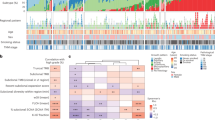
Abstract
The molecular alterations that occur in cells before cancer is manifest are largely uncharted. Lung carcinoma in situ (CIS) lesions are the pre-invasive precursor to squamous cell carcinoma. Although microscopically identical, their future is in equipoise, with half progressing to invasive cancer and half regressing or remaining static. The cellular basis of this clinical observation is unknown. Here, we profile the genomic, transcriptomic, and epigenomic landscape of CIS in a unique patient cohort with longitudinally monitored pre-invasive disease. Predictive modeling identifies which lesions will progress with remarkable accuracy. We identify progression-specific methylation changes on a background of widespread heterogeneity, alongside a strong chromosomal instability signature. We observed mutations and copy number changes characteristic of cancer and chart their emergence, offering a window into early carcinogenesis. We anticipate that this new understanding of cancer precursor biology will improve early detection, reduce overtreatment, and foster preventative therapies targeting early clonal events in lung cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Code availability
All code used in our analysis will be made available at http://github.com/ucl-respiratory/preinvasive on publication. All software dependencies, full version information, and parameters used in our analysis can be found here.Unless otherwise specified, all analyses were performed in an R statistical environment (v3.5.0; www.r-project.org/) using Bioconductor32 version 3.7.
Data availability
Whole-genome sequencing data have been deposited at the European Genome Phenome Archive (https://www.ebi.ac.uk/ega/ at the EBI) with accession number EGAD00001003883. All gene expression and methylation microarray data reported in this study have been deposited in the National Center for Biotechnology Information Gene Expression Omnibus (GEO, http://www.ncbi.nlm.nih.gov/geo/) public repository, and they are accessible through GEO accession number GSE108124.
References
Parkin, D. M., Bray, F., Ferlay, J. & Pisani, P. Global cancer statistics, 2002. CA Cancer J. Clin. 55, 74–108 (2005).
Torre, L. A., Siegel, R. L. & Jemal, A. Lung cancer statistics. Adv. Exp. Med. Biol. 893, 1–19 (2016).
Nicholson, A. G. et al. Reproducibility of the WHO/IASLC grading system for pre-invasive squamous lesions of the bronchus: a study of inter-observer and intra-observer variation. Histopathology 38, 202–208 (2001).
van der Heijden, E. H., Hoefsloot, W., van Hees, H. W. & Schuurbiers, O. C. High definition bronchoscopy: a randomized exploratory study of diagnostic value compared to standard white light bronchoscopy and autofluorescence bronchoscopy. Respir. Res. 16, 33 (2015).
Thakrar, R. M., Pennycuick, A., Borg, E. & Janes, S. M. Pre-invasive disease of the airway. Cancer Treat. Rev. 58, 77–90 (2017).
Pipinikas, C. P. et al. Cell migration leads to spatially distinct but clonally related airway cancer precursors. Thorax 69, 548–557 (2014).
Jeremy George, P. et al. Surveillance for the detection of early lung cancer in patients with bronchial dysplasia. Thorax 62, 43–50 (2007).
Futreal, P. A. et al. A census of human cancer genes. Nat. Rev. Cancer. 4, 177–183 (2004).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Alexandrov, L. B. & Stratton, M. R. Mutational signatures: the patterns of somatic mutations hidden in cancer genomes. Curr. Opin. Genet. Dev. 24, 52–60 (2014).
Cancer Genome Atlas Research, N. Comprehensive genomic characterization of squamous cell lung cancers. Nature 489, 519–525 (2012).
Jiang, F., Yin, Z., Caraway, N. P., Li, R. & Katz, R. L. Genomic profiles in stage I primary non small cell lung cancer using comparative genomic hybridization analysis of cDNA microarrays. Neoplasia 6, 623–635 (2004).
Chujo, M. et al. Comparative genomic hybridization analysis detected frequent overrepresentation of chromosome 3q in squamous cell carcinoma of the lung. Lung Cancer 38, 23–29 (2002).
Tonon, G. et al. High-resolution genomic profiles of human lung cancer. Proc. Natl Acad. Sci. USA 102, 9625–9630 (2005).
Petersen, I. et al. Patterns of chromosomal imbalances in adenocarcinoma and squamous cell carcinoma of the lung. Cancer Res. 57, 2331–2335 (1997).
Balsara, B. R. & Testa, J. R. Chromosomal imbalances in human lung cancer. Oncogene 21, 6877–6883 (2002).
Massion, P. P. et al. Genomic copy number analysis of non-small cell lung cancer using array comparative genomic hybridization: implications of the phosphatidylinositol 3-kinase pathway. Cancer Res. 62, 3636–3640 (2002).
Ried, T. et al. Mapping of multiple DNA gains and losses in primary small cell lung carcinomas by comparative genomic hybridization. Cancer Res. 54, 1801–1806 (1994).
Rodrigues, M. F., Esteves, C. M., Xavier, F. C. & Nunes, F. D. Methylation status of homeobox genes in common human cancers. Genomics 108, 185–193 (2016).
Matsubara, D. et al. Inactivating mutations and hypermethylation of the NKX2-1/TTF-1 gene in non-terminal respiratory unit-type lung adenocarcinomas. Cancer Sci. 108, 1888–1896 (2017).
Winslow, M. M. et al. Suppression of lung adenocarcinoma progression by Nkx2-1. Nature 473, 101–104 (2011).
Tata, P. R. et al. Developmental history provides a roadmap for the emergence of tumor plasticity. Dev. Cell 44, 679–693.e675 (2018).
van Boerdonk, R. A. et al. DNA copy number aberrations in endobronchial lesions: a validated predictor for cancer. Thorax 69, 451–457 (2014).
Lee, K., Kim, J. H. & Kwon, H. The actin-related protein baf53 is essential for chromosomal subdomain integrity. Mol. Cell 38, 789–795 (2015).
Carter, S. L., Eklund, A. C., Kohane, I. S., Harris, L. N. & Szallasi, Z. A signature of chromosomal instability inferred from gene expression profiles predicts clinical outcome in multiple human cancers. Nat. Genet. 38, 1043–1048 (2006).
Luo, W., Friedman, M. S., Shedden, K., Hankenson, K. D. & Woolf, P. J. GAGE: generally applicable gene set enrichment for pathway analysis. BMC Bioinformatics 10, 161 (2009).
Endesfelder, D. et al. Chromosomal instability selects gene copy-number variants encoding core regulators of proliferation in ER+ breast cancer. Cancer Res. 74, 4853–4863 (2014).
Blackburn, A. et al. Effects of copy number variable regions on local gene expression in white blood cells of Mexican Americans. Eur. J. Hum. Genet. 23, 1229–1235 (2015).
Mileyko, Y., Joh, R. I. & Weitz, J. S. Small-scale copy number variation and large-scale changes in gene expression. Proc. Natl Acad. Sci. USA 105, 16659–16664 (2008).
McGranahan, N., Burrell, R. A., Endesfelder, D., Novelli, M. R. & Swanton, C. Cancer chromosomal instability: therapeutic and diagnostic challenges. EMBO Rep. 13, 528–538 (2012).
Jamal-Hanjani, M. et al. Tracking the evolution of non-small-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
Jeremy George, P. et al. Surveillance for the detection of early lung cancer in patients with bronchial dysplasia. Thorax 62, 43–50 (2007).
Bibikova, M. et al. High density DNA methylation array with single CpG site resolution. Genomics 98, 288–295 (2011).
Sandoval, J. et al. Validation of a DNA methylation microarray for 450,000 CpG sites in the human genome. Epigenetics 6, 692–702 (2011).
Morris, T. J. et al. ChAMP: 450k chip analysis methylation pipeline. Bioinformatics 30, 428–430 (2014).
Teschendorff, A. E. et al. DNA methylation outliers in normal breast tissue identify field defects that are enriched in cancer. Nat. Commun. 7, 10478 (2016).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Benjamini, Y., Drai, D., Elmer, G., Kafkafi, N. & Golani, I. Controlling the false discovery rate in behavior genetics research. Behav. Brain Res 125, 279–284 (2001).
Kolde, R. Pheatmap: pretty heatmaps. R Package Version 61, 1–7 (2012).
Tibshirani, R., Hastie, T., Narasimhan, B. & Chu, G. Diagnosis of multiple cancer types by shrunken centroids of gene expression. Proc. Natl Acad. Sci. USA 99, 6567–6572 (2002).
Robin, X. et al. pROC: an open-source package for R and S + to analyze and compare ROC curves. BMC Bioinformatics 12, 77 (2011).
Keilwagen, J., Grosse, I. & Grau, J. Area under precision-recall curves for weighted and unweighted data. PLoS One. 9, e92209 (2014).
Raine, K. M. et al. ascatNgs: identifying somatically acquired copy-number alterations from whole-genome sequencing data. Curr. Protoc. Bioinformatics. 56, 15 19 11–15 19 17 (2016).
Feber, A. et al. Using high-density DNA methylation arrays to profile copy number alterations. Genome Biol. 15, R30 (2014).
van Boerdonk, R. A. et al. DNA copy number alterations in endobronchial squamous metaplastic lesions predict lung cancer. Am. J. Respir. Crit. Care. Med. 184, 948–956 (2011).
van Boerdonk, R. A. et al. DNA copy number aberrations in endobronchial lesions: a validated predictor for cancer. Thorax 69, 451–457 (2014).
Grossman, R. L. et al. Toward a shared vision for cancer genomic data. N. Engl. J. Med. 375, 1109–1112 (2016).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Luo, W., Friedman, M. S., Shedden, K., Hankenson, K. D. & Woolf, P. J. GAGE: generally applicable gene set enrichment for pathway analysis. BMC Bioinformatics 10, 161 (2009).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361 (2017).
Carter, S. L., Eklund, A. C., Kohane, I. S., Harris, L. N. & Szallasi, Z. A signature of chromosomal instability inferred from gene expression profiles predicts clinical outcome in multiple human cancers. Nat. Genet. 38, 1043–1048 (2006).
Whitfield, M. L. et al. Identification of genes periodically expressed in the human cell cycle and their expression in tumors. Mol. Biol. Cell 13, 1977–2000 (2002).
Jones, D. et al. cgpCaVEManWrapper: simple execution of caveman in order to detect somatic single nucleotide variants in ngs data. Curr. Protoc. Bioinformatics 56, 15.10.11–15.10.18 (2016).
Van Loo, P. et al. Allele-specific copy number analysis of tumors. Proc. Natl Acad. Sci USA 107, 16910–16915 (2010).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Skinner, M. E., Uzilov, A. V., Stein, L. D., Mungall, C. J. & Holmes, I. H. JBrowse: a next-generation genome browser. Genome Res. 19, 1630–1638 (2009).
Raine, K. M. et al. cgpPindel: identifying somatically acquired insertion and deletion events from paired end sequencing. Curr. Protoc. Bioinformatics 52, 15.7.1–15.7.12 (2015).
Ye, K., Schulz, M. H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Papaemmanuil, E. et al. RAG-mediated recombination is the predominant driver of oncogenic rearrangement in ETV6-RUNX1 acute lymphoblastic leukemia. Nat. Genetics 46, 116–125 (2014).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative genomics viewer (igv): high-performance genomics data visualization and exploration. Brief Bioinform. 14, 178–192 (2013).
Endesfelder, D. et al. Chromosomal instability selects gene copy-number variants encoding core regulators of proliferation in ER+ breast cancer. Cancer Res. 74, 4853–4863 (2014).
Forbes, S. A. et al. COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res. 45, D777–D783 (2017).
McLaren, W. et al. The ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041.e1021 (2017).
Miller, C. A. et al. SciClone: inferring clonal architecture and tracking the spatial and temporal patterns of tumor evolution. PLoS Comput. Biol. 10, e1003665 (2014).
McGranahan, N. et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351, 1463–1469 (2016).
Blokzijl, F., Janssen, R., van Boxtel, R. & Cuppen, E. MutationalPatterns: comprehensive genome-wide analysis of mutational processes. Genome Med .10, 33 (2018).
Alexandrov, L. B. et al. Clock-like mutational processes in human somatic cells. Nat. Genet. 47, 1402–1407 (2015).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Martincorena, I. & Campbell, P. J. Somatic mutation in cancer and normal cells. Science 349, 1483–1489 (2015).
Farmery, J. H. R. & Smith, M. L. NIHR BioResource - Rare Diseases & Lynch, A. G. Telomerecat: a ploidy-agnostic method for estimating telomere length from whole genome sequencing data. Sci. Rep. 8, 1300 (2018).
Acknowledgements
We thank all of the patients who participated in this study and K. Pearce, G. Chennel, D. Chambers, P. Mercer and K. Gowers for technical help and proofreading. We thank P. Rabbitts, A. Banerjee and C. Read for their early development of the study. The results published here are in part based on data generated by a TCGA pilot project established by the National Cancer Institute and National Human Genome Research Institute. Information about TCGA and the investigators and institutions that constitute the TCGA research network can be found at http://cancergenome.nih.gov. R.E.H., N.M., P.J.C., and S.M.J. are supported by the Wellcome Trust fellowships. S.M.J. is also supported by the Rosetrees Trust, the Welton Trust, the Garfield Weston Trust, the Stoneygate Trust and UCLH Charitable Foundation. V.T., C.P., R.E.H. and S.M.J. have been funded by the Roy Castle Lung Cancer Foundation. A.P. is funded by a Wellcome Trust clinical PhD training fellowship. H.L.-S. is funded by the Wellcome Trust Sanger Institute non-clinical PhD studentship. C.T. was a CRUK Clinician Scientist. This work was partially undertaken at UCLH/UCL, who received a proportion of funding from the Department of Health’s NIHR Biomedical Research Centre’s funding scheme (S.M.J. and N.N.). R.E.H., N.M., C.S., and S.M.J. are part of the CRUK Lung Cancer Centre of Excellence. A.S., C.S., and S.M.J. are supported by Stand Up to Cancer. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
V.H.T., C.P.P., and A.P. contributed equally to this work. S.M.J., P.J.C., V.H.T., A.P., R.E.H., H.L.-S., and C.P.P. co-wrote the manuscript. S.M.J., P.J.C., C.T., V.H.T., and C.P.P. conceived the study design. S.M.J., P.J.C., C.T., V.H.T., C.P.P., and A.P. designed the study protocols. V.H.T. performed gene expression, qPCR and LCM experiments, and analyzed and integrated clinicopathological data and gene expression data. C.P.P. performed methylation and LCM experiments, and analyzed and integrated clinicopathological data and methylation data. A.P. analyzed and integrated clinicopathological data, WGS data, gene expression data, and methylation data. H.L.-S., A.G.L., and H.F. analyzed WGS data. D.C. and P.N. performed LCM experiments. J.B. analyzed gene expression data. T.J.M., A.K., A.F., C.E.B., and D.S.P. analyzed methylation data. M.F. and A.C. conducted the pathological review. P.J.G., B.C., N.N., G.H., J.M.B., and R.M.T. performed bronchoscopies and collected the CIS and control biopsies. P.F.D. performed histological experiments. R.E.H., R.C.C., N.M., C.S., S.B., and A.S. gave advice and reviewed the manuscript. S.M.J. provided overall study oversight.
Corresponding author
Ethics declarations
Competing interests
A.S. is an employee of Johnson and Johnson. Discoveries within this manuscript have led S.M.J. to lead on Patent Applications 1819453.0 and 1819452.2 filed with the UK Intellectual Property Office through UCL Business PLC.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Experimental workflow.
Flow diagram illustrating which profiling techniques were applied to which samples. Biopsies taken from index CIS lesions were stored as fresh frozen (FF) and formalin-fixed paraffin embedded (FFPE). DNA was extracted from FF biopsies. The first 54 samples studied that had sufficient extracted DNA passing quality control (QC) underwent first methylation profiling, then whole-genome sequencing (WGS) when sufficient remaining DNA was available. Due to the low DNA quantity extracted from some biopsies, the methylation dataset (n = 54) was larger than the WGS data set (n = 29), therefore the subsequent 10 samples underwent WGS directly without methylation profiling. RNA was extracted from FFPE samples and underwent gene expression profiling when RNA passed QC. To ensure validity of our conclusions across orthogonal platforms we used Illumina microarrays to profile a discovery set of 33 samples, then subsequently used Affymetrix microarrays to profile an independent validation set of 18 further samples.
Extended Data Fig. 2 Mutational signatures of CIS lesions.
a–d, The contribution of each of five pre-selected mutational signatures to each lesion is shown. These five mutational signatures, associated with CpG deamination (1), APOBEC (2 and 13), tobacco (4) and unknown aetiology (5), were selected based on an initial run using all 30 mutational signatures, which showed that these were present in the data and in signature extractions from lung squamous cell cancer (LUSC) datasets. The number of substitutions attributed to each signature is shown (a-b) as well as the proportion of mutations attributed to each mutational signature (c-d). Samples from the same patient share the same identifier except for the final letter; for example, PD21883a and PD21883d are two samples from the same patient. e, Comparison of the mutational signatures of CIS lesions to those found in lung squamous cell cancer (LUSC). LUSC data were downloaded from TCGA and mutations called with our algorithms. All mutations from all samples from each cancer type were pooled for this analysis. The colour scale indicates the proportion of substitutions in each sample that are attributed to each signature. f-j, Comparison of the relative proportion of mutations attributed to each signature between progressive (red; n = 29) and regressive (green; n = 10) CIS samples. P values were calculated using likelihood ratio tests of a mixed effects model with outcome (progressive or regressive) included as a fixed effect versus a model that was identical but for the fact that outcome was not included as a fixed effect. Only signature 4 (smoking-associated) was significantly different between the two groups. Boxplots are generated using the R boxplot function, which displays the first and third quartile as hinges and places whiskers at the most extreme data point that is no more than 1.5 times the length of the box away from the box.
Extended Data Fig. 3 Genome-wide copy number changes of CIS lesions.
Visualization of copy number changes for 39 whole-genome-sequenced CIS samples. Rows represent samples, genomic position is represented on the x-axis. Local copy number gains are illustrated in red, losses in blue. We observe widespread changes in progressive CIS samples and a subset of regressive samples.
Extended Data Fig. 4 Documentation of biopsy history and chronology of lesion appearance in three misclassified regressive cases.
a, Case 1 (PD21893a) appeared to regress from a CIS lesion (07/2012) to squamous metaplasia (SqM; 11/2012). However, again, CIS was subsequently reconfirmed by biopsy (05/2013). b, Case 2 (PD21884a) had a lobectomy for T1N0 lung squamous cell cancer (LUSC) in the left upper lobe (LUL) and was under surveillance for carcinoma-in-situ (CIS) at the resection margins. A subsequent, high-grade CIS lesion (08/2009) profiled for genome-wide DNA methylation changes was considered regressive since a follow-up biopsy on the same anatomical site demonstrated the presence of a low-grade, moderately dysplastic (MoD) lesion (11/2009). A subsequent biopsy, however, was classified as CIS (02/2011) and the lesion then remained static for 26 months but eventually progressed into invasive cancer (04/2014). c, Case 3 (PD38326a) had an initial diagnosis of CIS (11/2015) followed by regression to normal epithelium (03/2016). CIS was subsequently identified at the same site (03/2017), with invasive cancer diagnosed on subsequent biopsy (07/2017).
Extended Data Fig. 5 Genomic aberrations in pre-invasive lung CIS lesions.
Comparisons of the number of substitutions (a), small insertions and deletions (b), genome rearrangements (c) and copy number changes (d), showing significantly more genomic changes in progressive (n = 29) than regressive (n = 10) lesions. Although there were more clonal substitutions in progressive than regressive lesions (e), the proportion of substitutions that were clonal and the number of clones were similar (f-g). Progressive lesions had more putative driver mutations (h). Telomere lengths (base pairs) were similar between the two groups (i). To confirm an association between CIN gene expression and copy number change we correlated Weighted Genome Integrity Index (wGII) with mean CIN gene expression for the CIS samples in which we have both gene expression and whole-genome sequencing data (n = 11). Pearson correlation coefficient r2 = 0.473 (j). All P values were calculated using likelihood ratio tests of a mixed effects model with outcome (progressive or regressive) included as a fixed effect versus a model that was identical but for the fact that outcome was not included as a fixed effect. Boxplots are generated using the R boxplot function, which displays the first and third quartile as hinges and places whiskers at the most extreme data point that is no more than 1.5 times the length of the box away from the box.
Extended Data Fig. 6 Subclonal mutational structure in progressive and regressive CIS lesions.
Heatmap showing the proportion of overlapping mutations between samples taken from the same patient. For four patients with lesions that would ultimately progress to cancer (denoted ‘P’), over half the mutations were shared between any two given samples, suggesting that the lesions were derived from a common ancestral clone. By contrast, for two patients with lesions that would ultimately regress (denoted ‘R’), almost no mutations were shared, suggesting that the lesions arose independently. Samples from the same patient are shown in the same color; PD38321a and PD38322a do belong to the same patient and were mislabelled during processing.
Extended Data Fig. 7 Differential molecular changes between progressive and regressive lesions.
Visualization of differential changes across the genome. A, shows all identified differentially methylated regions (DMRs) (hypermethylated regions in yellow, hypomethylated in blue) alongside a similar analysis comparing cancer and control samples from The Cancer Genome Atlas. We observe that 58% of DMRs identified in our progressive vs regressive analysis are also identified in cancer vs control. B, shows copy number changes across the genome in regressive CIS, progressive CIS and TCGA cancer samples. We observe congruency of copy number change, suggesting similar processes in the two cohorts.
Extended Data Fig. 8 Principal component analysis investigating effect of various biological, clinical and technical factors affecting correct case segregation for all DMPs and gene expression data.
a-f, Principal component analysis based on all methylation probes (n = 87; 36 progressive, 18 regressive, 33 control). (a) Smoking history (pack years). (b) Chronic obstructive pulmonary disease (COPD) status. (c) Previous lung cancer history referring to the presence of lung squamous cell cancer (LUSC) prior to identification of pre-invasive lesions. (d) Age at bronchoscopy (years); age of individual when pre-invasive lesion was first biopsied. (e) Gender. (f) Sentix ID. g-k, Principal component analysis for all gene expression data. (g) Smoking history (pack years). (h) COPD status. (i) Previous lung cancer history referring to the presence of LUSC prior to identification of pre-invasive lesions. (j) Age at bronchoscopy (years); age of individual when pre-invasive lesion was first biopsied. (k) Gender. P-values were calculated using multivariate ANOVA.
Extended Data Fig. 9 Predictive modeling and ROC analytics of gene expression and CNA data.
ROC and precision-recall curves for the predictive model based on gene expression data shown in Fig. 4A-C. Curves are shown for the CIS discovery set (a-b), CIS validation set (c-d) and application to TCGA LUSC data (e-f). Using an analogous method to gene expression and methylation we used copy number data derived from methylation arrays to predict lesion outcome. Probe-level copy number changes were aggregated over cytogenetic bands; these data were used as input to Prediction Analysis of Microarrays (PAM). g-i, Probability plot based on a 154 cytogenetic band signature for correct class prediction (red circles indicate progressive lesions, green circles indicate regressive lesions). The area under the curve for the 154-cytogenetic band signature is 0.86. j-l, Application of our predictive model to previously published data (van Boerdonk et al.) replicates their result, classifying all regressive and 9/12 progressive samples correctly. This dataset included pre-invasive samples of various histological grades, rather than only CIS. m-o, Application of our predictive model to TCGA copy number data. Samples were correctly classified into TCGA LUSC and TCGA control samples with an AUC of 0.98.
Extended Data Fig. 10 Predictive modeling of methylation data.
In addition to the predictive modeling based on probe variation shown in Fig. 5, we used differentially expressed methylation probes to create a predictor using a Prediction Analysis for Microarrays (PAM) method. The model was trained on a training set (a-c) consisting of 26 progressive samples, 11 regressive samples and 23 control samples, shown in red, green and blue, respectively. A predictor based on 141 DMPs was created. This was applied to a validation set of 10 progressive, 7 regressive and 10 control samples (d-f), predicting outcome with AUC = 0.99. g-i, Application of our predictive model to TCGA methylation data. Samples were correctly classified into TCGA LUSC and TCGA control samples with AUC = 0.99. j-m, ROC analytics and precision-recall curves for Methylation Heterogeneity Index (MHI) model presented in Fig. 4. Curves apply to cancer vs control (j-k) and progressive vs regressive (l-m), respectively. n, Histogram of AUC values using MHI model with random samples of 2000 probes, applied to progressive vs regressive data. This demonstrates that a similar AUC is achieved with a random sample of probes as when using the entire array.
Supplementary information
Supplementary Tables
Supplementary Tables 1–5
Rights and permissions
About this article
Cite this article
Teixeira, V.H., Pipinikas, C.P., Pennycuick, A. et al. Deciphering the genomic, epigenomic, and transcriptomic landscapes of pre-invasive lung cancer lesions. Nat Med 25, 517–525 (2019). https://doi.org/10.1038/s41591-018-0323-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0323-0
This article is cited by
-
Tissue of origin prediction for cancer of unknown primary using a targeted methylation sequencing panel
Clinical Epigenetics (2024)
-
Transient loss of Polycomb components induces an epigenetic cancer fate
Nature (2024)
-
Multimodal analysis of cfDNA methylomes for early detecting esophageal squamous cell carcinoma and precancerous lesions
Nature Communications (2024)
-
Molecular profiling and prognostic biomarkers in chinese non-small cell lung cancer cohort
Diagnostic Pathology (2023)
-
USP13 drives lung squamous cell carcinoma by switching lung club cell lineage plasticity
Molecular Cancer (2023)