Abstract
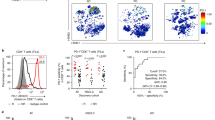
Evidence from mouse chronic viral infection models suggests that CD8+ T cell subsets characterized by distinct expression levels of the receptor PD-1 diverge in their state of exhaustion and potential for reinvigoration by PD-1 blockade. However, it remains unknown whether T cells in human cancer adopt a similar spectrum of exhausted states based on PD-1 expression levels. We compared transcriptional, metabolic and functional signatures of intratumoral CD8+ T lymphocyte populations with high (PD-1T), intermediate (PD-1N) and no PD-1 expression (PD-1–) from non-small-cell lung cancer patients. PD-1T T cells showed a markedly different transcriptional and metabolic profile from PD-1N and PD-1– lymphocytes, as well as an intrinsically high capacity for tumor recognition. Furthermore, while PD-1T lymphocytes were impaired in classical effector cytokine production, they produced CXCL13, which mediates immune cell recruitment to tertiary lymphoid structures. Strikingly, the presence of PD-1T cells was strongly predictive for both response and survival in a small cohort of non-small-cell lung cancer patients treated with PD-1 blockade. The characterization of a distinct state of tumor-reactive, PD-1-bright lymphocytes in human cancer, which only partially resembles that seen in chronic infection, provides potential avenues for therapeutic intervention.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Castle, J. C. et al. Exploiting the mutanome for tumor vaccination. Cancer Res. 72, 1081–1091 (2012).
Heemskerk, B., Kvistborg, P. & Schumacher, T. N. The cancer antigenome. EMBO J. 32, 194–203 (2013).
Mellman, I., Coukos, G. & Dranoff, G. Cancer immunotherapy comes of age. Nature 480, 480–489 (2011).
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 331, 1565–1570 (2011).
Schietinger, A. & Greenberg, P. D. Tolerance and exhaustion: defining mechanisms of T cell dysfunction. Trends Immunol. 35, 51–60 (2014).
Baitsch, L. et al. Exhaustion of tumor-specific CD8+ T cells in metastases from melanoma patients. J. Clin. Invest. 121, 2350–2360 (2011).
Zajac, A. J. et al. Viral immune evasion due to persistence of activated T cells without effector function. J. Exp. Med. 188, 2205–2213 (1998).
Wherry, E. J. T cell exhaustion. Nat. Immunol. 12, 492–499 (2011).
Wherry, E. J. et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity 27, 670–684 (2007).
Day, C. L. et al. PD-1 expression on HIV-specific T cells is associated with T-cell exhaustion and disease progression. Nature 443, 350–354 (2006).
Trautmann, L. et al. Upregulation of PD-1 expression on HIV-specific CD8+ T cells leads to reversible immune dysfunction. Nat. Med. 12, 1198–1202 (2006).
Golden-Mason, L. et al. Upregulation of PD-1 expression on circulating and intrahepatic hepatitis C virus-specific CD8+ T cells associated with reversible immune dysfunction. J. Virol. 81, 9249–9258 (2007).
Ahmadzadeh, M. et al. Tumor antigen-specific CD8 T cells infiltrating the tumor express high levels of PD-1 and are functionally impaired. Blood 114, 1537–1544 (2009).
Thommen, D. S. et al. Progression of lung cancer is associated with increased dysfunction of T cells defined by coexpression of multiple inhibitory receptors. CancerImmunol. Res. 3, 1344–1355 (2015).
Schreiner, J. et al. Expression of inhibitory receptors on intratumoral T cells modulates the activity of a T cell-bispecific antibody targeting folate receptor. Oncoimmunology 5, e1062969 (2015).
Zippelius, A. et al. Effector function of human tumor-specific CD8 T cells in melanoma lesions: a state of local functional tolerance. Cancer Res. 64, 2865–2873 (2004).
Blackburn, S. D., Shin, H., Freeman, G. J. & Wherry, E. J. Selective expansion of a subset of exhausted CD8 T cells by αPD-L1 blockade. Proc. Natl Acad. Sci. USA 105, 15016–15021 (2008).
Paley, M. A. et al. Progenitor and terminal subsets of CD8+ T cells cooperate to contain chronic viral infection. Science 338, 1220–1225 (2012).
Grosso, J. F. et al. Functionally distinct LAG-3 and PD-1 subsets on activated and chronically stimulated CD8 T cells. J. Immunol. 182, 6659–6669 (2009).
Sakuishi, K. et al. Targeting Tim-3 and PD-1 pathways to reverse T cell exhaustion and restore anti-tumor immunity. J. Exp. Med. 207, 2187–2194 (2010).
Kansy, B. A. et al. PD-1 status in CD8+ T cells associates with survival and anti-PD-1 therapeutic outcomes in head and neck cancer. Cancer Res. 77, 6353–6364 (2017).
Bolotin, D. A. et al. MiXCR: software for comprehensive adaptive immunity profiling. Nat. Methods 12, 380–381 (2015).
Wolfl, M. et al. Activation-induced expression of CD137 permits detection, isolation, and expansion of the full repertoire of CD8+ T cells responding to antigen without requiring knowledge of epitope specificities. Blood 110, 201–210 (2007).
Gros, A. et al. PD-1 identifies the patient-specific CD8+ tumor-reactive repertoire infiltrating human tumors. J. Clin. Invest. 124, 2246–2259 (2014).
Gros, A. et al. Prospective identification of neoantigen-specific lymphocytes in the peripheral blood of melanoma patients. Nat. Med. 22, 433–438 (2016).
Inozume, T. et al. Selection of CD8+PD-1+ lymphocytes in fresh human melanomas enriches for tumor-reactive T cells. J. Immunother. 33, 956–964 (2010).
Zheng, C. et al. Landscape of infiltrating T cells in liver cancer revealed by single-cell sequencing. Cell 169, 1342–1356.e1316 (2017).
Henson, S. M. et al. KLRG1 signaling induces defective Akt (Ser473) phosphorylation and proliferative dysfunction of highly differentiated CD8+ T cells. Blood 113, 6619–6628 (2009).
Crawford, A. et al. Molecular and transcriptional basis of CD4+ T cell dysfunction during chronic infection. Immunity 40, 289–302 (2014).
Schietinger, A. et al. Tumor-specific T cell dysfunction is a dynamic antigen-driven differentiation program initiated early during tumorigenesis. Immunity 45, 389–401 (2016).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Singer, M. et al. A distinct gene module for dysfunction uncoupled from activation in tumor-infiltrating T cells. Cell 166, 1500–1511.e1509 (2016).
Gupta, P. K. et al. CD39 expression identifies terminally exhausted CD8+ T cells. PLoS Pathog. 11, e1005177 (2015).
Quigley, M. et al. Transcriptional analysis of HIV-specific CD8+ T cells shows that PD-1 inhibits T cell function by upregulating BATF. Nat. Med. 16, 1147–1151 (2010).
Doering, T. A. et al. Network analysis reveals centrally connected genes and pathways involved in CD8+ T cell exhaustion versus memory. Immunity 37, 1130–1144 (2012).
Scharping, N. E. et al. The tumor microenvironment represses T cell mitochondrial biogenesis to drive intratumoral T cell metabolic insufficiency and dysfunction. Immunity 45, 701–703 (2016).
Chang, C. H. et al. Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell 162, 1229–1241 (2015).
Ho, P. C. et al. Phosphoenolpyruvate is a metabolic checkpoint of anti-tumor T cell responses. Cell 162, 1217–1228 (2015).
Sena, L. A. et al. Mitochondria are required for antigen-specific T cell activation through reactive oxygen species signaling. Immunity 38, 225–236 (2013).
MacIver, N. J., Michalek, R. D. & Rathmell, J. C. Metabolic regulation of T lymphocytes. Annu. Rev. Immunol. 31, 259–283 (2013).
van der Windt, G. J. et al. CD8 memory T cells have a bioenergetic advantage that underlies their rapid recall ability. Proc. Natl Acad. Sci. USA 110, 14336–14341 (2013).
Schurich, A. et al. Distinct metabolic requirements of exhausted and functional virus-specific CD8 T cells in the same host. Cell Rep. 16, 1243–1252 (2016).
Sukumar, M. et al. Inhibiting glycolytic metabolism enhances CD8+ T cell memory and antitumor function. J. Clin. Invest. 123, 4479–4488 (2013).
Wang, Y. et al. Autocrine complement inhibits IL10-dependent T-cell-mediated antitumor immunity to promote tumor progression. Cancer Discov. 6, 1022–1035 (2016).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
He, R. et al. Follicular CXCR5- expressing CD8+ T cells curtail chronic viral infection. Nature 537, 412–428 (2016).
Ansel, K. M. et al. A chemokine-driven positive feedback loop organizes lymphoid follicles. Nature 406, 309–314 (2000).
Zou, W., Wolchok, J. D. & Chen, L. PD-L1 (B7-H1) and PD-1 pathway blockade for cancer therapy: mechanisms, response biomarkers, and combinations. Sci. Transl. Med. 8, 328rv4 (2016).
Gu-Trantien, C. et al. CXCL13-producing TFH cells link immune suppression and adaptive memory in human breast cancer. JCI Insight 2, 91487 (2017).
Gunnlaugsdottir, B., Maggadottir, S. M. & Ludviksson, B. R. Anti-CD28-induced co-stimulation and TCR avidity regulates the differential effect of TGF-beta1 on CD4+ and CD8+ naïve human T-cells. Int. Immunol. 17, 35–44 (2005).
Allen, C. D. et al. Germinal center dark and light zone organization is mediated by CXCR4 and CXCR5. Nat. Immunol. 5, 943–952 (2004).
Wu, T. D., Reeder, J., Lawrence, M., Becker, G. & Brauer, M. J. GMAP and GSNAP for genomic sequence alignment: enhancements to speed, accuracy, and functionality. Methods Mol. Biol. 1418, 283–334 (2016).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Becker, R. A., Chambers, J. M. & Wilks, A. R. The New S Language (Wadsworth & Brooks/Cole, Monterey, CA, USA, 1988).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2007).
Nazarov, V. I. et al. tcR: an R package for T cell receptor repertoire advanced data analysis. BMC Bioinformatics 16, 175 (2015).
Acknowledgements
We thank D. Labes and E. Traunecker for exemplary technical assistance with cell sorting, F. Franco and T. Chao for performing electron microscopy analysis, L. Tietze (Ortenau Klinikum, Germany) for contribution of tumor samples, B. Dolder-Schlienger for technical assistance, and F. Uhlenbrock and D. Pinschewer for discussions and critical reading of the manuscript. This work was supported by grants from the Swiss National Science Foundation (P300PB_164755 to D.S.T., 320030_162575 to A.Z. and 31003A_163204 to P.C.H.), the Research Funds University of Basel (D.S.T.), the Lichtenstein-Stiftung (D.S.T.), the FAG-Basel (D.S.T.), the Dutch Cancer Society Queen Wilhelmina Award NKI 2013-6122 (T.N.S.) and ERC grant SENSIT (T.N.S.).
Author information
Authors and Affiliations
Contributions
D.S.T.: study design and supervision, design and execution of the experiments; data acquisition, analysis and interpretation; writing and revision of the manuscript; V.H.K.: execution of immunohistochemistry stainings, digital image analysis; contribution to manuscript drafting and revision; P.H.: execution of experiments; contribution to manuscript drafting and revision; A.R.: statistical analysis and interpretation, contribution to manuscript drafting; M.T.: execution of experiments; A.K.: RNA-seq analysis; S.D.: design and technical support with metabolism analysis; J.H.: execution of immunohistochemistry and digital image analysis; C.S.: collection and analysis of clinical data; C.H.: design of metabolism experiments, contribution to manuscript drafting; S.S.P.: collection and pathological characterization of patient samples; M.W. and D.L.: recruitment and characterization of patients; P.C.H.: execution of experiments, contribution to manuscript drafting; C.K. and V.K.: contribution to manuscript drafting; K.D.M.: execution of immunohistochemistry analysis; contribution to manuscript drafting; T.N.S.: study design and supervision; writing and revision of the manuscript; A.Z.: study design and supervision, writing and revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.R., A.K., C.K., V.K. are employed by Roche. A.Z. received research funding from Roche. Part of the work described in this manuscript is the subject of a patent application co-owned by NKI-AVL and the University of Basel. Based on NKI-AVL and the University of Basel policy on management of intellectual property, D.S.T., V.H.K., K.D.M., A.Z. and T.N.S. would be entitled to a portion of the royalty income received.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Figures
Supplementary Figures 1–6
Supplementary Table 1
TCR analysis data
Supplementary Table 2
Gene expression data
Supplementary Table 3
Tumor sample overview
Supplementary Table 4
Patient characteristics for predictive analysis
Rights and permissions
About this article
Cite this article
Thommen, D.S., Koelzer, V.H., Herzig, P. et al. A transcriptionally and functionally distinct PD-1+ CD8+ T cell pool with predictive potential in non-small-cell lung cancer treated with PD-1 blockade. Nat Med 24, 994–1004 (2018). https://doi.org/10.1038/s41591-018-0057-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0057-z
This article is cited by
-
Therapeutic and immunomodulatory potentials of mesenchymal stromal/stem cells and immune checkpoints related molecules
Biomarker Research (2024)
-
Single-cell RNA sequencing reveals myeloid and T cell co-stimulation mediated by IL-7 anti-cancer immunotherapy
British Journal of Cancer (2024)
-
Tumour-infiltrating lymphocyte therapy for patients with advanced-stage melanoma
Nature Reviews Clinical Oncology (2024)
-
Clinical and immunological relevance of SLAMF6 expression in the tumor microenvironment of breast cancer and melanoma
Scientific Reports (2024)
-
Multi-resolution deep learning characterizes tertiary lymphoid structures and their prognostic relevance in solid tumors
Communications Medicine (2024)