Abstract
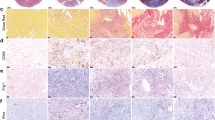
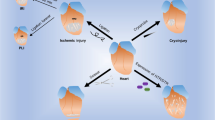
Paradigm-shifting studies in the mouse have identified tissue macrophage heterogeneity as a critical determinant of immune responses. In contrast, surprisingly little is known regarding macrophage heterogeneity in humans. Macrophages within the mouse heart are partitioned into CCR2− and CCR2+ subsets with divergent origins, repopulation mechanisms, and functions. Here, we demonstrate that the human myocardium also contains distinct subsets of CCR2− and CCR2+ macrophages. Analysis of sex-mismatched heart transplant recipients revealed that CCR2− macrophages are a tissue-resident population exclusively replenished through local proliferation, whereas CCR2+ macrophages are maintained through monocyte recruitment and proliferation. Moreover, CCR2− and CCR2+ macrophages have distinct functional properties, analogous to reparative CCR2− and inflammatory CCR2+ macrophages in the mouse heart. Clinically, CCR2+ macrophage abundance is associated with left ventricular remodeling and systolic function in heart failure patients. Collectively, these observations provide initial evidence for the functional importance of macrophage heterogeneity in the human heart.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Van Furth, R. & Cohn, Z. A. The origin and kinetics of mononuclear phagocytes. J. Exp. Med. 128, 415–435 (1968).
Volkman, A., Chang, N. C., Strausbauch, P. H. & Morahan, P. S. Differential effects of chronic monocyte depletion on macrophage populations. Lab. Invest. 49, 291–298 (1983).
Sawyer, R. T., Strausbauch, P. H. & Volkman, A. Resident macrophage proliferation in mice depleted of blood monocytes by strontium-89. Lab. Invest. 46, 165–170 (1982).
Ginhoux, F. et al. Fate mapping analysis reveals that adult microglia derive from primitive macrophages. Science 330, 841–845 (2010).
Schulz, C. et al. A lineage of myeloid cells independent of Myb and hematopoietic stem cells. Science 336, 86–90 (2012).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Hashimoto, D. et al. Tissue-resident macrophages self-maintain locally throughout adult life with minimal contribution from circulating monocytes. Immunity 38, 792–804 (2013).
Epelman, S. et al. Embryonic and adult-derived resident cardiac macrophages are maintained through distinct mechanisms at steady state and during inflammation. Immunity 40, 91–104 (2014).
Guilliams, M. et al. Alveolar macrophages develop from fetal monocytes that differentiate into long-lived cells in the first week of life via GM-CSF. J. Exp. Med. 210, 1977–1992 (2013).
Jakubzick, C. et al. Minimal differentiation of classical monocytes as they survey steady-state tissues and transport antigen to lymph nodes. Immunity 39, 599–610 (2013).
Gordon, S. & Pluddemann, A. Tissue macrophages: heterogeneity and functions. BMC Biol. 15, 53 (2017).
Boyer, S. W., Schroeder, A. V., Smith-Berdan, S. & Forsberg, E. C. All hematopoietic cells develop from hematopoietic stem cells through Flk2/Flt3-positive progenitor cells. Cell Stem Cell 9, 64–73 (2011).
Hoeffel, G. et al. Adult Langerhans cells derive predominantly from embryonic fetal liver monocytes with a minor contribution of yolk sac–derived macrophages. J. Exp. Med. 209, 1167–1181 (2012).
Sieweke, M. H. & Allen, J. E. Beyond stem cells: self-renewal of differentiated macrophages. Science 342, 1242974 (2013).
Davies, L. C., Jenkins, S. J., Allen, J. E. & Taylor, P. R. Tissue-resident macrophages. Nat. Immunol. 14, 986–995 (2013).
Wynn, T. A., Chawla, A. & Pollard, J. W. Macrophage biology in development, homeostasis and disease. Nature 496, 445–455 (2013).
Lavine, K. J. et al. Distinct macrophage lineages contribute to disparate patterns of cardiac recovery and remodeling in the neonatal and adult heart. Proc. Natl Acad. Sci. USA 111, 16029–16034 (2014).
Leid, J. et al. Primitive embryonic macrophages are required for coronary development and maturation. Circ. Res. 118, 1498–1511 (2016).
Li, W. et al. Heart-resident CCR2 + macrophages promote neutrophil extravasation through TLR9/MyD88/CXCL5 signaling. JCI Insight 1, e87315 (2016).
Xue, J. et al. Transcriptome-based network analysis reveals a spectrum model of human macrophage activation. Immunity 40, 274–288 (2014).
Hulsmans, M. Macrophages facilitate electrical conduction in the heart. Cell 169, 510–522 (2017).
Sager, H. B. et al. Proliferation and recruitment contribute to myocardial macrophage expansion in chronic heart failure. Circ. Res. 119, 853–864 (2016).
Gautier, E. L. et al. Gene-expression profiles and transcriptional regulatory pathways that underlie the identity and diversity of mouse tissue macrophages. Nat. Immunol. 13, 1118–1128 (2012).
Reynolds, G. Human and mouse mononuclear phagocyte networks: A tale of two species. Front. Immunol. 6, 330 (2015).
Zarif, J. C. A phased strategy to differentiate human CD14 + monocytes into classically and alternatively activated macrophages and dendritic cells. Biotechniques 61, 33–41 (2016).
Garlanda, C., Bottazzi, B., Bastone, A. & Mantovani, A. Pentraxins at the crossroads between innate immunity, inflammation, matrix deposition, and female fertility. Annu. Rev. Immunol. 23, 337–366 (2005).
Fujiu, K. et al. A heart–brain–kidney network controls adaptation to cardiac stress through tissue macrophage activation. Nat. Med. 23, 611–622 (2017).
Kubin, T. et al. Oncostatin M is a major mediator of cardiomyocyte dedifferentiation and remodeling. Cell Stem Cell 9, 420–432 (2011).
Vickers, A. E. et al. Organ slice viability extended for pathway characterization: an in vitro model to investigate fibrosis. Toxicol. Sci. 82, 534–544 (2004).
Brandenburger, M. et al. Organotypic slice culture from human adult ventricular myocardium. Cardiovasc. Res. 93, 50–59 (2012).
Diakos, N. A. et al. Myocardial atrophy and chronic mechanical unloading of the failing human heart: implications for cardiac assist device-induced myocardial recovery. J. Am. Coll. Cardiol. 64, 1602–1612 (2014).
Drakos, S. G. et al. Magnitude and time course of changes induced by continuous-flow left ventricular assist device unloading in chronic heart failure: insights into cardiac recovery. J. Am. Coll. Cardiol. 61, 1985–1994 (2013).
Epelman, S., Lavine, K. J. & Randolph, G. J. Origin and functions of tissue macrophages. Immunity. 41, 21–35 (2014).
Heidt, T. et al. Differential contribution of monocytes to heart macrophages in steady-state and after myocardial infarction. Circ. Res. 115, 284–295 (2014).
Molawi, K. et al. Progressive replacement of embryo-derived cardiac macrophages with age. J. Exp. Med. 211, 2151–2158 (2014).
Kaufmann, A., Salentin, R., Gemsa, D. & Sprenger, H. Increase of CCR1 and CCR5 expression and enhanced functional response to MIP-1 alpha during differentiation of human monocytes to macrophages. J. Leukoc. Biol. 69, 248–252 (2001).
Weber, C. et al. Differential chemokine receptor expression and function in human monocyte subpopulations. J. Leukoc. Biol. 67, 699–704 (2000).
Burchfield, J. S., Xie, M. & Hill, J. A. Pathological ventricular remodeling: mechanisms: part 1 of 2. Circulation 128, 388–400 (2013).
Kanitakis, J., Petruzzo, P. & Dubernard, J. M. Turnover of epidermal Langerhans’ cells. N. Engl. J. Med. 351, 2661–2662 (2004).
Eguiluz-Gracia, I. et al. Long-term persistence of human donor alveolar macrophages in lung transplant recipients. Thorax 71, 1006–1011 (2016).
Bittmann, I. et al. Cellular chimerism of the lung after transplantation. An interphase cytogenetic study. Am. J. Clin. Pathol. 115, 525–533 (2001).
Haniffa, M. Differential rates of replacement of human dermal dendritic cells and macrophages during hematopoietic stem cell transplantation. J. Exp. Med. 206, 371–385 (2009).
Dickinson, R. E. et al. Exome sequencing identifies GATA-2 mutation as the cause of dendritic cell, monocyte, B and NK lymphoid deficiency. Blood 118, 2656–2658 (2011).
Hambleton, S. IRF8 mutations and human dendritic-cell immunodeficiency. N. Engl. J. Med. 365, 127–138 (2011).
Bigley, V. et al. The human syndrome of dendritic cell, monocyte, B and NK lymphoid deficiency. J. Exp. Med. 208, 227–234 (2011).
Desch, A. N. Flow cytometric analysis of mononuclear phagocytes in nondiseased human lung and lung-draining lymph nodes. Am. J. Respir. Crit. Care Med. 193, 614–626 (2016).
Acknowledgements
We acknowledge the Translational Cardiovascular Biorepository at Washington University for providing cardiac tissue specimens. This project was made possible by funding provided from the Children’s Discovery Institute of Washington University and St. Louis Children’s Hospital (CH-II-2015-462, CH-II-2017-628), the Foundation for Barnes-Jewish Hospital (8038-88), and the NHLBI (R01 HL138466, R01 HL139714). K.J.L. is supported by National Institutes of Health (NIH) K08 HL123519 and Burroughs Welcome Fund (1014782). Histology was performed in the DDRCC advanced imaging and tissue analysis core supported by Grant #P30 DK52574. The Genome Technology Access Center in the Department of Genetics at Washington University School of Medicine is partially supported by NCI Cancer Center Support Grant #P30 CA91842 to the Siteman Cancer Center and by ICTS/CTSA Grant #UL1TR000448 from the National Center for Research Resources (NCRR), a component of the NIH, and NIH Roadmap for Medical Research. S.G.D. is supported by NHLBI R01 HL135121-01, AHA HF 16SFRN29020000-Project 1, and the Nora Eccles Treadwell Foundation. M.N. is supported by NIH HL139598 and the MGH Research Scholar Program. Y.L. is supported by NIH R01HL131908 and R01HL125655. D.K. is supported by NIH P01AI116501 and R01 HL094601, Veterans Administration Merit Review grant 1I01BX002730 and the Foundation for Barnes-Jewish Hospital. M.H. was supported by an MGH ECOR Tosteson and Fund for Medical Discovery Fellowship (2017A052660).
Author information
Authors and Affiliations
Contributions
G.B. performed the flow cytometry, gene expression profiling, and macrophage in vitro assays. C.S. performed and analyzed the immunostaining experiments. A.I. provided cardiac specimens. N.W. assisted with quantitative data analyses. S.G.D., C.H.S., and T.S.S. provided cardiac specimens and clinical data for the LVAD patient cohort. G.B., D.K., M.H., M.N., S.E., Y.L., and A.B. assisted with study design, data interpretation, and manuscript production. K.J.L. was responsible for all aspects of this study including study design, experimental execution, data analysis, data interpretation, and manuscript production.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1–3
Rights and permissions
About this article
Cite this article
Bajpai, G., Schneider, C., Wong, N. et al. The human heart contains distinct macrophage subsets with divergent origins and functions. Nat Med 24, 1234–1245 (2018). https://doi.org/10.1038/s41591-018-0059-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0059-x
This article is cited by
-
Coenzyme Q10 mitigates macrophage mediated inflammation in heart following myocardial infarction via the NLRP3/IL1β pathway
BMC Cardiovascular Disorders (2024)
-
Immune checkpoints in cardiac physiology and pathology: therapeutic targets for heart failure
Nature Reviews Cardiology (2024)
-
Immunomodulation and immunopharmacology in heart failure
Nature Reviews Cardiology (2024)
-
Animal models to study cardiac regeneration
Nature Reviews Cardiology (2024)
-
ATF3’s shielding hand to counter ischemia-reperfusion injury in cardiac macrophages
Nature Cardiovascular Research (2024)