Abstract
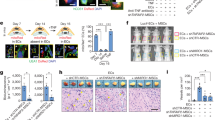
Recent studies have identified a specialized subset of CD31hiendomucinhi (CD31hiEMCNhi) vascular endothelium that positively regulates bone formation. However, it remains unclear how CD31hiEMCNhi endothelium levels are coupled to anabolic bone formation. Mice with an osteoblast-specific deletion of Shn3, which have markedly elevated bone formation, demonstrated an increase in CD31hiEMCNhi endothelium. Transcriptomic analysis identified SLIT3 as an osteoblast-derived, SHN3-regulated proangiogenic factor. Genetic deletion of Slit3 reduced skeletal CD31hiEMCNhi endothelium, resulted in low bone mass because of impaired bone formation and partially reversed the high bone mass phenotype of Shn3−/− mice. This coupling between osteoblasts and CD31hiEMCNhi endothelium is essential for bone healing, as shown by defective fracture repair in SLIT3-mutant mice and enhanced fracture repair in SHN3-mutant mice. Finally, administration of recombinant SLIT3 both enhanced bone fracture healing and counteracted bone loss in a mouse model of postmenopausal osteoporosis. Thus, drugs that target the SLIT3 pathway may represent a new approach for vascular-targeted osteoanabolic therapy to treat bone loss.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
29 May 2018
In the version of this article originally published, the Supplementary Information provided by the authors was not uploaded. The full version of the Supplementary Information now appears correctly.
References
Kusumbe, A. P., Ramasamy, S. K. & Adams, R. H. Coupling of angiogenesis and osteogenesis by a specific vessel subtype in bone. Nature 507, 323–328 (2014).
Ramasamy, S. K., Kusumbe, A. P., Wang, L. & Adams, R. H. Endothelial Notch activity promotes angiogenesis and osteogenesis in bone. Nature 507, 376–380 (2014).
Xie, H. et al. PDGF-BB secreted by preosteoclasts induces angiogenesis during coupling with osteogenesis. Nat. Med. 20, 1270–1278 (2014).
Shim, J. H. et al. Schnurri-3 regulates ERK downstream of WNT signaling in osteoblasts. J. Clin. Invest. 123, 4010–4022 (2013).
Jones, D. C. et al. Regulation of adult bone mass by the zinc finger adapter protein Schnurri-3. Science 312, 1223–1227 (2006).
Nguyen Ba-Charvet, K. T. et al. Slit2-Mediated chemorepulsion and collapse of developing forebrain axons. Neuron 22, 463–473 (1999).
Jaworski, A. & Tessier-Lavigne, M. Autocrine/juxtaparacrine regulation of axon fasciculation by Slit-Robo signaling. Nat. Neurosci. 15, 367–369 (2012).
Jones, C. A. et al. Robo4 stabilizes the vascular network by inhibiting pathologic angiogenesis and endothelial hyperpermeability. Nat. Med. 14, 448–453 (2008).
Rama, N. et al. Slit2 signaling through Robo1 and Robo2 is required for retinal neovascularization. Nat. Med. 21, 483–491 (2015).
Vasam, G. et al. Reversal of bone marrow mobilopathy and enhanced vascular repair by angiotensin-(1-7) in diabetes. Diabetes 66, 505–518 (2017).
Geutskens, S. B. et al. Control of human hematopoietic stem/progenitor cell migration by the extracellular matrix protein Slit3. Lab. Invest. 92, 1129–1139 (2012).
Zhou, W. J., Geng, Z. H., Spence, J. R. & Geng, J. G. Induction of intestinal stem cells by R-spondin 1 and Slit2 augments chemoradioprotection. Nature 501, 107–111 (2013).
Mehlen, P., Delloye-Bourgeois, C. & Chedotal, A. Novel roles for Slits and netrins: axon guidance cues as anticancer targets? Nat. Rev. Cancer 11, 188–197 (2011).
Zhang, B. et al. Repulsive axon guidance molecule Slit3 is a novel angiogenic factor. Blood 114, 4300–4309 (2009).
Paul, J. D. et al. SLIT3-ROBO4 activation promotes vascular network formation in human engineered tissue and angiogenesis in vivo. J. Mol. Cell Cardiol. 64, 124–131 (2013).
Zhang, J. & Link, D. C. Targeting of mesenchymal stromal cells by Cre-recombinase transgenes commonly used to target osteoblast lineage cells. J. Bone Miner. Res. 31, 2001–2007 (2016).
Chen, J. et al. Osx-Cre targets multiple cell types besides osteoblast lineage in postnatal mice. PLoS ONE 9, e85161 (2014).
Su, A. I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl Acad. Sci. USA 101, 6062–6067 (2004).
Yuan, W. et al. A genetic model for a central (septum transversum) congenital diaphragmatic hernia in mice lacking Slit3. Proc. Natl Acad. Sci. USA 100, 5217–5222 (2003).
Geutskens, S. B., Hordijk, P. L. & van Hennik, P. B. The chemorepellent Slit3 promotes monocyte migration. J. Immunol. 185, 7691–7698 (2010).
Naska, S., Lin, D. C., Miller, F. D. & Kaplan, D. R. p75NTR is an obligate signaling receptor required for cues that cause sympathetic neuron growth cone collapse. Mol. Cell Neurosci. 45, 108–120 (2010).
Howitt, J. A., Clout, N. J. & Hohenester, E. Binding site for Robo receptors revealed by dissection of the leucine-rich repeat region of Slit. EMBO J. 23, 4406–4412 (2004).
Liu, Z. et al. Extracellular Ig domains 1 and 2 of Robo are important for ligand (Slit) binding. Mol. Cell Neurosci. 26, 232–240 (2004).
Shin, M. et al. Vegfa signals through ERK to promote angiogenesis, but not artery differentiation. Development 143, 3796–3805 (2016).
Choi, H. J. et al. Yes-associated protein regulates endothelial cell contact-mediated expression of angiopoietin-2. Nat. Commun. 6, 6943 (2015).
Blockus, H. & Chedotal, A. Slit-Robo signaling. Development 143, 3037–3044 (2016).
Sakabe, M. et al. YAP/TAZ-CDC42 signaling regulates vascular tip cell migration. Proc. Natl Acad. Sci. USA 114, 10918–10923 (2017).
Bouxsein, M. L. et al. Ovariectomy-induced bone loss varies among inbred strains of mice. J. Bone Miner. Res. 20, 1085–1092 (2005).
Fukuda, T. et al. Sema3A regulates bone-mass accrual through sensory innervations. Nature 497, 490–493 (2013).
Zhang, Y. et al. Implant-derived magnesium induces local neuronal production of CGRP to improve bone-fracture healing in rats. Nat. Med. 22, 1160–1169 (2016).
Xu, R. Semaphorin 3A: a new player in bone remodeling. Cell Adh. Migr. 8, 5–10 (2014).
Ramalingam, P., Poulos, M. G. & Butler, J. M. Regulation of the hematopoietic stem cell lifecycle by the endothelial niche. Curr. Opin. Hematol. 24, 289–299 (2017).
Buza, J. A. 3rd & Einhorn, T. Bone healing in 2016. Clin. Cases Miner. Bone Metab. 13, 101–105 (2016).
Sun, H., Dai, K., Tang, T. & Zhang, X. Regulation of osteoblast differentiation by slit2 in osteoblastic cells. Cells Tissues Organs 190, 69–80 (2009).
Leder, B. Z. et al. Denosumab and teriparatide transitions in postmenopausal osteoporosis (the DATA-Switch study): extension of a randomised controlled trial. Lancet 386, 1147–1155 (2015).
Wein, M. N. et al. Control of bone resorption in mice by Schnurri-3. Proc. Natl Acad. Sci. USA 109, 8173–8178 (2012).
Chen, M. J., Yokomizo, T., Zeigler, B. M., Dzierzak, E. & Speck, N. A. Runx1 is required for the endothelial to haematopoietic cell transition but not thereafter. Nature 457, 887–891 (2009).
Rodda, S. J. & McMahon, A. P. Distinct roles for Hedgehog and canonical Wnt signaling in specification, differentiation and maintenance of osteoblast progenitors. Development 133, 3231–3244 (2006).
Lu, Y. et al. DMP1-targeted Cre expression in odontoblasts and osteocytes. J. Dent. Res. 86, 320–325 (2007).
Park, D. et al. Endogenous bone marrow MSCs are dynamic, fate-restricted participants in bone maintenance and regeneration. Cell Stem Cell 10, 259–272 (2012).
Xu, R. et al. c-Jun N-terminal kinases (JNKs) are critical mediators of osteoblast activity in vivo. J. Bone Miner. Res. 32, 1811–1815 (2017).
Tual-Chalot, S., Allinson, K.R., Fruttiger, M. & Arthur, H.M. Whole mount immunofluorescent staining of the neonatal mouse retina to investigate angiogenesis in vivo. J. Vis. Exp. 77, e50546 (2013).
Dempster, D. W. et al. Standardized nomenclature, symbols, and units for bone histomorphometry: a 2012 update of the report of the ASBMR Histomorphometry Nomenclature Committee. J. Bone Miner. Res. 28, 2–17 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Zhang, C. et al. Mosaic: making biological sense of complex networks. Bioinformatics 28, 1943–1944 (2012).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Greenblatt, M. B. et al. MEKK2 mediates an alternative beta-catenin pathway that promotes bone formation. Proc. Natl Acad. Sci. USA 113, E1226–E1235 (2016).
Bradaschia-Correa, V. et al. The selective serotonin reuptake inhibitor fluoxetine directly inhibits osteoblast differentiation and mineralization during fracture healing in mice. J. Bone Miner. Res. 32, 821–833 (2017).
Han, W. et al. The osteogenic potential of human bone callus. Sci. Rep. 6, 36330 (2016).
Acknowledgements
We thank the many individuals who provided valuable reagents. M.B.G. holds a Career Award for Medical Scientists from the Burroughs Wellcome Foundation and is supported by the Office of the Director of the NIH under award DP5OD021351, a Junior Investigator Award from the Musculoskeletal Transplant Foundation and a March of Dimes Basil O’Connor Award. J.-H.S. is supported by National Institute of Arthritis and Musculoskeletal and Skin Diseases/NIH under R01AR068983 and a pilot project program award from UMass Center for Clinical and Translational Science. A.D.L. is supported by NIH/National Heart, Lung, and Blood Institute under R01 HL126913. This content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH. We thank D. Ballon, B. He, B. -S. Ding and J. McCormick and Citigroup Biomedical Imaging Core, Weill Cornell Microscopy and Image Analysis and Flow Cytometry Core Facilities for technical support.
Author information
Authors and Affiliations
Contributions
R.X. and M.B.G. designed the experimental plan. R.X. executed most experiments. A.Y., Z.W., X.Y., P.K., N.L., Y.L., A.W., Y.Z., A.D.L., J.-H.S., J.M.B. and K.I. assisted with mouse studies. D.Y.S., J.-M.K., S.L. and B.Z. conducted in vitro experiments. S.D., G.J., H.D. and M.G.P. assisted with flow cytometry analysis and cell sorting. A.Q. collected human bone fracture samples. C.Z. performed RNA-seq analysis. M.P.B., B.Z. and J.-H.S. assisted with osteoclast studies. R.X. and M.B.G wrote the manuscript. M.B.G and L.H.G. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
L.H.G. is on the board of directors of and holds equity in the GlaxoSmithKline and Waters Corporations. She is also a founder of Quentis Pharmaceuticals.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13 and Supplementary Table 1
Rights and permissions
About this article
Cite this article
Xu, R., Yallowitz, A., Qin, A. et al. Targeting skeletal endothelium to ameliorate bone loss. Nat Med 24, 823–833 (2018). https://doi.org/10.1038/s41591-018-0020-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0020-z