Abstract
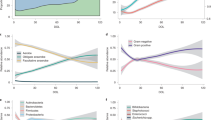
Commensal gut bacterial communities (microbiomes) are predicted to influence human health and disease1,2. Neonatal gut microbiomes are colonized with maternal and environmental flora and mature toward a stable composition over 2–3 years3,4. To study pre- and postnatal determinants of infant microbiome development, we analyzed 402 fecal metagenomes from 60 infants aged 0–8 months, using longitudinal generalized linear mixed models (GLMMs). Distinct microbiome signatures correlated with breastfeeding, formula ingredients, and maternal gestational weight gain (GWG). Amino acid synthesis pathway accretion in breastfed microbiomes complemented normative breastmilk composition. Prebiotic oligosaccharides, designed to promote breastfed-like microflora5, predicted functional pathways distinct from breastfed infant microbiomes. Soy formula in six infants was positively associated with Lachnospiraceae and pathways suggesting a short-chain fatty acid (SCFA)-rich environment, including glycerol to 1-butanol fermentation, which is potentially dysbiotic. GWG correlated with altered carbohydrate degradation and enriched vitamin synthesis pathways. Maternal and postnatal antibiotics predicted microbiome alterations, while delivery route had no persistent effects. Domestic water source correlates suggest water may be an underappreciated determinant of microbiome acquisition. Clinically important microbial pathways with statistically significant dietary correlates included dysbiotic markers6,7, core enterotype features8, and synthesis pathways for enteroprotective9 and immunomodulatory10,11 metabolites, epigenetic mediators1, and developmentally critical vitamins12, warranting further investigation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Sequence data supporting these findings have been deposited, along with relevant clinical metadata, in the SRA under BioProject ID PRJNA473126, with primary BioSample accession codes SAMN09259835–SAMN09260236 (study SRP148966). Source data for Figs. 1–4 are available online. Any additional data generated and analyzed in this study are available from the corresponding author upon reasonable request.
References
Indrio, F. et al. Epigenetic matters: the link between early nutrition, microbiome, and long-term health development. Front. Pediatr. 5, 178 (2017).
Clemente, J. C., Ursell, L. K., Parfrey, L. W. & Knight, R. The impact of the gut microbiota on human health: an integrative view. Cell 148, 1258–1270 (2012).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227 (2012).
Bäckhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17, 852 (2015).
Oozeer, R. et al. Intestinal microbiology in early life: specific prebiotics can have similar functionalities as human-milk oligosaccharides. Am. J. Clin. Nutr. 98, 561S–571S (2013).
de Weerth, C., Fuentes, S., Puylaert, P. & de Vos, W. M. Intestinal microbiota of infants with colic: development and specific signatures. Pediatrics 131, e550–e558 (2013).
Del Chierico, F. et al. Gut microbiota profiling of pediatric nonalcoholic fatty liver disease and obese patients unveiled by an integrated meta-omics-based approach. Hepatology 65, 451–464 (2017).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Yang, B., Feng, L., Wang, F. & Wang, L. Enterohemorrhagic Escherichia coli senses low biotin status in the large intestine for colonization and infection. Nat. Commun. 6, 6592 (2015).
Badurdeen, S., Mulongo, M. & Berkley, J. A. Arginine depletion increases susceptibility to serious infections in preterm newborns. Pediatr. Res. 77, 290–297 (2015).
Zhou, P., Li, Y., Ma, L. Y. & Lin, H. C. The role of immunonutrients in the prevention of necrotizing enterocolitis in preterm very low birth weight infants. Nutrients 7, 7256–7270 (2015).
Schwarzenberg, S. J. & Georgieff, M. K. The AAP Committee on Nutrition. Advocacy for improving nutrition in the first 1000 days to support childhood development and adult health. Pediatrics 141, e20173716 (2018).
Planer, J. D. et al. Development of the gut microbiota and mucosal IgA responses in twins and gnotobiotic mice. Nature 534, 263–266 (2016).
Zhang, Z., Adelman, A. S., Rai, D., Boettcher, J. & Lőnnerdal, B. Amino acid profiles in term and preterm human milk through lactation: a systematic review. Nutrients 5, 4800–4821 (2013).
Chu, D. M. et al. Maturation of the infant microbiome community structure and function across multiple body sites and in relation to mode of delivery. Nat. Med. 23, 314–326 (2017).
Butteiger, D. N. et al. Soy protein compared with milk protein in a Western diet increases gut microbial diversity and reduces serum lipids in golden Syrian hamsters. J. Nutr. 146, 697–705 (2016).
Yassour, M. et al. Natural history of the infant gut microbiome and impact of antibiotic treatment on bacterial strain diversity and stability. Sci. Transl. Med. 8, 343ra381 (2016).
Agostoni, C., Carratù, B., Boniglia, C., Riva, E. & Sanzini, E. Free amino acid content in standard infant formulas: comparison with human milk. J. Am. Coll. Nutr. 19, 434–438 (2000).
Sharon, G. et al. Specialized metabolites from the microbiome in health and disease. Cell Metab. 20, 719–730 (2014).
Haschke-Becher, E., Kainz, A. & Bachmann, C. Reference values of amino acids and of common clinical chemistry in plasma of healthy infants aged 1 and 4 months. J. Inherit. Metab. Dis. 39, 25–37 (2016).
Piacentini, G., Peroni, D., Bessi, E. & Morelli, L. Molecular characterization of intestinal microbiota in infants fed with soymilk. J. Pediatr. Gastroenterol. Nutr. 51, 71–76 (2010).
Vázquez, L., Flórez, A. B., Guadamuro, L. & Mayo, B. Effect of soy isoflavones on growth of representative bacterial species from the human gut. Nutrients 9, 727 (2017).
Li, S. et al. Continuously ingesting fructooligosaccharide can’t maintain rats’ gut Bifidobacterium at a high level. J. Food Sci. 80, M2530–M2534 (2015).
Bhatia, J. & Greer, F. The Committee on Nutrition. Use of soy protein-based formulas in infant feeding. Pediatrics 121, 1062–1068 (2008).
Vandenplas, Y. Prevention and management of cow’s milk allergy in non-exclusively breastfed infants. Nutrients 9, 731 (2017).
Bauchart-Thevret, C., Stoll, B., Chacko, S. & Burrin, D. G. Sulfur amino acid deficiency upregulates intestinal methionine cycle activity and suppresses epithelial growth in neonatal pigs. Am. J. Physiol. Endocrinol. Metab. 296, E1239–E1250 (2009).
Choe, E. K., Moon, J. S. & Park, K. J. Methionine enhances the contractile activity of human colon circular smooth muscle in vitro. J. Korean Med. Sci. 27, 777–783 (2012).
Neis, E. P., Dejong, C. H. & Rensen, S. S. The role of microbial amino acid metabolism in host metabolism. Nutrients 7, 2930–2946 (2015).
Alsaker, K. V., Paredes, C. & Papoutsakis, E. T. Metabolite stress and tolerance in the production of biofuels and chemicals: gene-expression-based systems analysis of butanol, butyrate, and acetate stresses in the anaerobe Clostridium acetobutylicum. Biotechnol. Bioeng. 105, 1131–1147 (2010).
Vitreschak, A. G., Rodionov, D. A., Mironov, A. A. & Gelfand, M. S. Regulation of riboflavin biosynthesis and transport genes in bacteria by transcriptional and translational attenuation. Nucleic Acids Res. 30, 3141–3151 (2002).
Stanislawski, M. A. et al. Pre-pregnancy weight, gestational weight gain, and the gut microbiota of mothers and their infants. Microbiome 5, 113 (2017).
Collado, M. C., Isolauri, E., Laitinen, K. & Salminen, S. Effect of mother’s weight on infant’s microbiota acquisition, composition, and activity during early infancy: a prospective follow-up study initiated in early pregnancy. Am. J. Clin. Nutr. 92, 1023–1030 (2010).
Antony, K. M. et al. The preterm placental microbiome varies in association with excess maternal gestational weight gain. Am. J. Obstet. Gynecol. 212, 653.e651–616 (2015).
Ma, J. et al. High-fat maternal diet during pregnancy persistently alters the offspring microbiome in a primate model. Nat. Commun. 5, 3889 (2014).
Hu, J. et al. Diversified microbiota of meconium is affected by maternal diabetes status. PLoS ONE 8, e78257 (2013).
Prince, A. L. et al. The perinatal microbiome and pregnancy: moving beyond the vaginal microbiome. Cold Spring Harb. Perspect. Med. 5, a023051 (2015).
Sacchetti, R., De Luca, G., Dormi, A., Guberti, E. & Zanetti, F. Microbial quality of drinking water from microfiltered water dispensers. Int. J. Hyg. Environ. Health 217, 255–259 (2014).
Dias, M. F. et al. Changes in mouse gut bacterial community in response to different types of drinking water. Water Res. 132, 79–89 (2017).
Poroyko, V. et al. Gut microbial gene expression in mother-fed and formula-fed piglets. PLoS ONE 5, e12459 (2010).
Charbonneau, M. R. et al. Sialylated milk oligosaccharides promote microbiota-dependent growth in models of infant undernutrition. Cell 164, 859–871 (2016).
Lim, E. S. et al. Early life dynamics of the human gut virome and bacterial microbiome in infants. Nat. Med. 21, 1228–1234 (2015).
Moore, A. M. et al. Gut resistome development in healthy twin pairs in the first year of life. Microbiome 3, 27 (2015).
Gurnee, E. A. et al. Gut colonization of healthy children and their mothers with pathogenic ciprofloxacin-resistant Escherichia coli. J. Infect. Dis. 212, 1862–1868 (2015).
Gibson, M. K. et al. Developmental dynamics of the preterm infant gut microbiota and antibiotic resistome. Nat. Microbiol. 1, 16024 (2016).
Fein, S. B. et al. Infant Feeding Practices Study II: study methods. Pediatrics 122(Suppl. 2)), S28–S35 (2008).
Segata, N. et al. Metagenomic microbial community profiling using unique clade-specific marker genes. Nat. Methods 9, 811–814 (2012).
McHardy, I. H. et al. Integrative analysis of the microbiome and metabolome of the human intestinal mucosal surface reveals exquisite inter-relationships. Microbiome 1, 17 (2013).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Schmieder, R. & Edwards, R. Fast identification and removal of sequence contamination from genomic and metagenomic datasets. PLoS ONE 6, e17288 (2011).
Abubucker, S. et al. Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLoS Comput. Biol. 8, e1002358 (2012).
Chu, D. M. et al. The early infant gut microbiome varies in association with a maternal high-fat diet. Genome Med. 8, 77 (2016).
Robinson, A. et al. Association of maternal gestational weight gain with the infant fecal microbiota. J. Pediatr. Gastroenterol. Nutr. 65, 509–515 (2017).
Singh, S., Karagas, M. R. & Mueller, N. T. Charting the maternal and infant microbiome: what is the role of diabetes and obesity in pregnancy? Curr. Diab. Rep. 17, 11 (2017).
American College of Obstetricians and Gynecologists. ACOG Committee opinion no. 548: weight gain during pregnancy. Obstet. Gynecol. 121, 210–212 (2013).
Joo, J. W., Hormozdiari, F., Han, B. & Eskin, E. Multiple testing correction in linear mixed models. Genome Biol. 17, 62 (2016).
Acknowledgements
This work is supported in part by awards to G.D. through the Edward Mallinckrodt, Jr. Foundation (Scholar Award), and the National Institute of General Medical Sciences (http://www.nigms.nih.gov/) of the National Institutes of Health (NIH) under award number R01GM099538. A.M.B.-D. was supported by the National Institutes of Diabetes and Digestive and Kidney Diseases of the NIH under award number K08-DK102673. A.W.D. received support from the Institutional Program Unifying Population and Laboratory-Based Sciences Burroughs Wellcome Fund grant to Washington University. B.B.W. and P.I.T. received support for the cohort and sample collection from the Children’s Discovery Institute of Washington University and St. Louis Children’s Hospital, and P.I.T. is supported by P30DK052574 (Biobank Core). P.I.T., B.B.W., and G.D. are also supported in part by a grant from the Eunice Kennedy Shriver National Institute of Child Health & Human Development (https://www.nichd.nih.gov/) of the NIH under award number R01HD092414. The content is solely the responsibility of the authors and does not necessarily represent the official views of the funding agencies. We would like to thank E. Martin, B. Koebbe, and J. Hoisington-López from the Edison Family Center for Genome Sciences & Systems Biology at Washington University School of Medicine for technical support in high-throughput computing and sequencing. We would like to thank A. J. Gasparrini, B. Wang, and B. Berla for technical assistance in experimental and computational protocol optimization for whole-metagenome shotgun sequencing of fecal samples. We would like to thank I. M. Ndao, N. Shaikh, S. Patel, B. Wang, and S. X. Sun for archival and maintenance of frozen fecal sample inventory. We would like to thank F. S. Cole and members of the Dantas lab for general helpful discussions regarding the research presented in this manuscript, and K. Guilonard for helpful comments on the text.
Author information
Authors and Affiliations
Contributions
A.M.B.-D., A.W.D., B.B.W., P.I.T., and G.D. conceived of experiments and design of work and analyses. B.B.W. and P.I.T. oversaw collection and stewardship of fecal samples and clinical metadata inventories. A.M.B.-D. performed wet-lab experiments with advice from G.D. A.M.B.-D. performed computational analyses with advice from A.W.D. and G.D. Article drafting was performed by A.M.B.-D. with critical revision performed by A.W.D., B.B.W., P.I.T., and G.D.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1 and 2
Supplementary Table 3
Maximum-likelihood longitudinal multivariate GLMM model information
Supplementary Table 4
Pathway-identified taxa
Supplementary Table 5
Qualitative summary of significant associations of clinical variables with taxa and pathways
Supplementary Table 6
Sample size for binary variables
Supplementary Table 7
Infant formula brands and ingredients
Supplementary Table 8
Taxa identified in zymobiomics community standard positive control samples
Rights and permissions
About this article
Cite this article
Baumann-Dudenhoeffer, A.M., D’Souza, A.W., Tarr, P.I. et al. Infant diet and maternal gestational weight gain predict early metabolic maturation of gut microbiomes. Nat Med 24, 1822–1829 (2018). https://doi.org/10.1038/s41591-018-0216-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0216-2
This article is cited by
-
Exploration of pathogenic microorganism within the small intestine of necrotizing enterocolitis
World Journal of Pediatrics (2024)
-
Vertical transmission of the gut microbiota influences glucose metabolism in offspring of mice with hyperglycaemia in pregnancy
Microbiome (2022)
-
The establishment of the gut microbiota in 1-year-aged infants: from birth to family food
European Journal of Nutrition (2022)
-
Dynamic colonization of gut microbiota and its influencing factors among the breast-feeding infants during the first two years of life
Journal of Microbiology (2022)
-
MFGM components promote gut Bifidobacterium growth in infant and in vitro
European Journal of Nutrition (2022)