Abstract
How lipidome changes support CD8+ effector T (Teff) cell differentiation is not well understood. Here we show that, although naive T cells are rich in polyunsaturated phosphoinositides (PIPn with 3–4 double bonds), Teff cells have unique PIPn marked by saturated fatty acyl chains (0–2 double bonds). PIPn are precursors for second messengers. Polyunsaturated phosphatidylinositol bisphosphate (PIP2) exclusively supported signaling immediately upon T cell antigen receptor activation. In late Teff cells, activity of phospholipase C-γ1, the enzyme that cleaves PIP2 into downstream mediators, waned, and saturated PIPn became essential for sustained signaling. Saturated PIP was more rapidly converted to PIP2 with subsequent recruitment of phospholipase C-γ1, and loss of saturated PIPn impaired Teff cell fitness and function, even in cells with abundant polyunsaturated PIPn. Glucose was the substrate for de novo PIPn synthesis, and was rapidly utilized for saturated PIP2 generation. Thus, separate PIPn pools with distinct acyl chain compositions and metabolic dependencies drive important signaling events to initiate and then sustain effector function during CD8+ T cell differentiation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
RNA sequencing data were accessed in the Gene Expression Omnibus under accession number GSE17124559. Source data are provided with this paper.
References
Waickman, A. T. & Powell, J. D. mTOR, metabolism, and the regulation of T cell differentiation and function. Immunol. Rev. 249, 43–58 (2012).
MacIver, N. J., Michalek, R. D. & Rathmell, J. C. Metabolic regulation of T lymphocytes. Annu Rev. Immunol. 31, 259–283 (2013).
Shyer, J. A., Flavell, R. A. & Bailis, W. Metabolic signaling in T cells. Cell Res 30, 649–659 (2020).
Carr, E. L. et al. Glutamine uptake and metabolism are coordinately regulated by ERK/MAPK during T lymphocyte activation. J. Immunol. 185, 1037–1044 (2010).
Wang, R. et al. The transcription factor Myc controls metabolic reprogramming upon T lymphocyte activation. Immunity 35, 871–882 (2011).
Maciver, N. J. et al. Glucose metabolism in lymphocytes is a regulated process with significant effects on immune cell function and survival. J. Leukoc. Biol. 84, 949–957 (2008).
Cham, C. M. & Gajewski, T. F. Glucose availability regulates IFN-γ production and p70S6 kinase activation in CD8+ effector T cells. J. Immunol. 174, 4670–4677 (2005).
Barneda, D., Cosulich, S., Stephens, L. & Hawkins, P. How is the acyl chain composition of phosphoinositides created and does it matter? Biochem. Soc. Trans. 47, 1291–1305 (2019).
Sasaki, T. et al. Mammalian phosphoinositide kinases and phosphatases. Prog. Lipid Res. 48, 307–343 (2009).
Czech, M. P. PIP2 and PIP3: complex roles at the cell surface. Cell 100, 603–606 (2000).
Huang, Y. H. & Sauer, K. Lipid signaling in T cell development and function. Cold Spring Harb. Perspect. Biol. 2, a002428 (2010).
Berridge, M. J. Inositol trisphosphate and calcium signalling mechanisms. Biochim. Biophys. Acta 1793, 933–940 (2009).
D’Souza, W. N., Chang, C. F., Fischer, A. M., Li, M. & Hedrick, S. M. The Erk2 MAPK regulates CD8 T cell proliferation and survival. J. Immunol. 181, 7617–7629 (2008).
Joseph, N., Reicher, B. & Barda-Saad, M. The calcium feedback loop and T cell activation: how cytoskeleton networks control intracellular calcium flux. Biochim. Biophys. Acta 1838, 557–568 (2014).
Sun, Y., Dandekar, R. D., Mao, Y. S., Yin, H. L. & Wulfing, C. Phosphatidylinositol (4,5) bisphosphate controls T cell activation by regulating T cell rigidity and organization. PLoS One 6, e27227 (2011).
Vickers, J. D. & Mustard, J. F. The phosphoinositides exist in multiple metabolic pools in rabbit platelets. Biochem. J. 238, 411–417 (1986).
Chakrabarti, P. et al. A dPIP5K dependent pool of phosphatidylinositol 4,5 bisphosphate (PIP2) is required for G-protein-coupled signal transduction in Drosophila photoreceptors. PLoS Genet. 11, e1004948 (2015).
Fujita, A., Cheng, J., Tauchi-Sato, K., Takenawa, T. & Fujimoto, T. A distinct pool of phosphatidylinositol 4,5-bisphosphate in caveolae revealed by a nanoscale labeling technique. Proc. Natl Acad. Sci. USA 106, 9256–9261 (2009).
Costello, P. S., Gallagher, M. & Cantrell, D. A. Sustained and dynamic inositol lipid metabolism inside and outside the immunological synapse. Nat. Immunol. 3, 1082–1089 (2002).
Latour, S. & Fischer, A. Signaling pathways involved in the T cell-mediated immunity against Epstein–Barr virus: lessons from genetic diseases. Immunol. Rev. 291, 174–189 (2019).
Imoto, M., Taniguchi, Y. & Umezawa, K. Inhibition of CDP-DG: inositol transferase by inostamycin. J. Biochem. 112, 299–302 (1992).
Bengsch, B. et al. Analysis of CD127 and KLRG1 expression on hepatitis C virus-specific CD8+ T cells reveals the existence of different memory T cell subsets in the peripheral blood and liver. J. Virol. 81, 945–953 (2007).
Ward, S. G. & Cantrell, D. A. Phosphoinositide 3-kinases in T lymphocyte activation. Curr. Opin. Immunol. 13, 332–338 (2001).
Hawse, W. F. & Cattley, R. T. T cells transduce T cell receptor signal strength by generating different phosphatidylinositols. J. Biol. Chem. 294, 4793–4805 (2019).
Hwang, J. R., Byeon, Y., Kim, D. & Park, S. G. Recent insights of T cell receptor-mediated signaling pathways for T cell activation and development. Exp. Mol. Med. 52, 750–761 (2020).
Janes, P. W., Ley, S. C., Magee, A. I. & Kabouridis, P. S. The role of lipid rafts in T cell antigen receptor (TCR) signalling. Semin. Immunol. 12, 23–34 (2000).
Holmgren, J., Lonnroth, I. & Svennerholm, L. Tissue receptor for cholera exotoxin: postulated structure from studies with GM1 ganglioside and related glycolipids. Infect. Immun. 8, 208–214 (1973).
Waddington, K. E., Pineda-Torra, I. & Jury, E. C. Analyzing T cell plasma membrane lipids by flow cytometry. Methods Mol. Biol. 1951, 209–216 (2019).
Maib, H. & Murray, D. H. A mechanism for exocyst-mediated tethering via Arf6 and PIP5K1C-driven phosphoinositide conversion. Curr. Biol. 32, 2821–2833 (2022).
Zhao, Y., Chen, Y. Q., Li, S., Konrad, R. J. & Cao, G. The microsomal cardiolipin remodeling enzyme acyl-CoA lysocardiolipin acyltransferase is an acyltransferase of multiple anionic lysophospholipids. J. Lipid Res. 50, 945–956 (2009).
Bone, L. N. et al. The acyltransferase LYCAT controls specific phosphoinositides and related membrane traffic. Mol. Biol. Cell 28, 161–172 (2017).
Lee, H. C. et al. Caenorhabditis elegans mboa-7, a member of the MBOAT family, is required for selective incorporation of polyunsaturated fatty acids into phosphatidylinositol. Mol. Biol. Cell 19, 1174–1184 (2008).
van der Windt, G. J. et al. CD8 memory T cells have a bioenergetic advantage that underlies their rapid recall ability. Proc. Natl Acad. Sci. USA 110, 14336–14341 (2013).
Bengsch, B. et al. Bioenergetic insufficiencies due to metabolic alterations regulated by the inhibitory receptor PD-1 are an early driver of CD8+ T cell exhaustion. Immunity 45, 358–373 (2016).
Chang, C. H. et al. Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell 162, 1229–1241 (2015).
Wolchok, J. D. et al. Nivolumab plus ipilimumab in advanced melanoma. N. Engl. J. Med. 369, 122–133 (2013).
Hawse, W. F., Boggess, W. C. & Morel, P. A. TCR signal strength regulates Akt substrate specificity to induce alternate murine Th and T regulatory cell differentiation programs. J. Immunol. 199, 589–597 (2017).
Lee, K. H. et al. T cell receptor signaling precedes immunological synapse formation. Science 295, 1539–1542 (2002).
Lee, K. H. et al. The immunological synapse balances T cell receptor signaling and degradation. Science 302, 1218–1222 (2003).
Traynor-Kaplan, A. et al. Fatty-acyl chain profiles of cellular phosphoinositides. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1862, 513–522 (2017).
Naguib, A. et al. p53 mutations change phosphatidylinositol acyl chain composition. Cell Rep. 10, 8–19 (2015).
van der Windt, G. J. & Pearce, E. L. Metabolic switching and fuel choice during T cell differentiation and memory development. Immunol. Rev. 249, 27–42 (2012).
Frauwirth, K. A. et al. The CD28 signaling pathway regulates glucose metabolism. Immunity 16, 769–777 (2002).
Ho, P. C. et al. Phosphoenolpyruvate is a metabolic checkpoint of anti-tumor T cell responses. Cell 162, 1217–1228 (2015).
Barneda, D. et al. Acyl chain selection couples the consumption and synthesis of phosphoinositides. EMBO J. 41, e110038 (2022).
Shulga, Y. V., Anderson, R. A., Topham, M. K. & Epand, R. M. Phosphatidylinositol-4-phosphate 5-kinase isoforms exhibit acyl chain selectivity for both substrate and lipid activator. J. Biol. Chem. 287, 35953–35963 (2012).
Blunsom, N. J. & Cockcroft, S. Phosphatidylinositol synthesis at the endoplasmic reticulum. Biochim. Biophys. Acta Mol. Cel. Biol. Lipids 1865, 158471 (2020).
D’Souza, K. & Epand, R. M. The phosphatidylinositol synthase-catalyzed formation of phosphatidylinositol does not exhibit acyl chain specificity. Biochemistry 54, 1151–1153 (2015).
Pike, L. J. Lipid rafts: bringing order to chaos. J. Lipid Res. 44, 655–667 (2003).
Myeong, J., Park, C. G., Suh, B. C. & Hille, B. Compartmentalization of phosphatidylinositol 4,5-bisphosphate metabolism into plasma membrane liquid-ordered/raft domains. Proc. Natl Acad. Sci. USA 118, e2025343118 (2021).
Veri, M. C. et al. Membrane raft-dependent regulation of phospholipase Cγ-1 activation in T lymphocytes. Mol. Cell. Biol. 21, 6939–6950 (2001).
Kallikourdis, M. et al. Phosphatidylinositol 4-phosphate 5-kinase beta controls recruitment of lipid rafts into the immunological synapse. J. Immunol. 196, 1955–1963 (2016).
Parry, R. V. et al. CTLA-4 and PD-1 receptors inhibit T cell activation by distinct mechanisms. Mol. Cell. Biol. 25, 9543–9553 (2005).
Callahan, M. K., Postow, M. A. & Wolchok, J. D. Targeting T cell co-receptors for cancer therapy. Immunity 44, 1069–1078 (2016).
Matyash, V., Liebisch, G., Kurzchalia, T. V., Shevchenko, A. & Schwudke, D. Lipid extraction by methyl-tert-butyl ether for high-throughput lipidomics. J. Lipid Res. 49, 1137–1146 (2008).
Wong, M., Xu, G., Park, D., Barboza, M. & Lebrilla, C. B. Intact glycosphingolipidomic analysis of the cell membrane during differentiation yields extensive glycan and lipid changes. Sci. Rep. 8, 10993 (2018).
Clark, J. et al. Quantification of PtdInsP3 molecular species in cells and tissues by mass spectrometry. Nat. Methods 8, 267–272 (2011).
Wills, J., Edwards-Hicks, J. & Finch, A. J. AssayR: a simple mass spectrometry software tool for targeted metabolic and stable isotope tracer analyses. Anal. Chem. 89, 9616–9619 (2017).
O’Sullivan, D. et al. Fever supports CD8+ effector T cell responses by promoting mitochondrial translation. Proc. Natl Acad. Sci. USA 118, e2023752118 (2021).
Bolstad, B. preprocessCore: a collection of pre-processing functions. R package version 1.42.0. https://github.com/bmbolstad/preprocessCore
Acknowledgements
The authors thank A. Shaw and Z. Katz for insightful discussion. This work was supported by the Max Planck Society, the Leibniz Prize, and the National Institutes of Health R01AI156274 (to E.L.P.), two Bloomberg Distinguished Professorships (to E.L.P. and E.J.P.), a research grant from Bristol Myers Squibb, a Marie-Sklodowska-Curie actions Individual Fellowship (to F.B.), a Sir Henry Wellcome Fellowship (to D.J.P.), two Alexander von Humboldt Fellowships (to M.V. and M.C.) and the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation)—project no 492259164 (to P.A.) and SFB-1479 project no. 441891347 (to R.Z.).
Author information
Authors and Affiliations
Contributions
Conceptualization, project insight and data interpretation: J.E.-H., P.A. and E.L.P. Performing and analyzing experiments: J.E.-H., P.A., J.M.B., H.M., M.A.S., M.C., R.I.K., M.V., G.E.C., D.E.S., F.B., B.K., J.D.C., F.H., A.P., C.S.F., G.C., R.L.K., M.S., M.C., H.P., J.M., K.G., L.F., M.M. and L.Z. Provided advice, material and scientific ideas: M.E.M., F.W., D.A.R.-S., F.M., B.B., R.Z., D.J.P., D.O.S. and E.J.P. Manuscript writing: J.E.H., P.A. and E.L.P.
Corresponding author
Ethics declarations
Competing interests
E.L.P. is an SAB member of ImmunoMet Therapeutics and E.L.P. and E.J.P are Founders and Scientific Advisors to Rheos Medicines. The other authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Michael Dustin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: N. Bernard, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Human CD8+ TE accumulate PIPn with saturated acyl chains.
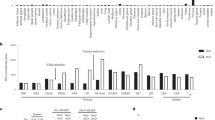
a, Pie chart showing the number of lipid species measured by LC-QqQ-MS/MS in each of the 18 lipid subclasses. b-c, WT CD8+ T cells were stimulated (Stim) with anti-CD3, anti-CD28 and IL-2 or cultured with IL-7 (unstim) for 3 days. b, Median lipid content per million cells before (raw) and after (normalized) quantile normalization is shown. n = 9 biologically independent samples across 3 independent experiments; unpaired two-tailed t-test. c, Relative total PI level calculated from quantile normalized lipid peak areas. n = 9 biologically independent samples across 3 independent experiments; unpaired two-tailed t-test. d, Experimental schematic. e-f, Volcano plot shows the log2 fold change (e) or total PI (f) between Unstim and Stim conditions using quantile normalized lipid peak areas. n = 3 biologically independent samples; one-way ANOVA or two-way ANOVA corrected for multiple comparisons (Sidak test). g, Relative intensity of saturated PI normalized to Unstim (log2 fold change). n = 3 biologically independent samples; one sample t-test. h, PI saturation percentage. n = 3 biologically independent samples; unpaired two-tailed t-test comparing saturated PI. I, Total PI. n = 3 biologically independent samples; unpaired two-tailed t-test. j, Total PI concentration (nmol/cell). n = 3 biologically independent samples; unpaired two-tailed t-test. k, Saturation of acyl chains in human TE is expressed as a relative proportion of the total PIP2 (left panel) and as a percentage of total PIP2 (right panel). n = 5 biologically independent samples; unpaired two-tailed t-test comparing the total or percentage saturated PIP2. Total polyunsaturated PIP2 was not significantly different. l, Saturation of acyl chains in human TE is expressed as a relative proportion of the total PIP3 (left panel) and as a percentage of PIP3 (right panel). n = 5 biologically independent samples; unpaired two-tailed t-test comparing the total or percentage saturated PIP3. Total polyunsaturated PIP3 was also significantly different (p < 0.001). m, Experimental schematic. n, Percentage CD8+ cells, activation marker HLA-DR+ cells, and proliferation marker Ki67+ cells are shown. MFI of HLA-DR and Ki67 are also shown. o, Relative intensity of saturated PI normalized to HD (log2 fold change). n = 3 HD and n = 3 EBV biologically independent samples; one sample t-test. p, PI saturation. n = 3 HD and n = 3 EBV biologically independent samples; unpaired two-tailed t-test comparing the summed percentage of saturated PI. q, Total PI concentration (nmol/cell). n = 3 HD and n = 3 EBV biologically independent samples; unpaired two-tailed t-test. Error bars show the standard error of the mean. a.u., arbitrary units.
Extended Data Fig. 2 CDIPT inhibition or deletion impairs TE fitness and function.
a, Simplified overview of PI de novo synthesis and remodeling. b, WT CD8+ T cells were activated and treated as in Fig. 2a. Total lysophosphatidlyinositol (LPI) content is shown. n = 4 biologically independent samples; unpaired two-tailed t-test. c, Gating strategy for flow cytometry analysis. d, Replication Index based on cell trace violet dilution was calculated using FlowJo software. n = 4 biologically independent samples; unpaired two-tailed t-test. e, PI species present in PI derived from Soy expressed by degree of acyl chain saturation. f-g, Cells were prepared as in (b), with the addition of BSA (10 mg/ml) control or BSA conjugated to Soy-derived PI (SoyPI, 100 μM) for the final day. f, Saturation of acyl chains expressed as a percentage of total PI. n = 3 biologically independent samples, one-way ANOVA corrected for multiple comparisons (Tukey test) comparing saturated PI. g, Percentage IFN-γ+ cells are shown, gated on Live/Dead-aqua- and CD8-Brilliant Violet 421+. n = 3 biologically independent samples; one-way ANOVA corrected for multiple comparisons (Tukey test). h-I, Cells were prepared as in (b), with the addition of BSA (10 mg/ml) control or BSA conjugated to PI 38:4 (100 µM) for the final day. h, Saturation of acyl chains expressed as a percentage of total PI. n = 3 biologically independent samples; unpaired two-tailed t-test comparing saturated PI. I, Percentage IFN-γ+ cells are shown, gated on Live/Dead-near-IR− and CD8-Brilliant Violet 421+. n = 5 biologically independent samples representative of three experiments; one-way ANOVA corrected for multiple comparisons (Tukey test). j, WT CD8+ T cells were activated as in Fig. 2a and the CDIPT inhibitor inostamycin was added at the indicated concentrations for 24 h starting either after either two, four or six days of activation. Upper panel: percentage of viable CD8+ cells. Cells were gated on FSC and SSC and single cells. Middle panel: percentage of EdU+ cells. Cells were gated on Live/Dead-aqua- CD8-APC-Cy7+. Lower panel: percentage of IFN-γ+ cells. Cells were gated on Live/Dead-aqua- CD8-APC-Cy7+. For each time point, the values were normalized to the values of the control samples (0 µg/ml CDIPTi). n = 4 biologically independent samples representative of two independent experiments. k, WT CD8+ T cells were activated and treated as in Fig. 2a. After 24 h of CDIPTi treatment, the cells were cultured in media without CDIPTi for the indicated time intervals. Upper panel: percentage of viable CD8+ cells. Cells were gated on FSC and SSC and single cells. Middle panel: percentage of EdU+ cells. Cells were gated on Live/Dead-aqua- CD8-APC-Cy7+. Lower panel: percentage of IFN-γ+ cells. Cells were gated on Live/Dead-aqua- CD8-APC-Cy7+. For each time point, the values were normalized to the values of the control samples at the respective time point (0 µg/ml CDIPTi). n = 3 biologically independent samples representative of two independent experiments. Error bars show s.e.m.
Extended Data Fig. 3 CRISPR-Cas9 deletion efficiency for CDIPT and ZAP70.
a, The efficiency of CRISPR-Cas9 for CDIPT and ZAP70 was verified by measuring protein expression.
Extended Data Fig. 4 Inhibition of CDIPT changes membrane lipid order and impairs lipid raft formation.
a, Phospholipase C activity in Jurkat T cells cultured in the presence or absence of 0.5 µg/ml CDIPT inhibitor and activated for 30 min with 50 ng/ml PMA and 500 ng/ml ionomycin. n = 5 biologically independent samples representative of 2 independent experiments; unpaired two-tailed t-test. b, WT CD8+ T cells were activated and cellular compartments were enriched as in Fig. 4i. Saturation of acyl chains expressed as a percentage of total PIP2 across organelles. n = 4 biologically independent samples. c, WT CD8+ T cells were isolated and activated or cultured in IL-7 as in Fig. 1a. Volcano plot shows the log2 fold change between Unstim and Stim using quantile normalized lipid peak areas. n = 5 biologically independent samples across 2 independent experiments; one-way ANOVA. Lipids from the following subclasses are highlighted: sphingomyelin (SM) in blue, phosphatidylinositol (PI) in orange, phosphatidylserine (PS) in purple and ganglioside GM1 in green. d, Cholera toxin subunit B measurement on primary CD8+ TE activated and treated as in Fig. 2a. n = 3 biologically independent samples representative of 3 independent experiments; unpaired two-tailed t-test. e, Cholera toxin subunit b measurement on Jurkat T cells treated as in (a). n = 5 biologically independent samples representative of 2 independent experiments; unpaired two-tailed t-test. f, WT CD8+ T cells were activated and treated as in Fig. 2a. Membrane lipid order was assessed by a di-4-ANEPPDHQ staining. Percentage of cells with high lipid order (di-4-ANEPPDHQ emission at 630 nm) and low lipid order (di-4-ANEPPDHQ emission at 570 nm). n = 3 biologically independent samples representative of 3 independent experiments; unpaired two-tailed t-test. g, Percentage of Jurkat T cells with low lipid order as determined by di-4-ANEPPDHQ flow cytometry staining after treatment as in (a). n = 3 biologically independent samples representative of two independent experiments; unpaired two-tailed t-test. (h) Generalized polarization (GP) index as a normalized intensity ratio of the two spectral emissions for CD8+ TE analyzed as in (e). n = 3 biologically independent samples representative of 3 independent experiments; unpaired two-tailed t-test. (I) WT CD8+ T cells were activated and treated as in Fig. 2a except that the CDIPT inhibitor was used at the indicated concentrations. Cells were gated on Live/Dead-aqua- CD8-APC-Cy7+. Left panel: Cholera toxin subunit B binding. Middle panel: mean fluorescence intensity of phosphorylated MEK1. Right panel: Percentage of IFN-γ+ cells. n = 4 biologically independent samples representative of two independent experiments; one-way ANOVA with Dunnett’s multiple comparisons test comparing all groups to ‘0 µg/ml’. Error bars show the standard error of the mean.
Extended Data Fig. 5 TE PIPn synthesis depends on glycolytic metabolism.
a, WT CD8+ T cells were isolated from spleens and lymph nodes of C57BL/6 mice then stimulated with plate-bound anti-CD3 (5 μg/ml), soluble anti-CD28 (0.5 μg/ml) and IL-2 (100 U/ml) for 2 days. Samples for RNA sequencing were harvested every 24 h. Heatmap depicting the gene expression levels of Lclat1 and Mboat7 in n = 3 biologically independent samples. b, WT CD8+ T cells were isolated and activated as in (a), followed by a treatment for 24 h with the Stearoyl-CoA-desaturase inhibitor A939572 (SCDi, 100 nM) in the presence of IL-2. Left panel: Replication Index based on Cell Trace Violet dilution calculated using FlowJo software. Middle and right panel: intracellular expression of IFN-γ. Cells were gated on Live/Dead-IR-, CD8-APC+. n = 3 biologically independent samples; unpaired two-tailed t-test. c, d WT CD8+ T cells were isolated and activated as in (a), followed by culture in IL-2 for 24 h. On d3, cells were either switched to IL-15 (100 U/ml, TM) or maintained in IL-2 until d6. TM were re-activated on d6 by stimulation with plate-bound anti-CD3 (5 μg/ml), soluble anti-CD28 (0.5 μg/ml) and IL-2 (100 U/ml) for 2 days to generate secondary TE. c, Relative amount of PI 36:2 measured every 24 h. n = 3 biologically independent samples; one-way ANOVA corrected for multiple comparisons (Dunnett test) compared all groups to 0 h timepoint. d, Saturation of acyl chains expressed as a proportion of total PIP2 (left panel) and as a percentage of PIP2 (right panel). Statistical analysis of the saturated PI on n = 5 independent biological samples; one-way ANOVA with Dunnett’s multiple comparisons’s test compared all groups to ‘Tn d3’. e, WT CD8+ T cells were differentiated into TM as in (c), with the addition of 100% U-13C-glucose 24 h before lipid analysis at each timepoint. Unstimulated cells were cultured in IL-7 with 100% U-13C-glucose. Fractional contribution of 13C-glucose-derived carbon to PI species was calculated. n = 3 biologically independent samples, two-way ANOVA with correction for multiple comparisons (Sidak’s test) comparing PI species in each group with the relevant 24 h unstimulated PI. Error bars show the standard error of the mean.
Extended Data Fig. 6 Elevated numbers and cytokine production by CD8+ T cells after immunotherapy.
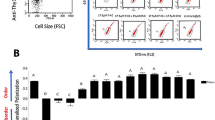
a–d, Sex-matched C57BL/6 mice were injected in the right flank with 1 × 106 B16-F10-OVA cells. After 3 days, mice were given i.p. injections of anti-PD1 and anti-CTLA4 (aPD1 + aCTLA4) or matching isotype controls (IgG1 + IgG2) every 3 days for 3 rounds in total. Tumor growth was measured up to day 16 post-tumor injection and CD8+ T cells were isolated on day 12 post-tumor injection to measure T cell function and lipid composition. a, CD8+ tumor infiltrating lymphocytes (TILs) on day 12 post-tumor injection normalized to tumor weight. n = 7 IgG1 + IgG2 and n = 8 aPD1 + aCTLA4 biologically independent samples; unpaired two-tailed t-test. b–c, TILs were isolated day 12 post-tumor injection and restimulated with PMA/ionomycin plus brefeldin A for 5 h in vitro. n = 7 biologically independent samples per group, unpaired two-tailed t-test. b, Representative contour plots of IFN-γ expression (left panel), percentage IFN-γ+ (middle panel) and IFN-γ expression (right panel) are shown. c, Representative contour plots of Granzyme B expression (left panel), percentage Granzyme B+ (middle panel) and Granzyme B expression (right panel) are shown. d, Relative total PI calculated from quantile normalized lipid peak areas. n = 7 IgG1 + IgG2 and n = 8 aPD1 + aCTLA4 biologically independent samples; unpaired two-tailed t-test. e-f, Sex-matched C57BL/6 mice were injected in the right flank with 1x106 B16-F10-OVA cells. After 3 days, mice were given i.p. injections of anti-PD1 antibody or matching isotype control (IgG2) every 3 days for 3 rounds in total. Tumor growth was measured up to day 12 post-tumor injection and CD8+ T cells were isolated on day 12 post-tumor injection to measure T cell function and lipid composition. e, Average tumor diameter measured at the indicated time points. n = 5 biologically independent samples per group; two-way ANOVA corrected for multiple comparisons (Sidak test) comparing the two groups at each time point across the dataset. f, Lipids were analyzed from TILs on day 12 post-tumor injection. Saturation of acyl chains is expressed as a percentage of total PI. n = 5 biologically independent samples per group; unpaired two-tailed t-test comparing saturated PI. Error bars show the standard error of the mean.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data and unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data and unprocessed western blots.
Source Data Fig. 5
Statistical source data and unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig./Table 1
Statistical source data.
Source Data Extended Data Fig./Table 2
Statistical source data.
Source Data Extended Data Fig./Table 3
Unprocessed western blots.
Source Data Extended Data Fig./Table 4
Statistical source data.
Source Data Extended Data Fig./Table 5
Statistical source data.
Source Data Extended Data Fig./Table 6
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Edwards-Hicks, J., Apostolova, P., Buescher, J.M. et al. Phosphoinositide acyl chain saturation drives CD8+ effector T cell signaling and function. Nat Immunol 24, 516–530 (2023). https://doi.org/10.1038/s41590-023-01419-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-023-01419-y
This article is cited by
-
Metabolic waypoints during T cell differentiation
Nature Immunology (2024)