Abstract
In lupus, Toll-like receptor 7 (TLR7) and TLR9 mediate loss of tolerance to RNA and DNA, respectively. Yet, TLR7 promotes disease, while TLR9 protects from disease, implying differences in signaling. To dissect this ‘TLR paradox’, we generated two TLR9 point mutants (lacking either ligand (TLR9K51E) or MyD88 (TLR9P915H) binding) in lupus-prone MRL/lpr mice. Ameliorated disease of Tlr9K51E mice compared to Tlr9−/− controls revealed a TLR9 ‘scaffold’ protective function that is ligand and MyD88 independent. Unexpectedly, Tlr9P915H mice were more protected than both Tlr9K51E and Tlr9WT mice, suggesting that TLR9 also possesses ligand-dependent, but MyD88-independent, regulatory signaling and MyD88-mediated proinflammatory signaling. Triple-mixed bone marrow chimeras showed that TLR9–MyD88-independent regulatory roles were B cell intrinsic and restrained differentiation into pathogenic age-associated B cells and plasmablasts. These studies reveal MyD88-independent regulatory roles of TLR9, shedding light on the biology of endosomal TLRs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
RNA-seq data of the TLR7-induced gene signature and of the in vitro TLR7-stimulated BALB/c B cells are deposited in the NCBI’s Gene Expression Omnibus (GEO) database and are publicly available under accession numbers GSE202108 and GSE202105.
All ATAC-seq data have been deposited in the NCBI GEO database and are publicly available under accession number GSE202103.
RNA-seq data of the sorted FO cells, MZ cells and ABCs across Tlr9 genotypes from the three-way bone marrow chimeric mice are deposited in the NCBI’s GEO and are publicly available under accession number GSE181283.
The mm10 genome database (https://www.ncbi.nlm.nih.gov/assembly/GCF_000001635.20/) was used to align sequences for the RNA-seq analysis and to align sequencing reads for the ATAC-seq analysis. Source data are provided with this paper.
References
Tsokos, G. C., Lo, M. S., Costa Reis, P. & Sullivan, K. E. New insights into the immunopathogenesis of systemic lupus erythematosus. Nat. Rev. Rheumatol. 12, 716–730 (2016).
Marshak-Rothstein, A. Toll-like receptors in systemic autoimmune disease. Nat. Rev. Immunol. 6, 823–835 (2006).
Teichmann, L. L., Schenten, D., Medzhitov, R., Kashgarian, M. & Shlomchik, M. J. Signals via the adaptor MyD88 in B cells and DCs make distinct and synergistic contributions to immune activation and tissue damage in lupus. Immunity 38, 528–540 (2013).
Christensen, S. R. et al. Toll-like receptor 7 and TLR9 dictate autoantibody specificity and have opposing inflammatory and regulatory roles in a murine model of lupus. Immunity 25, 417–428 (2006).
Bossaller, L. et al. TLR9 deficiency leads to accelerated renal disease and myeloid lineage abnormalities in pristane-induced murine lupus. J. Immunol. 197, 1044–1053 (2016).
Fairhurst, A. M. et al. Yaa autoimmune phenotypes are conferred by overexpression of TLR7. Eur. J. Immunol. 38, 1971–1978 (2008).
Nickerson, K. M., Wang, Y., Bastacky, S. & Shlomchik, M. J. Toll-like receptor 9 suppresses lupus disease in Fas-sufficient MRL mice. PLoS ONE 12, e0173471 (2017).
Jackson, S. W. et al. Opposing impact of B cell-intrinsic TLR7 and TLR9 signals on autoantibody repertoire and systemic inflammation. J. Immunol. 192, 4525–4532 (2014).
Scofield, R. H. et al. Klinefelter’s syndrome (47,XXY) in male systemic lupus erythematosus patients: support for the notion of a gene-dose effect from the X chromosome. Arthritis Rheum. 58, 2511–2517 (2008).
Lee, Y. H., Choi, S. J., Ji, J. D. & Song, G. G. Association between Toll-like receptor polymorphisms and systemic lupus erythematosus: a meta-analysis update. Lupus 25, 593–601 (2016).
Garcia-Ortiz, H. et al. Association of TLR7 copy number variation with susceptibility to childhood-onset systemic lupus erythematosus in Mexican population. Ann. Rheum. Dis. 69, 1861–1865 (2010).
Brown, G. J. et al. TLR7 gain-of-function genetic variation causes human lupus. Nature 605, 349–356 (2022).
Nickerson, K. M. et al. TLR9 regulates TLR7- and MyD88-dependent autoantibody production and disease in a murine model of lupus. J. Immunol. 184, 1840–1848 (2010).
Kim, Y. M., Brinkmann, M. M., Paquet, M. E. & Ploegh, H. L. UNC93B1 delivers nucleotide-sensing Toll-like receptors to endolysosomes. Nature 452, 234–238 (2008).
Fukui, R. et al. Unc93B1 biases Toll-like receptor responses to nucleic acid in dendritic cells toward DNA- but against RNA-sensing. J. Exp. Med. 206, 1339–1350 (2009).
Fukui, R. et al. Unc93B1 restricts systemic lethal inflammation by orchestrating Toll-like receptor 7 and 9 trafficking. Immunity 35, 69–81 (2011).
Stoehr, A. D. et al. TLR9 in peritoneal B-1b cells is essential for production of protective self-reactive IgM to control TH17 cells and severe autoimmunity. J. Immunol. 187, 2953–2965 (2011).
Tilstra, J. S. et al. B cell-intrinsic TLR9 expression is protective in murine lupus. J. Clin. Invest. 130, 3172–3187 (2020).
Peter, M. E., Kubarenko, A. V., Weber, A. N. & Dalpke, A. H. Identification of an N-terminal recognition site in TLR9 that contributes to CpG-DNA-mediated receptor activation. J. Immunol. 182, 7690–7697 (2009).
Christensen, S. R. et al. Toll-like receptor 9 controls anti-DNA autoantibody production in murine lupus. J. Exp. Med. 202, 321–331 (2005).
Nickerson, K. M., Cullen, J. L., Kashgarian, M. & Shlomchik, M. J. Exacerbated autoimmunity in the absence of TLR9 in MRL.Faslpr mice depends on Ifnar1. J. Immunol. 190, 3889–3894 (2013).
Poltorak, A. et al. Defective LPS signaling in C3H/HeJ and C57BL/10ScCr mice: mutations in Tlr4 gene. Science 282, 2085–2088 (1998).
Syrett, C. M., Sierra, I., Beethem, Z. T., Dubin, A. H. & Anguera, M. C. Loss of epigenetic modifications on the inactive X chromosome and sex-biased gene expression profiles in B cells from NZB/W F1 mice with lupus-like disease. J. Autoimmun. 107, 102357 (2020).
Souyris, M. et al. TLR7 escapes X chromosome inactivation in immune cells. Sci. Immunol. 3, eaap8855 (2018).
Azulay-Debby, H., Edry, E. & Melamed, D. CpG DNA stimulates autoreactive immature B cells in the bone marrow. Eur. J. Immunol. 37, 1463–1475 (2007).
Nickerson, K. M. et al. TLR9 promotes tolerance by restricting survival of anergic anti-DNA B cells, yet is also required for their activation. J. Immunol. 190, 1447–1456 (2013).
Scharer, C. D. et al. Epigenetic programming underpins B cell dysfunction in human SLE. Nat. Immunol. 20, 1071–1082 (2019).
Wang, S. et al. IL-21 drives expansion and plasma cell differentiation of autoreactive CD11chiT-bet+ B cells in SLE. Nat. Commun. 9, 1758 (2018).
Saelee, P., Kearly, A., Nutt, S. L. & Garrett-Sinha, L. A. Genome-wide identification of target genes for the key B cell transcription factor Ets1. Front Immunol. 8, 383 (2017).
Kikuchi, H., Nakayama, M., Takami, Y., Kuribayashi, F. & Nakayama, T. EBF1 acts as a powerful repressor of Blimp-1 gene expression in immature B cells. Biochem. Biophys. Res. Commun. 422, 780–785 (2012).
Schmidlin, H. et al. Spi-B inhibits human plasma cell differentiation by repressing BLIMP1 and XBP-1 expression. Blood 112, 1804–1812 (2008).
Willis, S. N. et al. Environmental sensing by mature B cells is controlled by the transcription factors PU.1 and SpiB. Nat. Commun. 8, 1426 (2017).
Carotta, S. et al. The transcription factors IRF8 and PU.1 negatively regulate plasma cell differentiation. J. Exp. Med. 211, 2169–2181 (2014).
Garrett-Sinha, L. A., Kearly, A. & Satterthwaite, A. B. The role of the transcription factor Ets1 in lupus and other autoimmune diseases. Crit. Rev. Immunol. 36, 485–510 (2016).
Thomsen, I. et al. RUNX1 regulates a transcription program that affects the dynamics of cell cycle entry of naive resting B cells. J. Immunol. 207, 2976–2991 (2021).
Emslie, D. et al. Oct2 enhances antibody-secreting cell differentiation through regulation of IL-5 receptor α chain expression on activated B cells. J. Exp. Med. 205, 409–421 (2008).
Scharer, C. D., Barwick, B. G., Guo, M., Bally, A. P. R. & Boss, J. M. Plasma cell differentiation is controlled by multiple cell division-coupled epigenetic programs. Nat. Commun. 9, 1698 (2018).
Goel, R. R. et al. Interferon λ promotes immune dysregulation and tissue inflammation in TLR7-induced lupus. Proc. Natl Acad. Sci. USA 117, 5409–5419 (2020).
Syedbasha, M. et al. Interferon-λ enhances the differentiation of naive B cells into plasmablasts via the mTORC1 pathway. Cell Rep. 33, 108211 (2020).
Huang, W., Ghisletti, S., Perissi, V., Rosenfeld, M. G. & Glass, C. K. Transcriptional integration of TLR2 and TLR4 signaling at the NCoR derepression checkpoint. Mol. Cell 35, 48–57 (2009).
Pascual, G. et al. A SUMOylation-dependent pathway mediates transrepression of inflammatory response genes by PPAR-γ. Nature 437, 759–763 (2005).
Setoguchi, K. et al. Peroxisome proliferator-activated receptor-γ haploinsufficiency enhances B cell proliferative responses and exacerbates experimentally induced arthritis. J. Clin. Invest. 108, 1667–1675 (2001).
Barish, G. D. et al. Bcl-6 and NF-κB cistromes mediate opposing regulation of the innate immune response. Genes Dev. 24, 2760–2765 (2010).
Xu, F. et al. Bcl6 sets a threshold for antiviral signaling by restraining IRF7 transcriptional program. Sci. Rep. 6, 18778 (2016).
Tan, C. et al. NR4A nuclear receptors restrain B cell responses to antigen when second signals are absent or limiting. Nat. Immunol. 21, 1267–1279 (2020).
Carroll, M. C. A protective role for innate immunity in systemic lupus erythematosus. Nat. Rev. Immunol. 4, 825–831 (2004).
Nakano-Yokomizo, T. et al. The immunoreceptor adapter protein DAP12 suppresses B lymphocyte-driven adaptive immune responses. J. Exp. Med. 208, 1661–1671 (2011).
Matsumura, T., Kawamura-Tsuzuku, J., Yamamoto, T., Semba, K. & Inoue, J. TRAF-interacting protein with a forkhead-associated domain B (TIFAB) is a negative regulator of the TRAF6-induced cellular functions. J. Biochem. 146, 375–381 (2009).
Mashima, R., Hishida, Y., Tezuka, T. & Yamanashi, Y. The roles of Dok family adapters in immunoreceptor signaling. Immunol. Rev. 232, 273–285 (2009).
Nakamura, A. et al. Increased susceptibility to LPS-induced endotoxin shock in secretory leukoprotease inhibitor (SLPI)-deficient mice. J. Exp. Med. 197, 669–674 (2003).
Sancho, D. & Reis e Sousa, C. Signaling by myeloid C-type lectin receptors in immunity and homeostasis. Annu. Rev. Immunol. 30, 491–529 (2012).
Komai, T. et al. Transforming growth factor-β and interleukin-10 synergistically regulate humoral immunity via modulating metabolic signals. Front. Immunol. 9, 1364 (2018).
van der Vuurst de Vries, A. R., Clevers, H., Logtenberg, T. & Meyaard, L. Leukocyte-associated immunoglobulin-like receptor-1 (LAIR-1) is differentially expressed during human B cell differentiation and inhibits B cell receptor-mediated signaling. Eur. J. Immunol. 29, 3160–3167 (1999).
Colombo, B. M. et al. Defective expression and function of the leukocyte associated Ig-like receptor 1 in B lymphocytes from systemic lupus erythematosus patients. PLoS ONE 7, e31903 (2012).
Silva, R. et al. CD300a is expressed on human B cells, modulates BCR-mediated signaling, and its expression is down-regulated in HIV infection. Blood 117, 5870–5880 (2011).
Pelka, K. et al. The chaperone UNC93B1 regulates Toll-like receptor stability independently of endosomal TLR transport. Immunity 48, 911–922 (2018).
Nundel, K. et al. Cell-intrinsic expression of TLR9 in autoreactive B cells constrains BCR/TLR7-dependent responses. J. Immunol. 194, 2504–2512 (2015).
Celhar, T. et al. Toll-like receptor 9 deficiency breaks tolerance to RNA-associated antigens and up-regulates Toll-like receptor 7 protein in Sle1 mice. Arthritis Rheumatol. 70, 1597–1609 (2018).
ten Broeke, T., Wubbolts, R. & Stoorvogel, W. MHC class II antigen presentation by dendritic cells regulated through endosomal sorting. Cold Spring Harb. Perspect. Biol. 5, a016873 (2013).
Akkaya, M. et al. Toll-like receptor 9 antagonizes antibody affinity maturation. Nat. Immunol. 19, 255–266 (2018).
Sindhava, V. J. et al. A TLR9-dependent checkpoint governs B cell responses to DNA-containing antigens. J. Clin. Invest. 127, 1651–1663 (2017).
Sanjuan, M. A. et al. Toll-like receptor signalling in macrophages links the autophagy pathway to phagocytosis. Nature 450, 1253–1257 (2007).
Dinarello, C. A. Overview of the IL-1 family in innate inflammation and acquired immunity. Immunol. Rev. 281, 8–27 (2018).
Pelletier, S., Gingras, S. & Green, D. R. Mouse genome engineering via CRISPR–Cas9 for study of immune function. Immunity 42, 18–27 (2015).
Teichmann, L. L. et al. B cell-derived IL-10 does not regulate spontaneous systemic autoimmunity in MRL.Faslpr mice. J. Immunol. 188, 678–685 (2012).
Blanco, F. K., Isenberg, J. & Analysis, D. A. of antibodies to RNA in patients with systemic lupus erythematosus and other autoimmune rheumatic diseases. Clin. Exp. Immunol. 86, 66–70 (1991).
Murakami, Y. et al. The protective effect of the anti-Toll-like receptor 9 antibody against acute cytokine storm caused by immunostimulatory DNA. Sci. Rep. 7, 44042 (2017).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Elsner, R. A. & Shlomchik, M. J. IL-12 blocks TFH cell differentiation during salmonella infection, thereby contributing to germinal center suppression. Cell Rep. 29, 2796–2809 (2019).
Acknowledgements
We thank A. Marshak-Rothstein (University of Massachusetts School of Medicine) and J. Tilstra for helpful discussions. We thank L. Conter, K. Minjung, M. Price and T. Marinov for technical support. We thank S. Watkins for advice on confocal analysis and Center for Biologic Imaging (CBI) for contributing as provider for instrumentation (funding 1S10OD019973-01). cDNA generation, library preparation and sequencing were performed by the University of Pittsburgh Health Science Sequencing Core at the University of Pittsburgh Medical Center Children’s Hospital of Pittsburgh. This work benefitted from ImageStreamX Mark II funded by NIH 1S10OD019942-01 (principal investigator L. Borghesi). This work was supported by NIH grant R37-AI118841.
Author information
Authors and Affiliations
Contributions
C.L., S.J. and M.J.S. conceived the project and designed experiments. S.J. designed and validated the Tlr9K51E and Tlr9P915H constructs. S.G. created the mice. C.L. and K.B.T. performed experiments, and C.L. analyzed data. S.S. assisted with ATAC-seq and RNA-seq data analysis. R.A.G., K.M.N. and R.A.E. assisted with data interpretation and experimental design. R.C.L. performed the in vitro RNA-seq experiments. R.A.E. and S.J. designed and performed the multiparametric flow cytometry panels. D.J.C. performed the ATAC-seq analysis. S.B. conducted pathologic analysis of the kidney tissue. R.F. and K.M. provided TLR9 antibody (Nar9 clone). C.L. and M.J.S. wrote the manuscript. M.J.S. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Marcus Clark and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. L.A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 TLR9 has a scaffold protective role in lupus.
a-b, Proportion of activated CD21int (a) or CD21hi (b) B cells with nuclear NF-κB (black gate, Fig. 1c) before (black bar) and after (red bar) stimulation. Five week old MRL/lpr mice, n = 3 per group, (* p < 0.05; *** p < 0.001, two-tailed paired-t test between the unstimulated and stimulated conditions for each Tlr9 genotype; (a) Tlr9 WT unstimulated versus stimulated p = 0.0007, Tlr9K51E/K51E unstimulated versus stimulated p = 0.0452, (b) Tlr9 WT unstimulated versus stimulated p = 0.0013, Tlr9K51E/K51E unstimulated versus stimulated p = 0.0480; # p < 0.05, ### p < 0.001, one-way ANOVA between the stimulated conditions). c-g, Evaluation of the clinical parameters, separated by gender (Tlr9WT male, n = 54; Tlr9−/− male, n = 21; Tlr9K51E/K51E male, n = 15; Tlr9WT female, n = 46; Tlr9−/− female, n = 22; Tlr9K51E/K51E female, n = 16; (c female) Tlr9WT versus Tlr9−/− p < 0.0001, Tlr9−/− versus Tlr9K51E/K51E p = 0.0001; (d) Tlr9WT versus Tlr9−/− p = 0.0056, Tlr9−/− versus Tlr9K51E/K51E p < 0.0001,Tlr9WT versus Tlr9K51E/K51E p = 0.038; (e) Tlr9WT versus Tlr9−/− p < 0.0001, Tlr9−/− versus Tlr9K51E/K51E p < 0.0001; (f) Tlr9WT versus Tlr9−/− p = 0.0173, Tlr9−/− versus Tlr9K51E/K51E p = 0.0003; (g) Tlr9WT versus Tlr9−/− p = 0.0632, Tlr9−/− versus Tlr9K51E/K51E p = 0.0115). h, Representative gating strategy of B cell populations (upper panel) and T cell (lower panel) subsets i, Percent TCR−CD19 + B cells of live splenocytes in male and female mice of the indicated genotypes (Tlr9WT, n = 86; Tlr9−/−, n = 36; Tlr9K51E/K51E, n = 29; (i) Tlr9WT versus Tlr9−/− p = 0.0420, Tlr9−/− versus Tlr9K51E/K51E p = 0.0078). j-k, Percent CD21hi CD23lo MZ, and CD11b+ CD11c+ ABC of live B cells in mice of the indicated genotypes. ((j) Tlr9WT, n = 86; Tlr9−/−, n = 36; Tlr9K51E/K51E, n = 29; Tlr9WT versus Tlr9−/− p = 0.0081, Tlr9−/− versus Tlr9K51E/K51E p = 0.0196; (k) Tlr9WT, n = 86; Tlr9−/−, n = 36; Tlr9K51E/K51E, n = 24; (k) Tlr9WT versus Tlr9−/− p = 0.0871, Tlr9−/− versus Tlr9K51E/K51E p = 0.0215) l, Percent TCR−CD44hiCD138hi plasmablasts of live splenocytes in mice of the indicated genotypes (Tlr9WT, n = 86; Tlr9−/−, n = 36; Tlr9K51E/K51E, n = 29; Tlr9WT versus Tlr9−/− p = 0.0372, Tlr9−/− versus Tlr9K51E/K51E p = 0.0015). m, Percent TCR+CD4+ and TCR+CD8+ cells of live splenocytes in male and female mice of the indicated genotypes (Tlr9WT, n = 79; Tlr9−/−, n = 36; Tlr9K51E/K51E, n = 28; for CD4 + , Tlr9−/− versus Tlr9K51E/K51E p = 0.0002, Tlr9WT versus Tlr9K51E/K51E p = 0.0023; for CD8 + , Tlr9WT versus Tlr9−/− p = 0.001, Tlr9−/− versus Tlr9K51E/K51E p = 0.0304). n-o, Percent naïve (CD62Lhi CD44lo), activated (CD62hi CD44hi) and memory (CD62Llo CD44hi) of CD4 and CD8 splenocytes (Tlr9WT, n = 79; Tlr9−/−, n = 36; Tlr9K51E/K51E, n = 28; for naïve CD4 + , Tlr9WT versus Tlr9−/− p = 0.0017, Tlr9−/− versus Tlr9K51E/K51E p = 0.0071; for activated CD4 + , Tlr9WT versus Tlr9−/− p < 0.0001, Tlr9−/− versus Tlr9K51E/K51E p = 0.0089; for naïve CD8 + , Tlr9WT versus Tlr9−/− p = 0.0012; for memory CD8 + , Tlr9WT versus Tlr9−/− p = 0.0095). For all panels, data points indicate individual mice and bars the mean ± SEM. For statistics in (c-o) * p < 0.05; ** p < 0.01; *** p < 0.001; **** p < 0.0001, one-way ANOVA with Tukey’s multiple comparisons test).
Extended Data Fig. 2 TLR9-MyD88 signaling promotes disease.
a-e, Evaluation of the clinical parameters separated by gender (Tlr9−/− male, n = 21; Tlr9P915H/P915H male, n = 37; Tlr9+/− male, n = 24; Tlr9−/− female, n = 22; Tlr9P915H/P915H female, n = 25; Tlr9+/− female, n = 28; (a, female) Tlr9−/− versus Tlr9P915H/P915H p = 0.0022, Tlr9−/− versus Tlr9+/− p = 0.0431; (b) Tlr9−/− versus Tlr9P915H/P915H p < 0.0001, Tlr9P915H/P915H versus Tlr9+/− p = 0.0005; (c) Tlr9−/− versus Tlr9P915H/P915H p = 0.0007, Tlr9P915H/P915H versus Tlr9+/− p = 0.0445; (d) Tlr9−/− versus Tlr9P915H/P915H p < 0.0001, Tlr9P915H/P915H versus Tlr9+/− p = 0.0005; (e) Tlr9P915H/P915H versus Tlr9+/− p = 0.0521) f, Percent TCR−CD19+ B cells of live splenocytes in male and female mice of the indicated genotypes (Tlr9−/−, n = 36; Tlr9P915H/P915H, n = 55; Tlr9+/−, n = 44; Tlr9−/− versus Tlr9+/− p = 0.0260). g-h, Percent CD21hi CD23lo MZ (Tlr9−/−, n = 36; Tlr9P915H/P915H, n = 55; Tlr9+/−, n = 44), and CD11b+ CD11c+ ABC of live B cells in mice of the indicated genotypes (Tlr9−/−, n = 36; Tlr9P915H/P915H, n = 53; Tlr9+/−, n = 41; (g) Tlr9−/− versus Tlr9P915H/P915H p = 0.0109, Tlr9P915H/P915H versus Tlr9+/− p = 0.0032; (h) Tlr9−/− versus Tlr9P915H/P915H p = 0.0228). i, Percent TCR−CD44hiCD138hi plasmablasts of live splenocytes in mice of the indicated genotypes (Tlr9−/−, n = 36; Tlr9P915H/P915H, n = 55; Tlr9+/−, n = 44; Tlr9−/− versus Tlr9P915H/P915H p = 0.0264). j, Percent TCR+CD4+ and TCR+CD8+ cells of live splenocytes in male and female mice of the indicated genotypes (Tlr9−/−, n = 36; Tlr9P915H/P915H, n = 52; Tlr9+/−, n = 44; CD4+, Tlr9−/− versus Tlr9P915H/P915H p = 0.0047, Tlr9P915H/P915H versus Tlr9+/− p = 0.0131; CD8+, Tlr9−/− versus Tlr9P915H/P915H p = 0.0001, Tlr9P915H/P915H versus Tlr9+/− p = 0.0005). k-l, Percent naïve (CD62Lhi CD44lo), activated (CD62Lhi CD44hi) and memory (CD62Llo CD44hi) of CD4 and CD8 splenocytes (Tlr9−/−, n = 36; Tlr9P915H/P915H, n = 52; Tlr9+/−, n = 44, (k) naïve Tlr9−/− versus Tlr9P915H/P915H p < 0.0001, Tlr9P915H/P915H versus Tlr9+/− p = 0.0018; activated Tlr9−/− versus Tlr9P915H/P915H p = 0.0345; (l) activated Tlr9−/− versus Tlr9P915H/P915H p = 0.0324, Tlr9P915H/P915H versus Tlr9+/− p = 0.0034; memory Tlr9−/− versus Tlr9P915H/P915H p = 0.0013, Tlr9P915H/P915H versus Tlr9+/− p = 0.0006). (a-l, For all panels, data points indicate individual mice and bars the mean ± SEM. For statistics in (a-l) * p < 0.05; ** p < 0.01; *** p < 0.001; **** p < 0.0001, one-way ANOVA with Tukey’s multiple comparisons test).
Extended Data Fig. 3 Effects of TLR9 and gender on TLR7 expression and distribution.
a-d, Representative flow cytometry plots of live, singlet, TCRβ−CD19+ B cells of 13 week old BALB/c (a) or 5 week old MRL/lpr (d) mice. Numbers indicate the frequencies in the gate. b,c, Comparison of TLR7 and TLR9 MFI between Tlr9 WT male and Tlr9 WT female BALB/c mice is shown for each of the defined B cell subsets (male, n = 8; female, n = 8 from two experiments; (b) male versus female MZ B cells p = 0.0333; male FO versus male MZ B cells p < 0.0001; (c) male FO versus male MZ B cells p = 0.0009). e,f, Comparison of TLR7 and TLR9 MFI between Tlr9 WT male and Tlr9 WT female 5 week old MRL/lpr mice is shown for each of the defined B cell subsets (male, n = 9 mice; female, n = 9 mice from two experiments; (e) male versus female FO B cells p < 0.0001; male versus female MZ B cells p < 0.0001; male versus female ABC B cells p = 0.0282; male FO versus male MZ B cells p < 0.0001; male FO versus male ABC B cells p < 0.0001; male MZ versus male ABC B cells p = 0.0330 (f) male FO versus male MZ B cells p < 0.0001; male FO versus male ABC B cells p < 0.0001; male MZ versus male ABC B cells p < 0.0001). (For b,c,e,f, data points indicate individual mice and bars the mean ± SEM of two experiments pooled. * p < 0.05, **** p < 0.0001, two tailed unpaired t-test between male and female groups; b,c, # p < 0.05; ## p < 0.01; ### p < 0.001; #### p < 0.0001, two tailed paired-t-test between subsets in the male group; e,f # p < 0.05; ## p < 0.01; ### p < 0.001; #### p < 0.0001 RM one way ANOVA with Tukey’s multiple comparisons test between subsets in the male group.) g-i, Quantification of TLR7 MFI (ratio of TLR7 MFI over average TLR7 MFI of the corresponding male or female WT group), in FO, MZ and ABC B cells of 5 week old female and male MRL/lpr mice (Tlr9K51E/K51E male, n = 4; female, n = 4; Tlr9 WT male, n = 4, female, n = 8; Tlr9P915H/P915H male, n = 4, female, n = 7; Tlr9+/− male, n = 3, female, n = 4; Tlr9−/− male, n = 3, female, n = 7; (g, male) Tlr9K51E/K51E versus Tlr9P915H/P915H p = 0.0097; Tlr9K51E/K51E versus Tlr9−/− p = 0.0061, Tlr9+/− versus Tlr9−/− p = 0.0430; (g, female) Tlr9 WT versus Tlr9+/− p = 0.0232; Tlr9 WT versus Tlr9−/− p < 0.0001; Tlr9K51E/K51E versus Tlr9+/− p = 0.0386, Tlr9K51E/K51E versus Tlr9−/- p < 0.0001, Tlr9P915H/P915H versus Tlr9 -/- p = 0.0002; (h, male) Tlr9 WT versus Tlr9P915H/P915H p = 0.0018; Tlr9 WT versus Tlr9 -/- p = 0.0261; Tlr9K51E/K51E versus Tlr9P915H/P915H p < 0.0001; Tlr9K51E/K51E versus Tlr9−/− p = 0.0005; Tlr9P915H/P915H versus Tlr9+/− p = 0.0001; Tlr9−/− versus Tlr9+/− p = 0.0012; (h, female) Tlr9 WT versus Tlr9−/− p = 0.0033; Tlr9K51E/K51E versus Tlr9−/− p < 0.0021; Tlr9−/− versus Tlr9+/− p = 0.0074; (i, male) Tlr9 WT versus Tlr9K51E/K51E p = 0.0398, Tlr9K51E/K51E versus Tlr9P915H/P915H p = 0.0143, Tlr9K51E/K51E versus Tlr9−/− p = 0.0284; (i, female) Tlr9 WT versus Tlr9+/− p = 0.0011, Tlr9 WT versus Tlr9−/− P < 0.0001, Tlr9K51E/K51E versus Tlr9+/− p = 0.0004, Tlr9K51E/K51E versus Tlr9−/− p < 0.0001, Tlr9P915H/P915H versus Tlr9+/− p = 0.004, Tlr9P915H/P915H versus Tlr9−/− p = 0.0002). j-l, Quantification of TLR9 MFI (ratio of TLR9 MFI over mean TLR9 MFI of the corresponding male or female WT group), in FO, MZ and ABC B cells of 5 week old female and male MRL/lpr mice (Tlr9K51E/K51E male, n = 4; female, n = 4; Tlr9 WT male, n = 4, female, n = 8; Tlr9P915H/P915H male, n = 4, female, n = 7; Tlr9+/− male, n = 3, female, n = 4; Tlr9−/− male, n = 3, female, n = 7; (j, male) Tlr9 WT versus Tlr9P915H/P915H p = 0.0001, Tlr9 WT versus Tlr9+/− p < 0.0001, Tlr9 WT versus Tlr9−/− p < 0.0001, Tlr9K51E/K51E versus Tlr9P915H/P915H p = 0.0004, Tlr9K51E/K51E versus Tlr9+/− p < 0.0001, Tlr9K51E/K51E versus Tlr9−/− p < 0.0001, Tlr9P915H/P915H versus Tlr9−/− p = 0.0004, Tlr9+/− versus Tlr9−/− p = 0.0104; (j, female) Tlr9 WT versus Tlr9P915H/P915H p < 0.0001, Tlr9 WT versus Tlr9+/− p < 0.0001, Tlr9 WT versus Tlr9−/− p < 0.0001, Tlr9K51E/K51E versus Tlr9P915H/P915H p < 0.0001, Tlr9K51E/K51E versus Tlr9+/− p < 0.0001, Tlr9K51E/K51E versus Tlr9−/− p < 0.0001, Tlr9P915H/P915H versus Tlr9−/− p < 0.0001, Tlr9+/− versus Tlr9−/− p < 0.0001; (k, male) Tlr9 WT versus Tlr9P915H/P915H p < 0.0001, Tlr9 WT versus Tlr9+/− p < 0.0001, Tlr9 WT versus Tlr9−/− p < 0.0001, Tlr9K51E/K51E versus Tlr9P915H/P915H p < 0.0001, Tlr9K51E/K51E versus Tlr9+/− p < 0.0001, Tlr9K51E/K51E versus Tlr9−/− p < 0.0001, Tlr9P915H/P915H versus Tlr9−/− p = 0.0008, Tlr9+/− versus Tlr9−/− p = 0.0015; (k, female) Tlr9 WT versus Tlr9P915H/P915H p < 0.0001, Tlr9 WT versus Tlr9+/− p < 0.0001, Tlr9 WT versus Tlr9−/− p < 0.0001, Tlr9K51E/K51E versus Tlr9P915H/P915H p < 0.0001, Tlr9K51E/K51E versus Tlr9+/− p < 0.0001, Tlr9K51E/K51E versus Tlr9−/− p < 0.0001, Tlr9P915H/P915H versus Tlr9+/− p = 0.0467, Tlr9P915H/P915H versus Tlr9−/− p < 0.0001, Tlr9+/− versus Tlr9−/− p < 0.0001; (l, male) Tlr9 WT versus Tlr9P915H/P915H p < 0.0001, Tlr9 WT versus Tlr9+/− p < 0.0001, Tlr9 WT versus Tlr9−/− p < 0.0001, Tlr9K51E/K51E versus Tlr9P915H/P915H p < 0.0001, Tlr9K51E/K51E versus Tlr9+/− p < 0.0001, Tlr9K51E/K51E versus Tlr9−/− p < 0.0001, Tlr9P915H/P915H versus Tlr9−/− p < 0.0001, Tlr9+/− versus Tlr9−/− p < 0.0001; (l, female) Tlr9 WT versus Tlr9P915H/P915H p < 0.0001, Tlr9 WT versus Tlr9+/− p < 0.0001, Tlr9 WT versus Tlr9−/− p < 0.0001, Tlr9K51E/K51E versus Tlr9P915H/P915H p = 0.0018, Tlr9K51E/K51E versus Tlr9+/− p = 0.0108, Tlr9K51E/K51E versus Tlr9−/− p < 0.0001, Tlr9P915H/P915H versus Tlr9−/− p = 0.0008, Tlr9+/− versus Tlr9−/− p = 0.0024). (For g-l, data points indicate individual mice and bars the mean ± SEM of one or two experiments pooled; * p < 0.05; ** p < 0.01; *** p < 0.001; **** p < 0.0001, one-way ANOVA with Tukey’s multiple comparisons test). m, The percent of TLR7 positivity in EEA1+ or LAMP-1+ endosomes of TLR9WT B cells is unchanged after a 2 hour stimulation with TLR7 agonist (CL097). (TLR7+EEA1+ unstimulated B cells, n = 17 from one mouse; TLR7+EEA1+ stimulated B cells, n = 25 from one mouse; TLR7+LAMP1+ unstimulated B cells, n = 27 from two mice; TLR7+LAMP1+ stimulated B cells, n = 40 from two mice). Data points indicate individual fields (1-3 cells per field) analyzed, and bars the mean ± SEM of two mice.
Extended Data Fig. 4 TLR9–MyD88-independent functions are B cell-intrinsic.
a-b, The competitive advantage of Tlr9WT vs Tlr9−/− or Tlr9P915H/P915H in different subsets of immature (n = 11 mice from three cohorts), transitional and naïve B cells (n = 15 mice from three cohorts), at the late time point (19-20 weeks post chimerism). Values greater than one on the log2 scale indicate TLR9WT advantage and below one indicate Tlr9−/− or Tlr9P915H/P915H advantage ((a) T2, p = 0.066; FO, p < 0.0001; (b) Fr. A-C, p < 0.0434; Fr. D, p = 0.0063; Fr. E, p = 0.0092). c,d, The competitive advantage of Tlr9WT vs Tlr9−/− or Tlr9P915H/P915H in different subsets of mature and differentiated B cells at the late time point (FO, MZ, GC, ABC, PB, CD80 + PDL2 + n = 15 mice from three cohorts; PC, n = 11 mice from three cohorts). Values greater than one on the log2 scale indicate TLR9WT advantage and below one indicate Tlr9−/− or Tlr9P915H advantage ((c) FO, p < 0.0001; CD80+PDL2+, p = 0.0162; PB, p = 0.0003; PC, p = 0.0356; FO versus MZ, p < 0.0001 (d) ABC, p = 0.0025; CD80+PDL2+, p = 0.0052; PB, p = 0.0014; PC, p = 0.0288). e, The competitive advantage of Tlr9−/− vs Tlr9P915H/P915H in different subsets of mature and differentiated B cells at the late time point (FO, MZ, GC, ABC, PB, CD80 + PDL2 + n = 15 mice from three cohorts; PC, n = 11 mice from three cohorts). Values greater than one indicate Tlr9−/− advantage and below one indicate Tlr9P915H advantage (FO, p < 0.0001; GC, p = 0.0002; ABC, p = 0.0005; CD80+PDL2+, p = 0.0002; PB, p < 0.0001; PC, p = 0.0056). (a-e) Data were pooled from three cohorts (two females and one male). Symbols represent individual mice and bars the mean ± SEM. Each group was compared to the hypothetical mean of no competitive advantage (one), by two-tailed one-sample t test (# p < 0.05; ## p < 0.01; ### p < 0.001; #### p < 0.0001), shown above each individual bar. Additionally, for (c), statistical significance between FO and the other B cell subsets was calculated by a mixed-effect analysis with Dunnett’s multiple comparisons test (* p < 0.05). f,g, Quantification of TLR7 (f) and TLR9 (g) in Tlr9WT transitional (T1,T2) (TLR7 n = 5; TLR9, n = 5 mice), FO (TLR7, n = 18; TLR9, n = 12 mice), MZ (TLR7, n = 18; TLR9, n = 12 mice), ABC (TLR7, n = 18; TLR9, n = 12 mice) and memory-like CD80+PDL2+ (TLR7, n = 11; TLR9, n = 5 mice) B cells of the chimera mice, at the early time point. TLR7 expression is minimal in immature B cells and maximal in ABCs ((f) T1 versus FO, p = 0.0180; T1 versus MZ, p = 0.0122; T1 versus ABC, p = 0.0017; T1 versus CD80+PDL2+, p = 0.0059; T2 versus FO, p = 0.0127; T2 versus MZ, p = 0.0119; T2 versus ABC, p = 0.0015; T2 versus CD80+PDL2+, p = 0.0064; FO versus MZ, p < 0.0001; FO versus ABC, p < 0.0001; FO versus CD80+PDL2+, p < 0.0001; MZ versus ABC, p < 0.0001; ABC versus CD80+PDL2+, p < 0.0001; (g) T1 versus ABC, p = 0.0193; T2 versus ABC, p = 0.0240; FO versus ABC, p < 0.0001; MZ versus ABC, p < 0.0001; ABC versus CD80+PDL2+, p = 0.0144). ((f-g), Symbols represent individual mice and bars the mean ± SEM. * p < 0.05; ** p < 0.01; *** p < 0.001; **** p < 0.0001 Mixed-effects analysis between subsets, with Tukey’s multiple comparisons test). h,i, Quantification of TLR7 MFI (ratio of TLR7 MFI in Tlr9−/− or Tlr9P915H/P915H cells over Tlr9WT subset per mouse), in FO (early, n = 18; late, n = 14), MZ (early, n = 18; late, n = 14), ABC (early, n = 18; late, n = 14) and CD80+PDL2+ (early, n = 11; late, n = 14) B cells, at the early (h) and late (i) time point. Each group was compared to the hypothetical mean of one (indicating similar TLR7 expression in mutated Tlr9 and Tlr9WT mice), by two-tailed one-sample t test (# p < 0.05; ## p < 0.01; ### p < 0.001; #### p < 0.0001), shown above each individual bar; (Symbols represent individual mice and bars the mean ± SEM; (h) Tlr9−/− FO, p < 0.0001; Tlr9P915H/P915H FO, p = 0.0002; Tlr9−/− MZ, p < 0.0001; Tlr9P915H/P915H MZ, p < 0.0001; Tlr9−/− ABC, p < 0.0001; Tlr9P915H/P915H ABC, p = 0.0024; Tlr9P915H/P915H CD80+PDL2+, p = 0.0174; (i) Tlr9−/− FO, p = 0.0019; Tlr9P915H/P915H FO, p < 0.0001; Tlr9−/− MZ, p = 0.0008; Tlr9P915H/P915H MZ, p < 0.0001; Tlr9−/− ABC, p < 0.0001; Tlr9P915H/P915H ABC, p = 0.0065). j,k, Quantification of TLR9 MFI in T1 (n = 5), T2 (n = 5), FO (n = 12), MZ (n = 12), ABC (n = 12) and CD80+PDL2+ (n = 5) B cells of the 3-way BM chimera mice at the early time point. Symbols represent individual mice and bars the mean ± SEM; For statistics, * p < 0.05; ** p < 0.01; *** p < 0.001; **** p < 0.0001, RM one-way ANOVA with Tukey’s multiple comparisons test; (j) T1, Tlr9WT versus Tlr9−/−, p = 0.0004; Tlr9WT versus Tlr9P915H/P915H, p = 0.0022; Tlr9−/− versus Tlr9P915H/P915H, p = 0.0004; T2, Tlr9WT versus Tlr9−/−, p = 0.0006; Tlr9WT versus Tlr9P915H/P915H, p = 0.0003; Tlr9−/− versus Tlr9P915H/P915H, p = 0.0009; FO, Tlr9WT versus Tlr9−/−, p < 0.0001; Tlr9WT versus Tlr9P915H/P915H, p < 0.0001; Tlr9−/− versus Tlr9P915H/P915H, p < 0.0001; (k) MZ, Tlr9WT versus Tlr9−/−, p < 0.0001; Tlr9WT versus Tlr9P915H/P915H, p < 0.0001; Tlr9−/− versus Tlr9P915H/P915H, p < 0.0001; ABC, Tlr9WT versus Tlr9−/−, p < 0.0001; Tlr9WT versus Tlr9P915H/P915H, p < 0.0001; Tlr9−/− versus Tlr9P915H/P915H, p < 0.0001; CD80+PDL2+, Tlr9WT versus Tlr9−/−, p = 0.0001; Tlr9WT versus Tlr9P915H/P915H, p = 0.0026; Tlr9−/− versus Tlr9P915H/P915H, p = 0.0006; l-p, Frequencies of T1 (n = 5), T2 (n = 3), FO (n = 4), ABC (n = 5) B cells and PB (n = 5) of the 3-way BM chimera mice in S or G2/M phase, determined by Ki67 and DAPI staining, and frequencies of activated caspase 3+ cells for each B cell subset (n = 6 mice). ((l-p), data points indicate individual mice and bars the mean ± SEM. For statistics * p < 0.05; ** p < 0.01; *** p < 0.001; **** p < 0.0001, RM one-way ANOVA with Turkey’s multiple comparisons test. (l), G2/M T1, Tlr9WT versus Tlr9P915H/P915H, p = 0.0240; Caspase 3+ Tlr9−/− versus Tlr9P915H/P915H, p = 0.0020; Caspase 3+ Tlr9WT versus Tlr9P915H/P915H, p = 0.0076; (m), S T2, Tlr9−/− versus Tlr9P915H/P915H, p = 0.0485; G2/M T2, Tlr9WT versus Tlr9−/−, p = 0.0172; Caspase 3+ Tlr9−/− versus Tlr9P915H/P915H, p = 0.0333; Caspase 3+ Tlr9WT versus Tlr9P915H/P915H, p = 0.0361; (n) G2/M FO, Tlr9WT versus Tlr9−/−, p = 0.0109; Tlr9WT versus Tlr9P915H/P915H, p = 0.0113; (o) Caspase 3+ Tlr9−/− versus Tlr9P915H/P915H, p = 0.0473).
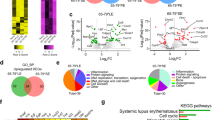
Extended Data Fig. 5 TLR9 controls ABC gene programs independently of MyD88.
a, PCA plot showing differences in accessible regions between the ABCs of the three genotypes Tlr9WT, Tlr9−/− and Tlr9P915H, sorted from the 3-way BM chimera mice. The different genotypes are represented as different shapes. b, PCA plot showing transcriptional differences between the ABCs of the three genotypes Tlr9WT, Tlr9−/− and Tlr9P915H, sorted from the 3-way BM chimera mice. c, Venn diagram representing the number of up-regulated genes (from Fig. 7d), shared or unique between Tlr9WT and Tlr9P915H ABCs in comparison to Tlr9−/− ABCs. d, Gene Set Enrichment Analysis (GSEA) was performed to look for plasma cell gene signature. GSEA shows significant enrichment of a plasma cell gene signature in Tlr9WT compared to Tlr9P915H ABCs (left panel), but no significant enrichment between Tlr9WT and Tlr9−/− ABCs (right panel). The p value was calculated using rankSumTestWithCorrelation function from the R limma package. e, XY plot showing the transcripts of the inhibitory pathway identified in Fig. 7d that were uniquely upregulated in Tlr9P915H ABCs vs. the transcripts upregulated by both Tlr9P915H and Tlr9WT ABCs, compared to Tlr9−/− ABCs. f, Quantification by qPCR (lower panels) of the transcripts of key inhibitory markers, found uniquely upregulated in Tlr9P915H ABCs compared to Tlr9WT and Tlr9−/− ABCs by RNA seq (upper panels), in sorted Tlr9−/−, Tlr9P915H and Tlr9WT ABCs from another cohort of 3-way BM chimera MRL/lpr mice (n = 4). g, Gene Set Enrichment Analysis (GSEA) was performed to look for TLR7-induced gene-set signature. GSEA shows no significant enrichment of TLR7-induced gene-set signature in Tlr9−/− ABCs compared to Tlr9P915H ABCs of the 3-way BM chimera MRL/lpr mice. The p value was calculated using rankSumTestWithCorrelation function from the R limma package. h, Venn diagram representing that only 20 genes were upregulated in Tlr9−/− ABCs compared to both Tlr9P915H and Tlr9WT ABCs.
Extended Data Fig. 6 Effects of TLR9 genotypes on FO and MZ B cell gene programs.
RNA-seq analysis of sorted FO B cells (TCRβ− CD19+ CD23hi CD21lo) and MZ B cells (TCRβ− CD19+ CD23lo CD21hi) of the different Tlr9 genotypes from the 3-way BM chimera MRL/lpr mice (described in Fig. 6a), all over 19 weeks post-chimerization. a, PCA plot shows transcriptional differences between the FO B cells of the three genotypes Tlr9WT, Tlr9−/− and Tlr9P915H, sorted from the 3-way BM chimera mice. b, Number of DEGs identified using Limma R package (Log2 fold-change >1 and FDR-corrected p value < 0.05) between the different Tlr9 genotypes from sorted FO B cells of the 3-way BM chimera MRL/lpr mice (n = 4 replicates). c, Venn diagram representing the number of up-regulated genes (from Fig. E6b), in Tlr9WT FO B cells compared to Tlr9P915H and Tlr9−/− FO B cells. d, XY plot showing the DEG from Fig. E6b: (i) uniquely upregulated in Tlr9WT FO B cells compared to Tlr9−/− FO B cells; (ii) upregulated in Tlr9WT FO B cells compared to Tlr9P915H FO B cells; or (iii) upregulated in comparison to both Tlr9P915H and Tlr9−/− FO B cells. e, PCA plot showing transcriptional differences between the MZ B cells of the three genotypes Tlr9WT, Tlr9−/− and Tlr9P915H, sorted from the 3-way BM chimera mice. f, Number of DEGs identified using Limma R package (Log2 fold-change > 1 and FDR-corrected p value < 0.05) between the different Tlr9 genotypes from sorted MZ B cells of the 3-way BM chimera MRL/lpr mice (n = 4 replicates). g, Venn diagram representing the number of up-regulated genes from Fig. E6f, in Tlr9WT MZ B cells compared to Tlr9P915H and Tlr9−/− MZ B cells. h, XY plot showing the DEG from Fig. E6f): (i) uniquely upregulated in Tlr9WT MZ B cells compared to Tlr9−/− MZ B cells, (ii) upregulated in Tlr9WT MZ B cells compared to Tlr9P915H MZ B cells, or (iii) upregulated in comparison to both Tlr9P915H and Tlr9−/− MZ B cells.
Extended Data Fig. 7 Tlr9P915H and Tlr9−/− induce distinct ABC and PB states.
a,b, Costimulatory molecules and PB-associated transcripts from the RNA-seq analysis that were more highly expressed in Tlr9−/− and Tlr9WT ABCs compared to Tlr9P915H ABCs (upper row, n = 4) were investigated at the protein level by flow cytometry (lower row) in a separate cohort of 3-way BM chimera MRL/lpr mice (n = 12) ((a) CD80, Tlr9−/− versus Tlr9P915H, p < 0.0001; Tlr9P915H versus Tlr9WT,p = 0.0023; CD86, Tlr9−/− versus Tlr9P915H, p < 0.0001; Tlr9P915H versus Tlr9WT,p = 0.0017; CD200, Tlr9−/− versus Tlr9P915H, p = 0.0040; Tlr9−/− versus Tlr9WT,p = 0.0073; Itgb7, Tlr9−/− versus Tlr9P915H, p = 0.0003; Tlr9P915H versus Tlr9WT,p = 0.0057; (b) CD44, Tlr9−/− versus Tlr9P915H, p = 0.0003; Tlr9P915H versus Tlr9WT,p = 0.0076; CXCR4, Tlr9−/− versus Tlr9P915H, p < 0.0001; Tlr9P915H versus Tlr9WT,p = 0.0008; CD138, Tlr9−/− versus Tlr9P915H, p < 0.0001; Tlr9P915H versus Tlr9WT,p = 0.0094). c,d, Flow cytometry analysis shows differential expression (MFI) of selected transcription factors (c) and the inhibitory molecule CD300a (d) among Tlr9 genotypes in ABCs of the 3-way BM chimera MRL/lpr mice ((c) IRF4, Tlr9−/− versus Tlr9P915H, p = 0.0002; BCL6, Tlr9−/− versus Tlr9P915H, p = 0.0015; Tlr9P915H versus Tlr9WT, p = 0.0011; Tlr9−/− versus Tlr9WT,p = 0.0025; TBET, Tlr9−/− versus Tlr9P915H, p = 0.0002; Tlr9−/− versus Tlr9WT,p = 0.0107; (d) CD300a, Tlr9−/− versus Tlr9P915H, p < 0.0001; Tlr9P915H versus Tlr9WT,p = 0.0188). (a-d) Splenocytes from twelve 3-way BM chimera mice (n = 6 females, and n = 6 males from 2 experiments) were stained for the indicated markers and the CD45.1/CD45.2 congenic markers. (Each line represents a 3-way BM chimera MRL/lpr mouse, * p < 0.05; ** p < 0.01; *** p < 0.001; **** p < 0.0001, RM one-way ANOVA with Turkey’s multiple comparisons test).
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Leibler, C., John, S., Elsner, R.A. et al. Genetic dissection of TLR9 reveals complex regulatory and cryptic proinflammatory roles in mouse lupus. Nat Immunol 23, 1457–1469 (2022). https://doi.org/10.1038/s41590-022-01310-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-022-01310-2
This article is cited by
-
Cyclical palmitoylation regulates TLR9 signalling and systemic autoimmunity in mice
Nature Communications (2024)
-
Phospholipase D4 as a signature of toll-like receptor 7 or 9 signaling is expressed on blastic T-bet + B cells in systemic lupus erythematosus
Arthritis Research & Therapy (2023)
-
HMGB1 and Toll-like receptors: potential therapeutic targets in autoimmune diseases
Molecular Medicine (2023)
-
Genetics of SLE: mechanistic insights from monogenic disease and disease-associated variants
Nature Reviews Nephrology (2023)