Abstract
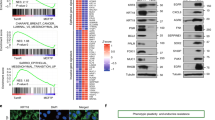
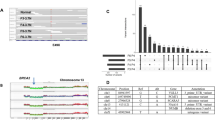
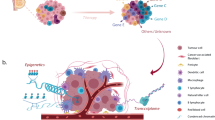
The degree of intrinsic and interpatient phenotypic heterogeneity and its role in tumor evolution is poorly understood. Phenotypic drifts can be transmitted via inheritable transcriptional programs. Cell-type specific transcription is maintained through the activation of epigenetically defined regulatory regions including promoters and enhancers. Here we have annotated the epigenome of 47 primary and metastatic estrogen-receptor (ERα)-positive breast cancer clinical specimens and inferred phenotypic heterogeneity from the regulatory landscape, identifying key regulatory elements commonly shared across patients. Shared regions contain a unique set of regulatory information including the motif for transcription factor YY1. We identify YY1 as a critical determinant of ERα transcriptional activity promoting tumor growth in most luminal patients. YY1 also contributes to the expression of genes mediating resistance to endocrine treatment. Finally, we used H3K27ac levels at active enhancer elements as a surrogate of intra-tumor phenotypic heterogeneity to track the expansion and contraction of phenotypic subpopulations throughout breast cancer progression. By tracking the clonality of SLC9A3R1-positive cells, a bona fide YY1-ERα-regulated gene, we show that endocrine therapies select for phenotypic clones under-represented at diagnosis. Collectively, our data show that epigenetic mechanisms significantly contribute to phenotypic heterogeneity and evolution in systemically treated breast cancer patients.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ferlay, J. et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 136, E359–E386 (2015).
Ali, S., Buluwela, L. & Coombes, C. Antiestrogens and their therapeutic applications in breast cancer and other diseases. Ann. Rev. Med. 62, 217–232 (2010).
Perou, C. et al. Molecular portraits of human breast tumours. Nature 406, 747–752 (2000).
Genestie, C. et al. Comparison of the prognostic value of Scarff–Bloom–Richardson and Nottingham histological grades in a series of 825 cases of breast cancer: major importance of the mitotic count as a component of both grading systems. Anticancer Res. 18, 571–576 (1998).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Koboldt, D. et al. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
EBCTCG. Aromatase inhibitors versus tamoxifen in early breast cancer: patient-level meta-analysis of the randomised trials. Lancet 386, 1341–1352 (2015).
Magnani et al. Acquired CYP19A1 amplification is an early specific mechanism of aromatase inhibitor resistance in ERα metastatic breast cancer. Nat. Genet. 49, 444–450 (2017).
Yates, L. et al. Genomic evolution of breast cancer metastasis and relapse. Cancer Cell 32, 169–184 (2017).
Nguyen, V. et al. Differential epigenetic reprogramming in response to specific endocrine therapies promotes cholesterol biosynthesis and cellular invasion. Nat. Commun. 6, 10044 (2015).
Roadmap Epigenomics Consortium. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
ENCODE Project Consortium. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Whyte, W. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Heintzman, N. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 39, 311–318 (2007).
Falahi, F. et al. Towards sustained silencing of HER2/neu in cancer by epigenetic editing. Mol. Cancer Res. 11, 1029–1039 (2013).
Laprell, F., Finkl, K. & Müller, J. Propagation of polycomb-repressed chromatin requires sequence-specific recruitment to DNA. Science 356, 85–88 (2017).
Wang, X. & Moazed, D. DNA sequence-dependent epigenetic inheritance of gene silencing and histone H3K9 methylation. Science 356, 88–91 (2017).
Coleman, R. T. & Struhl, G. Causal role for inheritance of H3K27me3 in maintaining the OFF state of a Drosophila HOX gene. Science 356, eaai8236 (2017).
Magnani, L., Eeckhoute, J. & Lupien, M. Pioneer factors: directing transcriptional regulators within the chromatin environment. Trends Genet. 27, 465–474 (2011).
Jozwik, K. M. & Carroll, J. S. Pioneer factors in hormone-dependent cancers. Nat. Rev. Cancer 12, 381–385 (2012).
Hnisz, D. et al. Convergence of developmental and oncogenic signaling pathways at transcriptional super-enhancers. Mol. Cell 58, 362–370 (2015).
Heintzman, N. D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009).
Yates, L. R. et al. Subclonal diversification of primary breast cancer revealed by multiregion sequencing. Nat. Med. 21, 751–759 (2015).
Williams, M. J., Werner, B., Barnes, C., Graham, T. & Sottoriva, A. Identification of neutral tumor evolution across cancer types. Nat. Genet. 48, 238–244 (2016).
Lan, X. et al. Fate mapping of human glioblastoma reveals an invariant stem cell hierarchy. Nature 549, 227–232 (2017).
Tirosh, I. et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313 (2016).
Harvey, J. M., Clark, G. M., Osborne, C. K. & Allred, D. C. Estrogen receptor status by immunohistochemistry is superior to the ligand-binding assay for predicting response to adjuvant endocrine therapy in breast cancer. J. Clin. Oncol. 17, 1474–1481 (1999).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Cowper-Sal Iari, R. et al. Breast cancer risk-associated SNPs modulate the affinity of chromatin for FOXA1 and alter gene expression. Nat. Genet. 44, 1191–1198 (2012).
Cohen, A. J. et al. Hotspots of aberrant enhancer activity punctuate the colorectal cancer epigenome. Nat. Commun. 8, 14400 (2017).
Michailidou, K. et al. Association analysis identifies 65 new breast cancer risk loci. Nature 551, 92–94 (2017).
Levsky, J. M. & Singer, R. H. Gene expression and the myth of the average cell. Trends Cell Biol. 13, 4–6 (2003).
Wang, S. et al. Target analysis by integration of transcriptome and ChIP–seq data with BETA. Nat. Protoc. 8, 2502–2515 (2013).
Gyorffy, B. et al. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res. 123, 725–731 (2010).
Neph, S. et al. An expansive human regulatory lexicon encoded in transcription factor footprints. Nature 489, 83–90 (2012).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Paakinaho, V. et al. Single-molecule analysis of steroid receptor and cofactor action in living cells. Nat. Commun. 8, 15896 (2017).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Carroll, J. S. et al. Chromosome-wide mapping of estrogen receptor binding reveals long-range regulation requiring the Forkhead protein FoxA1. Cell 122, 33–43 (2005).
Beagan, J. A. et al. YY1 and CTCF orchestrate a 3D chromatin looping switch during early neural lineage commitment. Genome Res. 27, 1139–1152 (2017).
Weintraub, A. et al. YY1 is a structural regulator of enhancer–promoter loops. Cell 171, 1573–1588 (2017).
Vella, P., Barozzi, I., Cuomo, A., Bonaldi, T. & Pasini, D. Yin Yang 1 extends the Myc-related transcription factors network in embryonic stem cells. Nucleic Acids Res. 40, 3403–3418 (2012).
Jeon, Y. & Lee, J. T. YY1 tethers Xist RNA to the inactive X nucleation center. Cell 146, 119–133 (2011).
Sigova, A. A. et al. Transcription factor trapping by RNA in gene regulatory elements. Science 350, 978–981 (2015).
Klymenko, T. et al. A polycomb group protein complex with sequence-specific DNA-binding and selective methyl-lysine-binding activities. Genes Dev. 20, 1110–1122 (2006).
Tang, Z. et al. CTCF-mediated human 3D genome architecture reveals chromatin topology for transcription. Cell 163, 1611–1627 (2015).
Hurtado, A., Holmes, K., Ross-Innes, C., Schmidt, D. & Carroll, J. FOXA1 is a key determinant of estrogen receptor function and endocrine response. Nat. Genet. 43, 27–33 (2011).
Cardone, R. A., Casavola, V. & Reshkin, S. J. The role of disturbed pH dynamics and the Na+/H+ exchanger in metastasis. Nat. Rev. Cancer 5, 786–795 (2005).
Gerlinger, M. et al. Genomic architecture and evolution of clear cell renal cell carcinomas defined by multiregion sequencing. Nat. Genet. 46, 225–233 (2014).
McGranahan, N. & Swanton, C. Biological and therapeutic impact of intratumor heterogeneity in cancer evolution. Cancer Cell 27, 15–26 (2015).
Juric, D. et al. Convergent loss of PTEN leads to clinical resistance to a PI(3)Kα inhibitor. Nature 518, 240–244 (2015).
Shah, S. et al. Mutational evolution in a lobular breast tumour profiled at single nucleotide resolution. Nature 461, 809–813 (2009).
Arduino, S. et al. Reduced IL-2 level concentration in patients with breast cancer as a possible risk factor for relapse. Eur. J. Gynaecol. 17, 535–537 (1996).
Cai, Y. et al. YY1 functions with INO80 to activate transcription. Nat. Struct. Mol. Biol. 14, 872–874 (2007).
Onder, T. et al. Chromatin-modifying enzymes as modulators of reprogramming. Nature 486, 598–602 (2012).
Whalen, S., Truty, R. M. & Pollard, K. S. Enhancer–promoter interactions are encoded by complex genomic signatures on looping chromatin. Nat. Genet. 48, 488–496 (2016).
Schmidt, D. et al. ChIP–seq: using high-throughput sequencing to discover protein–DNA interactions. Methods 48, 240–248 (2009).
Acknowledgements
The authors acknowledge and thanks all patients and their families for support and for donating research samples. The authors thank the Breast Cancer Now Tissue Bank (project TR0121), Imperial Tissue Bank and the LEGACY study for contributing tissues. The authors acknowledge infrastructure support from the Cancer Research UK Imperial Centre, the Imperial Experimental Cancer Medicine Centre and the National Institute for Health Research Imperial Biomedical Research Centre. L.M. was supported by a CRUK fellowship (C46704/A23110) and an Imperial Junior Fellowship (G53019). D.P. was supported by a Wellcome Trust PhD studentship (103034/Z/13/Z). G.C. was supported by a Marie Skłodowska Curie Training Grant (642691, EpiPredict). G.P. was supported by AIRC IG 2016-18696. The authors thank J.A. Buendia, L. Watson and J. Carrol for constructive comments on the manuscript.
Author information
Authors and Affiliations
Contributions
L.M. conceived the study. D.K.P., E.E., N.S. and Y.P. performed the experiments. L.M., G.C., B.G., A.S., L.S.P., I.B. and P.S. developed and performed bioinformatic analyses. K.G. organized tissue collection. D.J.H., G.S., P.B., C.P. and R.C.C. recruited patients and supplied tissues. S.S. performed pathology assessment of ChIP–seq processed samples. G.P. provided matched material. A.V. and G.P. performed IHC staining and scoring. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–19 and Supplementary Methods
Supplementary Table 1
Clinical characteristics of the patient dataset
Supplementary Table 2
Summary statistics for ChIP-seq in clinical samples
Supplementary Dataset
Coding mutations at cancer genes
Rights and permissions
About this article
Cite this article
Patten, D.K., Corleone, G., Győrffy, B. et al. Enhancer mapping uncovers phenotypic heterogeneity and evolution in patients with luminal breast cancer. Nat Med 24, 1469–1480 (2018). https://doi.org/10.1038/s41591-018-0091-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0091-x
This article is cited by
-
circRNA-SFMBT2 orchestrates ERα activation to drive tamoxifen resistance in breast cancer cells
Cell Death & Disease (2023)
-
Molecular classification of hormone receptor-positive HER2-negative breast cancer
Nature Genetics (2023)
-
Heterogeneity of the tumor immune microenvironment and clinical interventions
Frontiers of Medicine (2023)
-
Heterogeneity of the tumor immune microenvironment and its clinical relevance
Experimental Hematology & Oncology (2022)
-
Endocrine resistance and breast cancer plasticity are controlled by CoREST
Nature Structural & Molecular Biology (2022)