Abstract
Quantitative changes in leptin concentration lead to alterations in food intake and body weight, but the regulatory mechanisms that control leptin gene expression are poorly understood. Here we report that fat-specific and quantitative leptin expression is controlled by redundant cis elements and trans factors interacting with the proximal promoter together with a long noncoding RNA (lncOb). Diet-induced obese mice lacking lncOb show increased fat mass with reduced plasma leptin levels and lose weight after leptin treatment, whereas control mice do not. Consistent with this finding, large-scale genetic studies of humans reveal a significant association of single-nucleotide polymorphisms (SNPs) in the region of human lncOb with lower plasma leptin levels and obesity. These results show that reduced leptin gene expression can lead to a hypoleptinemic, leptin-responsive form of obesity and provide a framework for elucidating the pathogenic mechanism in the subset of obese patients with low endogenous leptin levels.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request. For the CoLaus study, owing to restrictions by the cantonal ethical committee (CER-VD) no individual data for the CoLaus study can be made publicly available. However, such data can be requested for research purposes from the scientific committee of the CoLaus study (peter.vollenweider@chuv.ch). For the Inter99 study, the source data are available from authors at the Novo Nordisk Foundation Center for Basic Metabolic Research, who may be contacted at tuomas.kilpelainen@sund.ku.dk.
References
Haslam, D. W. & James, W. P. Obesity. Lancet 366, 1197–1209 (2005).
Maffei, M. et al. Increased expression in adipocytes of ob RNA in mice with lesions of the hypothalamus and with mutations at the db locus. Proc. Natl. Acad. Sci. USA 92, 6957–6960 (1995).
Friedman, J. M. Leptin at 14 y of age: an ongoing story. Am. J. Clin. Nutr. 89, 973S–979S (2009).
Ioffe, E., Moon, B., Connolly, E. & Friedman, J. M. Abnormal regulation of the leptin gene in the pathogenesis of obesity. Proc. Natl Acad. Sci. USA 95, 11852–11857 (1998).
Farooqi, I. S. et al. Partial leptin deficiency and human adiposity. Nature 414, 34–35 (2001).
Maffei, M. et al. Leptin levels in human and rodent: measurement of plasma leptin and ob RNA in obese and weight-reduced subjects. Nat. Med. 1, 1155–1161 (1995).
Considine, R. V. et al. Serum immunoreactive-leptin concentrations in normal-weight and obese humans. N. Engl. J. Med. 334, 292–295 (1996).
Mandrup, S., Loftus, T. M., MacDougald, O. A., Kuhajda, F. P. & Lane, M. D. Obese gene expression at in vivo levels by fat pads derived from s.c. implanted 3T3-F442A preadipocytes. Proc. Natl Acad. Sci. USA 94, 4300–4305 (1997).
de la Brousse, F. C., Shan, B. & Chen, J. L. Identification of the promoter of the mouse obese gene. Proc. Natl Acad. Sci. USA 93, 4096–4101 (1996).
Mason, M. M., He, Y., Chen, H., Quon, M. J. & Reitman, M. Regulation of leptin promoter function by Sp1, C/EBP, and a novel factor. Endocrinology 139, 1013–1022 (1998).
Pennacchio, L. A. et al. In vivo enhancer analysis of human conserved non-coding sequences. Nature 444, 499–502 (2006).
Levine, M., Cattoglio, C. & Tjian, R. Looping back to leap forward: transcription enters a new era. Cell 157, 13–25 (2014).
Miura, S. et al. Nuclear factor 1 regulates adipose tissue–specific expression in the mouse GLUT4 gene. Biochem. Biophys. Res. Commun. 325, 812–818 (2004).
Waki, H. et al. Global mapping of cell type–specific open chromatin by FAIRE-seq reveals the regulatory role of the NFI family in adipocyte differentiation. PLoS Genet. 7, e1002311 (2011).
Sun, X., Haider Ali, M. S. S. & Moran, M. The role of interactions of long non-coding RNAs and heterogeneous nuclear ribonucleoproteins in regulating cellular functions. Biochem. J. 474, 2925–2935 (2017).
Zhao, X. Y., Li, S., Wang, G. X., Yu, Q. & Lin, J. D. A long noncoding RNA transcriptional regulatory circuit drives thermogenic adipocyte differentiation. Mol. Cell 55, 372–382 (2014).
Carpenter, S. et al. A long noncoding RNA mediates both activation and repression of immune response genes. Science 341, 789–792 (2013).
Sun, L. et al. Long noncoding RNAs regulate adipogenesis. Proc. Natl Acad. Sci. USA 110, 3387–3392 (2013).
Ulitsky, I., Shkumatava, A., Jan, C. H., Sive, H. & Bartel, D. P. Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell 147, 1537–1550 (2011).
Mercer, T. R., Dinger, M. E. & Mattick, J. S. Long non-coding RNAs: insights into functions. Nat. Rev. Genet. 10, 155–159 (2009).
Lam, M. T., Li, W., Rosenfeld, M. G. & Glass, C. K. Enhancer RNAs and regulated transcriptional programs. Trends Biochem. Sci. 39, 170–182 (2014).
Step, S. E. et al. Anti-diabetic rosiglitazone remodels the adipocyte transcriptome by redistributing transcription to PPARγ-driven enhancers. Genes Dev. 28, 1018–1028 (2014).
Kilpelainen, T. O. et al. Genome-wide meta-analysis uncovers novel loci influencing circulating leptin levels. Nat. Commun. 7, 10494 (2016).
Loh, P. R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Halaas, J. L. et al. Physiological response to long-term peripheral and central leptin infusion in lean and obese mice. Proc. Natl Acad. Sci. USA 94, 8878–8883 (1997).
Ottaway, N. et al. Diet-induced obese mice retain endogenous leptin action. Cell Metab. 21, 877–882 (2015).
Knight, Z. A., Hannan, K. S., Greenberg, M. L. & Friedman, J. M. Hyperleptinemia is required for the development of leptin resistance. PLoS One 5, e11376 (2010).
Lu, Y. H., Dallner, O. S., Birsoy, K., Fayzikhodjaeva, G. & Friedman, J. M. Nuclear factor-Y is an adipogenic factor that regulates leptin gene expression. Mol. Metab. 4, 392–405 (2015).
Sallam, T. et al. Transcriptional regulation of macrophage cholesterol efflux and atherogenesis by a long noncoding RNA. Nat. Med. 24, 304–312 (2018).
Gong, S. et al. A gene expression atlas of the central nervous system based on bacterial artificial chromosomes. Nature 425, 917–925 (2003).
Hogan, B., Beddington, R., Constantini, F. & Lacy, E. Manipulating the Mouse Embryo 2nd edn (Cold Spring Harbor Press, 1994).
Cope, N. F. & Fraser, P. Chromosome conformation capture. Cold Spring Harb. Protoc. 2009, pdbprot5137 (2009).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Hong, S. et al. Dissociation of muscle insulin sensitivity from exercise endurance in mice by HDAC3 depletion. Nat. Med. 23, 223–234 (2017).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Emmett, M. J. et al. Histone deacetylase 3 prepares brown adipose tissue for acute thermogenic challenge. Nature 546, 544–548 (2017).
Firmann, M. et al. The CoLaus study: a population-based study to investigate the epidemiology and genetic determinants of cardiovascular risk factors and metabolic syndrome. BMC Cardiovasc. Disord. 8, 6 (2008).
Aune, D., Norat, T., Leitzmann, M., Tonstad, S. & Vatten, L. J. Physical activity and the risk of type 2 diabetes: a systematic review and dose–response meta-analysis. Eur. J. Epidemiol. 30, 529–542 (2015).
Jorgensen, T. et al. A randomized non-pharmacological intervention study for prevention of ischaemic heart disease: baseline results Inter99. Eur. J. Cardiovasc. Prev. Rehabil. 10, 377–386 (2003).
Elliott, P., Peakman, T. C. & Biobank, U. K. The UK Biobank sample handling and storage protocol for the collection, processing and archiving of human blood and urine. Int. J. Epidemiol. 37, 234–244 (2008).
Peakman, T. C. & Elliott, P. The UK Biobank sample handling and storage validation studies. Int. J. Epidemiol. 37(Suppl. 1), i2–i6 (2008).
Downey, P. & Peakman, T. C. Design and implementation of a high-throughput biological sample processing facility using modern manufacturing principles. Int. J. Epidemiol. 37(Suppl. 1), i46–i50 (2008).
Collins, R. What makes UK Biobank special? Lancet 379, 1173–1174 (2012).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Acknowledgements
We thank S. Malik and R. G. Roeder for valuable discussions. We acknowledge technical support from S. Schmidt and R. Sarnoff. We also thank the Gene Targeting Core at Rockefeller University for assistance with CRISPR targeting. O.S.D. acknowledges support from the Swedish Research Council, the Swedish Medical Research Society, and the Sweden-America Foundation. This work was supported by NIH grant R01-DK49780 (M.A.L.), the JPB Foundation (M.A.L.), and CEN 5402133 (J.M.F.). The study was supported in part by the NCATS CTSA program (UL1-TR001866) (R.V.). The Sohn Conference Foundation and the Leona M. and Harry B. Helmsley Charitable Trust are acknowledged for mass spectrometry instrumentation. The Novo Nordisk Foundation Center for Basic Metabolic Research is an independent Research Center at the University of Copenhagen partially funded by an unrestricted donation from the Novo Nordisk Foundation (https://www.metabol.ku.dk).
Author information
Authors and Affiliations
Contributions
O.S.D. performed the majority of the experiments. O.S.D. planned experiments and drafted the manuscript with J.M.F. J.M.M. performed experiments (LNA and GRO-seq). Additional experiments were performed by Y.L. and K.B. E.W. provided support for animal husbandry and some mouse experiments. G.F. performed pronuclear injections for BACTGs and CRISPR, and B.D.D. and H.M. performed mass spectrometry and subsequent analysis. Z.K. and P.M.-V. contributed the CoLaus data, and T.O.K., N.G., and A.L. provided the Inter99 data. Y.Z. provided some experimental advice. R.V. performed a major part of the statistical analysis. A.M. and R.J.F.L. performed the CoLaus and Inter99 analysis. M.A.L. conceived part of the study and supervised the LNA and GRO-seq experiments. J.M.F. conceived the majority of the study and drafted the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Initial BACTG reporter lines and additional data.
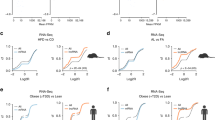
a, The first two BACTG lines covering 304 kb were imaged with IVIS comparing fasted or fed mice or mice crossed to Lepob/Lepob mice. Tissues were analyzed for luciferase reporter activity (−156 kb to +18 kb, n = 4; −22 kb to +148 kb, n = 4). Sp, spleen; St, stomach; Br, brain; Li, liver; Ki, kidney; In, intestine; eW, eWAT; iW, iWAT. b, Additional luciferase biochemistry for Fig. 1a,d for mice fasted for 48 h, fed, or crossed to Lepob/Lepob mice for the −16.5 to +8.8 kb BACTG (fasted, n = 7; fed, n = 6; ob, n = 6) and the −762 bp to +13.9 kb BACTG (fasted, n = 14; fed, n = 13; ob iWAT, n = 4; ob eWAT, n = 6. Values are means ± s.e.m.
Extended Data Fig. 2 Overview of BACTGs.
An overview of the BACTG deletion series generated indicating the length of the reporter gene and the measured luciferase activity for mice crossed to Lepob/Lepob mice, fed, or fasted for 48 h. ob eWAT/iWAT: −22 to +8.8 (n = 3), −17.4 to +8.8 (n = 7), −16.5 to +8.8 (n = 10), +762 to +13.9 (n = 3), +762 to +14.2 (n = 4), +762 to +14.8 (n = 3), +762 to +15.8 (n = 6), +762 to +18 (n = 2 or 3); fed eWAT/iWAT: −22 to +8.8 (n = 8), −17.4 to +8.8 (n = 13), −16.5 to +8.8 (n = 12), −16.4 to +8.8 (n = 2), −16.1 to +8.8 (n = 11), −762 to +8.8 (n = 7), +762 to +13.6 (n = 14), +762 to +13.9 (n = 3), +762 to +14.2 (n = 9), +762 to +14.8 (n = 3), +762 to +15.8 (n = 3), +762 to +18 (n = 5 or); fasted eWAT/iWAT: −22 to +8.8 (n = 6), −17.4 to +8.8 (n = 4), −16.5 to +8.8 (n = 11), +762 to +13.9 (n = 2), +762 to +14.2 (n = 11), +762 to +14.8 (n = 3), +762 to +15.8 (n = 3), +762 to +18 (n = 3). Values are means ± s.e.m.
Extended Data Fig. 3 Predicted interactions of identified proteins.
String interaction prediction was performed (https://string-db.org/) indicating interactions between RNA-binding proteins (red) and DNA-binding proteins (blue).
Extended Data Fig. 4 RNA expression of leptin and lncOb in human subcutaneous adipose tissue.
RNA levels of leptin and lncOb were measured by TaqMan qPCR in human samples (Sm, skeletal muscle; Li, liver; Lu, lung; AdiMix, mixed human fat samples; BMI25–30, three separate samples of human subcutaneous adipose tissue with known BMI). n = 7. Data were analyzed by linear regression.
Extended Data Fig. 5 A mutation in lncOb causes dysregulation of quantitative leptin mRNA expression in mice.
Leptin mRNA levels were measured in subcutaneous adipose tissue with a TaqMan assay and are plotted against fat mass (grams) for HFD-fed mice (WT, n = 10; lncOb–/–, n = 9) (squares, males; circles, females). Data were analyzed by linear regression.
Extended Data Fig. 6 A mutation in lncOb causes dysregulation of quantitative leptin expression (regular diet).
a, Seven-week-old male (WT, n = 9; lncOb–/–, n = 7) and female (WT, n = 6; lncOb–/–, n = 9) mice were fed a regular diet for 20 weeks and body weight was monitored (shaded regions represent means ± s.e.m). b, Fat mass was measured by EchoMRI in male (WT, n = 9; lncOb–/–, n = 7) and female (WT, n = 6; lncOb–/–, n = 9) mice after 20 weeks of a regular diet. Values are means ± s.e.m. P values were calculated by two-tailed Student’s t-test. c, Leptin levels (ng/ml) were measured by ELISA and are plotted against fat mass (grams) for regular diet (RD)-fed mice (WT, n = 13; lncOb–/–, n = 14) (squares, males; circles, females). For a data were analyzed by a linear mixed model with an AR(1) correlation structure, and for c data were analyzed by linear regression.
Extended Data Fig. 7 Regional plots for the leptin locus.
a,b, Regional plots for ‘comparative body size at age 10 years’ in UK Biobank (n ≃ 450,000) (a) and BMI-adjusted leptin levels from Kilpelainen et al.23 at the LEP locus (n > 46,000) (b). P values were calculated using a linear regression model.
Extended Data Fig. 8 rs10487505 is a significant eQTL in the GTEx portal.
a,b, The GTEx portal (https://gtexportal.org/) was queried for rs10487505, indicating a single eQTL (a) that impacts expression of IMPDH1 in thyroid (b) (n = 103 reference homozygotes, n = 204 heterozygotes, n = 92 alternative homozygotes). Box plots depict the interquartile range (IQR) and mean, and whiskers depict minimum and maximum values (1.5 times IQR). P value was calculated with a linear regression model. The normalized effect size (NES) of the eQTL is defined as the slope of the linear regression.
Supplementary information
Supplementary Information
Supplementary Tables 1–6
Supplementary Table 7
Proteins identified by mass spectrometry in Fig. 3a
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 2
Unprocessed gel shift images
Source Data Extended Data Fig. 4
Statistical Source Data
Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 6
Statistical Source Data
Rights and permissions
About this article
Cite this article
Dallner, O.S., Marinis, J.M., Lu, YH. et al. Dysregulation of a long noncoding RNA reduces leptin leading to a leptin-responsive form of obesity. Nat Med 25, 507–516 (2019). https://doi.org/10.1038/s41591-019-0370-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0370-1
This article is cited by
-
The interaction between MALAT1 and TUG1 with dietary fatty acid quality indices on visceral adiposity index and body adiposity index
Scientific Reports (2024)
-
Transcriptional and epigenetic changes after dietary and surgical weight loss interventions in an animal model of obesity
International Journal of Obesity (2023)
-
Identification and characterization of the long non-coding RNA NFIA-AS2 as a novel locus for body mass index in American Indians
International Journal of Obesity (2023)
-
Downregulation of hepatic lncRNA Gm19619 improves gluconeogenesis and lipogenesis following vertical sleeve gastrectomy in mice
Communications Biology (2023)
-
The long noncoding RNA ADIPINT regulates human adipocyte metabolism via pyruvate carboxylase
Nature Communications (2022)