Abstract
Leber congenital amaurosis type 10 is a severe retinal dystrophy caused by mutations in the CEP290 gene1,2. We developed EDIT-101, a candidate genome-editing therapeutic, to remove the aberrant splice donor created by the IVS26 mutation in the CEP290 gene and restore normal CEP290 expression. Key to this therapeutic, we identified a pair of Staphylococcus aureus Cas9 guide RNAs that were highly active and specific to the human CEP290 target sequence. In vitro experiments in human cells and retinal explants demonstrated the molecular mechanism of action and nuclease specificity. Subretinal delivery of EDIT-101 in humanized CEP290 mice showed rapid and sustained CEP290 gene editing. A comparable surrogate non-human primate (NHP) vector also achieved productive editing of the NHP CEP290 gene at levels that met the target therapeutic threshold, and demonstrated the ability of CRISPR/Cas9 to edit somatic primate cells in vivo. These results support further development of EDIT-101 for LCA10 and additional CRISPR-based medicines for other inherited retinal disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Code availability
Custom analysis software is freely available at the following websites: https://github.com/editasmedicine/uditas https://github.com/editasmedicine/digenomitas
Data availability
Detailed data analysis is available in the Supplementary Tables published with this manuscript. All requests for raw data are promptly reviewed by the CSO or CTO of Editas Medicine, and any data that can be shared will be released tothe requestor. WGS data are available via the NCBI SRA database under accession code PRJNA497602.
References
Chang, B. et al. In-frame deletion in a novel centrosomal/ciliary protein CEP290/NPHP6 perturbs its interaction with RPGR and results in early-onset retinal degeneration in the rd16 mouse. Hum. Mol. Genet. 15, 1847–1857 (2006).
Drivas, T. G. & Bennett, J. CEP290 and the primary cilium. Adv. Exp. Med. Biol. 801, 519–525 (2014).
Stone, E. M. Leber congenital amaurosis - a model for efficient genetic testing of heterogeneous disorders: LXIV Edward Jackson Memorial Lecture. Am. J. Ophthalmol. 144, 791–811 (2007).
Stone, E. M. et al. Clinically focused molecular investigation of 1000 consecutive families with inherited retinal disease. Ophthalmology 124, 1314–1331 (2017).
Drivas, T. G., Wojno, A. P., Tucker, B. A., Stone, E. M. & Bennett, J. Basal exon skipping and genetic pleiotropy: a predictive model of disease pathogenesis. Sci. Transl. Med. 7, 291ra297 (2015).
den Hollander, A. I. et al. Mutations in the CEP290 (NPHP6) gene are a frequent cause of Leber congenital amaurosis. Am. J. Hum. Genet. 79, 556–561 (2006).
Collin, R. W. et al. Antisense oligonucleotide (AON)-based therapy for Leber congenital amaurosis caused by a frequent mutation in CEP290. Mol. Ther. Nucleic Acids 1, e14 (2012).
Gerard, X. et al. AON-mediated exon skipping restores ciliation in fibroblasts harboring the common Leber congenital amaurosis CEP290 mutation. Mol. Ther. Nucleic Acids 1, e29 (2012).
Parfitt, D. A. et al. Identification and correction of mechanisms underlying inherited blindness in human iPSC-derived optic cups. Cell Stem Cell 18, 769–781 (2016).
Coppieters, F., Lefever, S., Leroy, B. P. & De Baere, E. CEP290, a gene with many faces: mutation overview and presentation of CEP290base. Hum. Mutat. 31, 1097–1108 (2010).
Boye, S. E. et al. Natural history of cone disease in the murine model of Leber congenital amaurosis due to CEP290 mutation: determining the timing and expectation of therapy. PLoS ONE 9, e92928 (2014).
Geller, A. M. & Sieving, P. A. Assessment of foveal cone photoreceptors in Stargardt’s macular dystrophy using a small dot detection task. Vision Res. 33, 1509–1524 (1993).
Geller, A. M., Sieving, P. A. & Green, D. G. Effect on grating identification of sampling with degenerate arrays. J. Opt. Soc. Am. A. 9, 472–477 (1992).
Garanto, A. et al. Unexpected CEP290 mRNA splicing in a humanized knock-in mouse model for Leber congenital amaurosis. PLoS ONE 8, e79369 (2013).
Cideciyan, A. V. et al. Centrosomal-ciliary gene CEP290/NPHP6 mutations result in blindness with unexpected sparing of photoreceptors and visual brain: implications for therapy of Leber congenital amaurosis. Hum. Mutat. 28, 1074–1083 (2007).
Cideciyan, A. V. et al. Cone photoreceptors are the main targets for gene therapy of NPHP5 (IQCB1) or NPHP6 (CEP290) blindness: generation of an all-cone Nphp6 hypomorph mouse that mimics the human retinal ciliopathy. Hum. Mol. Genet. 20, 1411–1423 (2011).
Boye, S. E. et al. The human rhodopsin kinase promoter in an AAV5 vector confers rod- and cone-specific expression in the primate retina. Hum. Gene Ther. 23, 1101–1115 (2012).
LUXTURNA (voretigene neparvovec-rzyl, package insert) (Spark Therapeutics, 2017).
Giannoukos, G. et al. UDiTaS, a genome editing detection method for indels and genome rearrangements. BMC Genomics 19, 212 (2018).
Burnight, E. R. et al. CEP290 gene transfer rescues Leber congenital amaurosis cellular phenotype. Gene Ther. 21, 662–672 (2014).
Feodorova, Y., Koch, M., Bultman, S., Michalakis, S. & Solovei, I. Quick and reliable method for retina dissociation and separation of rod photoreceptor perikarya from adult mice. MethodsX 2, 39–46 (2015).
Niesen, F. H., Berglund, H. & Vedadi, M. The use of differential scanning fluorimetry to detect ligand interactions that promote protein stability. Nat. Protoc. 2, 2212–2221 (2007).
Kim, D. et al. Digenome-seq: genome-wide profiling of CRISPR-Cas9 off-target effects in human cells. Nat. Methods 12, 237–243 (2015).
Park, J. et al. Digenome-seq web tool for profiling CRISPR specificity. Nat. Methods 14, 548–549 (2017).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Friedland, A. E. et al. Characterization of Staphylococcus aureus Cas9: a smaller Cas9 for all-in-one adeno-associated virus delivery and paired nickase applications. Genome Biol. 16, 257 (2015).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBO J. 17, 10–12 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Bothmer, A. et al. Characterization of the interplay between DNA repair and CRISPR/Cas9-induced DNA lesions at an endogenous locus. Nat. Commun. 8, 13905 (2017).
Kleinstiver, B. P. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Bae, S., Park, J. & Kim, J. S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475 (2014).
Acknowledgements
We thank B. Tucker and colleagues at the University of Iowa for the LCA10 patient fibroblasts, and the patients for their generous donation of biopsy samples. We thank B. Rabe for advice and technological assistance in establishing retinal explant model. We thank M. McCartney and N. Sprehe from Lions Eye Institute for Transplant and Research for invaluable help and services with human retinal explant experiments, and we thank the donors and their families for their generous donation. We thank our Editas colleagues P. Baciu, D. Balderson and G. Cox for helpful discussions and scientific input, and K. LeClair, W. Jaworowicz and G. Wilmes for helpful discussions and for manufacturing expertise and materials supply. This work was fully funded by Editas Medicine.
Author information
Authors and Affiliations
Contributions
M.L.M., M.S., C.J.W., D.B., A.G., P.S., V.E.M., C.F.A. and H.J. conceived the strategy and designed experiments. M.L.M., R.B., A.D., A.E.F., S.W.G., J.E.H., S.J., R.M., D.R., S.S. and M.N.S. designed, performed and analyzed in vitro experiments. M.L.M., M.S., G.S.B., H.C., S.S., C.Y. and H.J. designed, performed and analyzed in vivo experiments. S.J.H., C.F.A. and H.J. designed, performed and analyzed immunogenicity experiments. M.L.M., C.J.W., L.A.B., D.M.C., J.A.D., V.D., T.J.F., G.G., G.M.G., H.J., E.M., T.T., D.G.T., T.W. and V.E.M. designed, performed and analyzed specificity studies. B.H. and S.J.S. performed and analyzed histology studies. M.L.M., M.S., C.J.W., C.F.A. and H.J. wrote, and all authors reviewed, the manuscript.
Corresponding author
Ethics declarations
Competing interests
C.J.W., R.B., G.S.B., D.M.C., J.A.D., A.D., V.D., A.E.F., G.G., S.W.G., G.M.G., B.H., S.J., E.M., D.R., T.T., D.G.T., S.J.S., S.S., P.S., T.W., C.Y., V.E.M. and C.F.A. are current employees and shareholders of Editas Medicine. M.L.M., M.S., H.C., L.A.B., D.B., A.G., S.J.H., H.J., J.E.H., R.M., M.N.S. and H.J. are former employees and shareholders of Editas Medicine and were employeed by Editas at the time this work was conducted. T.J.F. is a paid consultant of Editas Medicine. Editas Medicine has filed patents pertaining to the work described in this manuscript.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Splicing reporter assay demonstrates functionality of deletions and inversions.
a, Schematic depicting design of GFP reporter constructs to determine functionality of edits and ability to correct splicing. Cytomegalovirus promoter drives split GFP interrupted by a portion of CEP290 intron 26. Correct splicing is required to reconstitute GFP. b, Quantification of GFP expression (mean fluorescence intensity), normalized to PGK-mCherry also present on reporter plasmid, by flow cytometry following transfection into human U2OS cells. n = 3 independent transfections performed on separate days. Bars represent mean and standard deviation.
Extended Data Fig. 2 In vitro validation of editing strategy in primary patient fibroblasts.
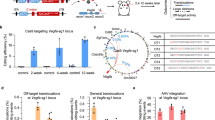
a, Targeted deletion at the CEP290 locus in primary patient fibroblasts transfected with plasmids encoding SaCas9 and seven different pairs of CEP290 gRNAs, as quantified by Droplet Digital PCRR. Left, cell line IVS26#36, n = 4 independent transfections performed on different days, error bars represent standard deviation. Right, cell line IVS26#35, n = 1. b, Quantification of WT 26-27 (blue) and mutant 26-X-27 (orange) CEP290 mRNA transcripts in IVS26#35 primary patient fibroblasts by qRT–PCR. CEP290 expression is normalized to beta-actin. n = 2 independent transfections performed on different days, qRT–PCR run in triplicate. Line represents mean. c, representative western blot from IVS26 primary patient fibroblasts transfected with plasmids encoding SaCas9 and seven different pairs of CEP290 gRNAs. Experiment was performed twice in each cell line with similar results. d, Quantification of CEP290 full-length protein expression in IVS26#36 and -#35 cell lines transfected with plasmids encoding SaCas9 and seven different pairs of CEP290 gRNAs. Expression level fold change over control calculated by densitometry of western blot bands and normalized to control. n = 4 biological replicates (two different transfections on different days, each in the two different cell lines), error bars represent mean and standard deviation.
Extended Data Fig. 3 Specificity profiling of CEP290 gRNAs 323 and 64.
a, Digenome-Seq was used to identify cuts across the genome. The positive control, SpCas9-EMX1, showed dose responsiveness with an increased number of cut sites as RNP concentration increased. No cut sites, other than the on-target site, were identified for guide 64 and only one cut site was detected at the highest concentration (1,000 nM) for guide 323. That candidate off-target site was located in intron 2 of gene CTD-2058B24.2, an 'anti-sense RNA' with no ascribed function; the cut site is close to, but distinct from, an annotated DNase-hypersensitive region. The raw data associated with this figure are available in Supplementary Tables 6 and 7. b, Targeted amplicon sequencing of candidate off-target sites in cell lines nucleofected with plasmids and human retinal explants transduced with EDIT-101. Off-target sites are plotted on the x axis and grouped into categories (on-target sites, detection below the lowest level of detection, and detection above the lowest level of detection with no change relative to control), then sorted by mean indel percentage. The y axis is a log scale plot of indels detected at the predicted cut site ±2 bases. The raw data associated with this figure are available in Supplementary Table 8. c, Summary table of candidate off-target site-targeted sequencing. See detailed list in Supplementary Table 8.
Extended Data Fig. 4 Transduction and editing efficiency of mouse neural retina by subretinal injection of 1 μl of AAV5 vectors.
a, A representative image of a flat-mounted retina from an HuCEP290 IVS26 KI mouse administered AAV5-GKR1-GFP. The red line outlines the retina, and the GFP-positive area is colored white. b, Total editing rates, as quantified by UDiTaS, in genomic DNA isolated from either total retinal cells or fluorescent-activated cell sorter-isolated GFP-positive retinal cells following subretinal injection of EDIT-101 and AAV5-GRK1-GFP in mice. Each bar represents one mouse eye.
Extended Data Fig. 5 Comparison of human CEP290 gRNAs and NHP surrogate guides.
HuCEP290 IVS26 KI mice were subretinally injected with 1 μl of either EDIT-101 (blue) or NHP surrogate vector (VIR067, orange) at varying doses. Total editing was quantified by UDiTaS. Each point represents a single mouse eye and error bars represent mean and standard deviation. There was no significant difference found between EDIT-101 and VIR067 at any dose.
Extended Data Fig. 6 Localization of AAV genomes and SaCas9 protein in NHPs subretinally injected with VIR026.
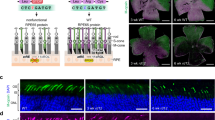
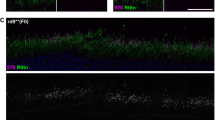
a,b, Detection of AAV5 vector genome in photoreceptor cells of NHP retina from animals treated with either 1 × 1011 vg ml–1 (a, animal no. I16464) or 1 × 1012 vg ml–1 (b, animal no. I16467) and assayed at 13 weeks post-dosing. In situ hybridization with a probe specific for the vector genome shows positive staining enriched in the outer nuclear layer (red, arrow). The area of positive staining was quantified on ×20 stitched tiles. Scale bar, 100 μm. The experiment was performed on six retinas from six different animals treated with VIR026. c,d, Anti-Cas9 immunohistochemistry in monkey no. I6467 showing positive staining in the photoreceptor nuclear layer (arrows) within the bleb region (c) but not outside of the bleb on the opposite side of the optic nerve (d). ×20 stitched tiles. Scale bar, 50 μm. The experiment was performed on six retinas from six different animals treated with VIR026. e, Detection of AAV5 vector genome in photoreceptor cells of NHP 1012 OD showing positive staining encompassing the foveal area (red). ×20 stitched tiles. OD: right eye.
Extended Data Fig. 7 Immunogenicity assessment of AAV5-CRISPR/Cas9-based in vivo genome editing in NHPs.
a,b, Antibodies against AAV5 capsid protein (a) and Cas9 protein (b) were measured in sera from the study animals using a Luminex bead-based assay. Results are presented as concentration of IgG against AAV5 and Cas9 in each individual animal’s serum at each time point. All samples were run in triplicate. c,d, ELISpots were performed to measure Cas9-specific CD8 T-cell responses (c) and Cas9-specific CD4 T-cell responses (d). Peripheral blood mononuclear cells from individual animals were stimulated with Cas9 peptide pools and assayed for IFN-γ production. Data are presented as the number of IFN-γ spot-forming cells per million cells.
Extended Data Fig. 8 Inhibition of anti-Cas9 and anti-AAV5 antibody binding with excess antigen.
a,b, To confirm antibody specificity, animal serum was pre-incubated with excess AAV5 capsid protein, 1 × 1013 viral particles ml–1 (a) or excess Cas9 protein, 160 µg ml–1 (b). Reduction in antibody binding, denoted by a decrease in median fluorescence intensity, was measured using the Luminex bead platform.
Extended Data Fig. 9 Ocular tolerability.
Scoring of anterior and posterior changes based on modifications of the Standardization of Uveitis Nomenclature, Hackett–McDonald and Semi-quantitative Preclinical Ocular Toxicology Scoring systems.
Extended Data Fig. 10 Representative fundus image.
Representative fundus images from animal no. 1005 (treated OD with EDIT-101 at 1 × 1012 vg ml–1 and OS with vehicle) at pre-test, post-dosing and days 8 and 59, showing submacular bleb formation. Fundus imaging was performed with similar results for all 16 NHPs reported in this study, and is detailed in Supplementary Table 9. OD: right eye; OS: left eye.
Supplementary Information
Supplementary Information
Supplementary Results
Supplementary Tables
Supplementary Tables 1–12
Source data
Extended Data Fig. 2
The unprocessed gel image for the CEP290 portion of Extended Data Figure 2c
Extended Data Fig. 2
The unprocessed gel image for the GAPDH portion of Extended Data Figure 2c
Rights and permissions
About this article
Cite this article
Maeder, M.L., Stefanidakis, M., Wilson, C.J. et al. Development of a gene-editing approach to restore vision loss in Leber congenital amaurosis type 10. Nat Med 25, 229–233 (2019). https://doi.org/10.1038/s41591-018-0327-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0327-9
This article is cited by
-
Decoding the complexity of on-target integration: characterizing DNA insertions at the CRISPR-Cas9 targeted locus using nanopore sequencing
BMC Genomics (2024)
-
Between hope and reality: treatment of genetic diseases through nucleic acid-based drugs
Communications Biology (2024)
-
Adeno-associated virus as a delivery vector for gene therapy of human diseases
Signal Transduction and Targeted Therapy (2024)
-
Targeted Gene Insertion: The Cutting Edge of CRISPR Drug Development with Hemophilia as a Highlight
BioDrugs (2024)
-
In vivo LNP-CRISPR Approaches for the Treatment of Hemophilia
Molecular Diagnosis & Therapy (2024)