.G.C., M. Schroeder, B.T. and S.C. analyzed data, M.D. reprogrammed patients’ samples, M.G. was involved in the start of the project, contributed to data discussion and supervision of J.K. S.P.R. was involved in patient sample collection and critical discussion, J.K., S.K., C.K., B.T. and S.C. wrote the manuscript. All authors provided ongoing critical review of results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Expression of DCHS1 and FAT4 in cerebral organoids and temporal development of DCHS1- and FAT4-mutant organoids.
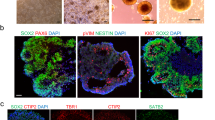
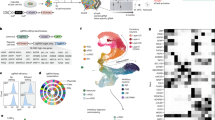
a, Axial T1 image demonstrating laminar PH lining the occipital horns and peritrigonal region of the ventricles (arrows). b, Axial T2 demonstrating linear lesions iso-intense with cortical gray matter adjacent to the lateral walls of the bodies of the lateral ventricles2. c, Schematic representation of the main experimental approach used. d, Timeline of the organoid generation. EBs, embryoid bodies; hESC, low-bFGF human embryonic stem cell medium; NIM, neural induction medium; Y, Rock inhibitor; NDM, neural differentiation medium; +/– A: B27 supplement with or without vitamin; IHC, immunohistochemistry; RNA-seq, RNA sequencing. e–h′, Detection of DCHS1 and FAT4 by in situ hybridization, b = 2, o = 5 per condition. i,j, mRNA expression of DCHS1 and FAT4 in single cells (c = 316) derived from control organoids, showing similar expression patterns between DCHS1 and FAT4 but often in different cells. i, Cells (columns) are ordered based on their PC 2 loading, corresponding to the trajectory from NPCs to neurons. Side bar shows maximal zone correlation for each single cell (ventricular zone, VZ, yellow; inner subventricular zone, iSVZ, orange; outer SVZ, oSVZ, red; cortical plate, CP, purple). j, Biplot showing transcript levels (in log2(FPKM)) of FAT4 (x axis) and DCHS1 (y axis) in 316 single cells of control organoids. k,l, Temporal development of patient-derived cerebral organoids compared with control organoids (o = 14 CTRL, 17 DCHS1, 17 FAT4); the diameters of organoids derived from DCHS1- and FAT4-mutant cells are slightly smaller compared to those of control organoids until day 12 (d12), shown in k. Significance based on two-way ANOVA, P = 0.000, Tukey HSD post hoc for multiple comparisons was performed for defining statistical differences between the three genotypes. Dotted lines highlight ventricles (V). Data in graphs are represented as mean ± s.e.m. Scale bars, 100 µm in e–h′,k.
Extended Data Fig. 2 Heterotopically located neurons in DCHS1 and FAT4-mutant and KO organoids.
a–c,e–j,l–o, Micrographs of sections of mutant or KO organoids immunostained as indicated in the panels. Note the mispositioning of neurons marked by arrows in mutant or in electroporated cerebral organoids. a–c, b = 2, o = 6 per condition; e–j, b = 2, o = 6 per condition; l–lʹʹʹ, b = 5, o = 15 per condition; m–o, b = 5, o = 15 per condition. d, MAP2 fluorescence intensity measured only in the ventricular zones (VZ) (v = 6 CTRL, 18 DCHS1, 11 FAT4; significance based on one-way ANOVA, P = 0.0166, Tukey HSD post hoc for multiple comparisons for defining statistical differences between the three genotypes). k, Quantification measured by qPCR of the knockdown of DCHS1 and FAT4 by microRNAs (miRNA) against DCHS1 or FAT4, respectively (b = 2, independent cultures per time = 3 CTRL, 3 miRNA DCHS1, 3 miRNA FAT4, significance based on one-way ANOVA, P = 0.0377, Tukey HSD post hoc for multiple comparisons for defining statistical differences between the three genotypes) in SH-SY5Y cells 48 h after nucleofection. l–lʹʹʹ, Nodule of TUBB3+ neurons intermingling with NESTIN+ processes of NPCs in the germinal zone of DCHS1-mutant organoids. m–o DCHS1- and FAT4-mutant organoids show changes in the morphology and thickness of their neuritis, as depicted by arrowheads. Dotted lines highlight ventricles (V). Data in graphs are represented as mean ± s.e.m. Scale bars, 100 µm in a–c and e–j and 30 µm in l–o. Source data.
Extended Data Fig. 3 Neural progenitor proliferation and signatures in mutant organoids.
a–i,k–m, Micrographs of sections of mutant cerebral organoids from day 20 immunostained for NESTIN (a–c) and DCX (d–f) and from day 42 immunostained for PAX6 (g–i) and PH3 (k–m). a–f, b = 1, o = 3 per condition; g–i,k–m, b = 3, o = 9 per condition. j, Quantification of thickness of ventricular zone (VZ) and cortical plate (CP) structures in cerebral organoids (v = 6 CTRL, 11 DCHS1, 12 FAT4). n, Quantification of PH3+ cells per length of apical surface (o = 3, v = 23 CTRL, 41 DCHS1, 17 FAT4, F(2,80) = 2.41, P = 0.097, comparison between CTRL and FAT4 F(1,39) = 5.18, P = 0.029). o, Hierarchical clustering visualizing for all NPCs (338 single cells), expression of genes identified by PCA (top 50 positively and negatively correlating with PC 1) on all NPCs. p, Number of NPCs and neurons for each experiment shown in o. q, FACS plots depicting the definition of the sorting gates (secondary antibodies control) and sorting of KI67+ or DCX+ cells in control, DCHS1- and FAT4-mutant organoids (b = 2, o = 6 CTRL, 6 DCHS1, 3 FAT4). r,s, Z scores of the quantification of KI67+ cells (r) and DCX+ cells (s) from FACs analysis shown in q. Statistical analysis was performed using two-tailed Mann Whitney test. r, CTRL to DCHS1 P = 0.0022, CTRL to FAT4 P = 0.0238. s, CTRL to DCHS1 P = 0.0317, CTRL to FAT4 P = 0.0357. Results are mean ± s.e.m. (j,n) or z scores as mean ± s.e.m. (r,s). Dotted lines highlight ventricles (V). Scale bars, 30 µm for a–f and 50 µm for g–m.
Extended Data Fig. 4 Morphological changes in the NPCs upon DCHS1 and FAT4 deletion.
a–c, Micrographs of sections of KO cerebral organoids day 40 immunostained for NESTIN. Arrows indicate the disrupted morphology of NPCs (b,c). d–f, Micrographs of sections of control organoids electroporated with miRNA against DCHS1 or FAT4 at day 42 and analyzed at day 49. Arrows indicate the disrupted morphology upon downregulation of DCHS1 and FAT4 (e,f). a–c, b = 2, o = 6 per condition; d–f, b = 2, o = 6 per condition. Dotted lines highlight ventricles (V). Scale bars, 30 µm.
Extended Data Fig. 5 Apicobasal polarity in cerebral organoids.
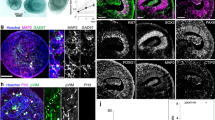
a–c, Micrographs of sections of organoids (day 42) immunostained for ACETYLATED TUBULIN, b = 2, o = 6 per condition. d,e, Western blot and quantification of ACETYLATED TUBULIN (d) (independent cultures = 5 CTRL, 4 DCHS1, 4 FAT4, significance based on one sample two-tailed t test, P = 0.562 CTRL vs DCHS1, P = 0.013 CTRL vs FAT4) and TYROSINATED TUBULIN (e) (independent cultures = 3 CTRL, 3 DCHS1, 3 FAT4) levels in NPCs, significance based on one sample two-tailed t test, P = 0.967 CTRL vs DCHS1, P = 0.728 CTRL vs FAT4. f–n,p–r, Micrographs of sections of cerebral organoids (day 42) immunostained as indicated in the panels. f–n,p–r, b = 3, o = 9 per condition. o, Quantification of the distance from the apical surface (positive for β-CATENIN and PALS1) and DAPI+ nuclei of NPCs (v = 27 CTRL, 26 DCHS1, 26 FAT4; 5 different positions were measured and averaged for each ventricle; significance based on one-sample two-tailed t test, P = 0.0025 CTRL vs DCHS1, P = 0.001 CTRL vs FAT4). s, Ratio of ARL13B fluorescence intensity measured at the apical surface (cilia facing the ventricular lumen) and at basal position (all the rest of the cilia in the germinal and cortical zones) (b = 5, o = 15, v = 9 CTRL, 4 DCHS1, 14 FAT4, significance based on one-sample t test). Results are mean ± s.e.m. Dotted lines highlight ventricles (V). Scale bar, 50 µm (a–c) and 20 µm (f–r).
Extended Data Fig. 6 2D time-lapse imaging experimental design and morphological changes of mutant neurons.
a, Experimental design for 2D live imaging of migrating neurons. Neural progenitors were differentiated for 7 d and imaged for 3 d every 5 min. b, Quantification of the percentage of cells expressing DCX, MAP2 or both markers in neuronal cultures after 10 d in culture. c–cʹʹ, Immunostaining for DoubleCortin (DCX), MAP2 (mature neurons) and VGLUT (glutamatergic neurons) in 2D neurons derived from control cells in monolayer culture, b = 3, independent cultures = 9 per condition. d–f, Immunostaining for TUBB3 in 2D neurons derived from control and mutant cells in monolayer culture, b = 3, independent cultures = 9 per condition. Results are mean ± s.e.m. Scale bar, 20 µm.
Extended Data Fig. 7 Characterization of mutant and KO organoid regions.
a–l, Micrographs of sections of mutant and KO organoids immunostained as depicted in the panels. a–c,g–i, b = 3, o = 9 per condition; d–f,j–l, b = 2, o = 6 per condition. m, Heat map showing expression of genes marking neurons in the cortex/forebrain (columns) for all neuronal cells from control and mutant organoids (rows, 467 single cells). Scale bar, 30 µm.
Supplementary information
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 5
Unprocessed Western Blots
Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 6
Statistical Source Data
Rights and permissions
About this article
Cite this article
Klaus, J., Kanton, S., Kyrousi, C. et al. Altered neuronal migratory trajectories in human cerebral organoids derived from individuals with neuronal heterotopia. Nat Med 25, 561–568 (2019). https://doi.org/10.1038/s41591-019-0371-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0371-0