Abstract
Tissue-resident memory T cells (TRM cells) provide rapid and superior control of localized infections. While the transcription factor Runx3 is a critical regulator of CD8+ T cell tissue residency, its expression is repressed in CD4+ T cells. Here, we show that, as a direct consequence of this Runx3-deficiency, CD4+ TRM cells lacked the transforming growth factor (TGF)-β-responsive transcriptional network that underpins the tissue residency of epithelial CD8+ TRM cells. While CD4+ TRM cell formation required Runx1, this, along with the modest expression of Runx3 in CD4+ TRM cells, was insufficient to engage the TGF-β-driven residency program. Ectopic expression of Runx3 in CD4+ T cells incited this TGF-β-transcriptional network to promote prolonged survival, decreased tissue egress, a microanatomical redistribution towards epithelial layers and enhanced effector functionality. Thus, our results reveal distinct programming of tissue residency in CD8+ and CD4+ TRM cell subsets that is attributable to divergent Runx3 activity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All original data is available from the corresponding author upon reasonable request. RNA-seq and ATAC-seq data is available in the Gene Expression Omnibus database under accession codes GSE182511 and GSE198611, respectively. Source data are provided with this paper.
Code availability
The code generated and used for the analysis of sequencing data are available from the corresponding author on reasonable request.
References
Jameson, S. C. & Masopust, D. Understanding subset diversity in T cell memory. Immunity 48, 214–226 (2018).
Masopust, D. & Soerens, A. G. Tissue-resident T cells and other resident leukocytes. Annu. Rev. Immunol. 37, 521–546 (2019).
Mackay, L. K. et al. The developmental pathway for CD103+CD8+ tissue-resident memory T cells of skin. Nat. Immunol. 14, 1294–1301 (2013).
Kok, L. et al. A committed tissue-resident memory T cell precursor within the circulating CD8+ effector T cell pool. J. Exp. Med. 217, e20191711 (2020).
Kurd, et al.Early precursors and molecular determinants of tissue resident memory CD8+ T lymphocytes revealed by single-cell RNA sequencing. Sci. Immunol. 5, eaaz6894 (2020).
Kok, L., Masopust, D. & Schumacher, T. N. The precursors of CD8+ tissue resident memory T cells: from lymphoid organs to infected tissues. Nat. Rev. Immunol. 22, 283–293 (2022).
Hirai, T. et al. Keratinocyte-mediated activation of the cytokine TGF-β maintains skin recirculating memory CD8+ T cells. Immunity 50, 1249–1261.e1245 (2019).
Zhang, N. & Bevan, J. M. Transforming growth factor-β signaling controls the formation and maintenance of gut-resident memory T cells by regulating migration and retention. Immunity 39, 687–696 (2013).
Christo, S. N. et al. Discrete tissue microenvironments instruct diversity in resident memory T cell function and plasticity. Nat. Immunol. 22, 1140–1151 (2021).
Skon, C. N. et al. Transcriptional downregulation of S1pr1 is required for the establishment of resident memory CD8+ T cells. Nat. Immunol. 14, 1285–1293 (2013).
Mackay, L. K. et al. Hobit and Blimp1 instruct a universal transcriptional program of tissue residency in lymphocytes. Science 352, 459–463 (2016).
Milner, J. J. et al. Runx3 programs CD8+ T cell residency in non-lymphoid tissues and tumours. Nature 552, 253–257 (2017).
Beura, L. K. et al. CD4+ resident memory T cells dominate immunosurveillance and orchestrate local recall responses. J. Exp. Med. 216, 1214–1229 (2019).
Nguyen, Q. P., Deng, T. Z., Witherden, D. A. & Goldrath, A. W. Origins of CD4+ circulating and tissue‐resident memory T‐cells. Immunology 157, 3–12 (2019).
Schreiner, D. & King, C. G. CD4+ memory T cells at home in the tissue: mechanisms for health and disease. Front Immunol. 9, 2394 (2018).
Hondowicz, B. D., Kim, K. S., Ruterbusch, M. J., Keitany, G. J. & Pepper, M. IL-2 is required for the generation of viral-specific CD4+ Th1 tissue-resident memory cells and B cells are essential for maintenance in the lung. Eur. J. Immunol. 48, 80–86 (2018).
Turner, D. L. & Farber, D. L. Mucosal resident memory CD4 T cells in protection and immunopathology. Front Immunol. 5, 331 (2014).
Kumar, B. V. et al. Human tissue-resident memory T cells are defined by core transcriptional and functional signatures in lymphoid and mucosal sites. Cell Rep. 20, 2921–2934 (2017).
Collins, N. et al. Skin CD4+ memory T cells exhibit combined cluster-mediated retention and equilibration with the circulation. Nat. Commun. 7, 11514 (2016).
Iijima, N. & Iwasaki, A. A local macrophage chemokine network sustains protective tissue-resident memory CD4 T cells. Science 346, 93–98 (2014).
Gebhardt, T. et al. Different patterns of peripheral migration by memory CD4+ and CD8+ T cells. Nature 477, 216–219 (2011).
Ariotti, S. et al. Tissue-resident memory CD8+ T cells continuously patrol skin epithelia to quickly recognize local antigen. Proc. Natl Acad. Sci. USA 109, 19739–19744 (2012).
Steinert, E. M. et al. Quantifying memory CD8 T cells reveals regionalization of immunosurveillance. Cell 161, 737–749 (2015).
Shin, B. et al. Runx1 and Runx3 drive progenitor to T-lineage transcriptome conversion in mouse T cell commitment via dynamic genomic site switching. Proc. Natl Acad. Sci. USA 118, e2019655118 (2021).
Setoguchi, R. et al. Repression of the transcription factor Th-POK by Runx complexes in cytotoxic T cell development. Science 319, 822–825 (2008).
Luckey, M. A. et al. The transcription factor ThPOK suppresses Runx3 and imposes CD4+ lineage fate by inducing the SOCS suppressors of cytokine signaling. Nat. Immunol. 15, 638–645 (2014).
Ciucci, T. et al. The emergence and functional fitness of memory CD4+ T cells require the transcription factor Thpok. Immunity 50, 91–105.e104 (2019).
Djuretic, I. M. et al. Transcription factors T-bet and Runx3 cooperate to activate Ifng and silence Il4 in T helper type 1 cells. Nat. Immunol. 8, 145–153 (2007).
Komine, O. et al. The Runx1 transcription factor inhibits the differentiation of naive CD4+ T cells into the Th2 lineage by repressing GATA3 expression. J. Exp. Med. 198, 51–61 (2003).
Reis, B. S., Rogoz, A., Costa-Pinto, F. A., Taniuchi, I. & Mucida, D. Mutual expression of the transcription factors Runx3 and ThPOK regulates intestinal CD4+ T cell immunity. Nat. Immunol. 14, 271–280 (2013).
Mackay, L. et al. T-box transcription factors combine with the cytokines TGF-β and IL-15 to control tissue-resident memory T cell fate. Immunity 43, 1101–1111 (2015).
Wang, D. et al. The transcription factor Runx3 establishes chromatin accessibility of cis-regulatory landscapes that drive memory cytotoxic T lymphocyte formation. Immunity 48, 659–674.e656 (2018).
Schenkel, J. M. et al. T cell memory. Resident memory CD8 T cells trigger protective innate and adaptive immune responses. Science 346, 98–101 (2014).
Ariotti, S. et al. T cell memory. Skin-resident memory CD8+ T cells trigger a state of tissue-wide pathogen alert. Science 346, 101–105 (2014).
Taniuchi, I. et al. Differential requirements for Runx proteins in CD4 repression and epigenetic silencing during T lymphocyte development. Cell 111, 621–633 (2002).
Moreau, J. M., Velegraki, M., Bolyard, C., Rosenblum, M. D. & Li, Z. Transforming growth factor-β1 in regulatory T cell biology. Sci. Immunol. 7, eabi4613 (2022).
Korn, T., Bettelli, E., Oukka, M. & Kuchroo, V. K. IL-17 and Th17 cells. Annu. Rev. Immunol. 27, 485–517 (2009).
Mucida, D. et al. Transcriptional reprogramming of mature CD4+ helper T cells generates distinct MHC class II–restricted cytotoxic T lymphocytes. Nat. Immunol. 14, 281–289 (2013).
Keller, H. R. et al. The molecular basis and cellular effects of distinct CD103 expression on CD4 and CD8 T cells. Cell. Mol. Life Sci. 78, 5789–5805 (2021).
Gebhardt, T. et al. Memory T cells in nonlymphoid tissue that provide enhanced local immunity during infection with herpes simplex virus. Nat. Immunol. 10, 524–530 (2009).
Fonseca, R. et al. Developmental plasticity allows outside-in immune responses by resident memory T cells. Nat. Immunol. 21, 412–421 (2020).
Stolley, J. M. et al. Retrograde migration supplies resident memory T cells to lung-draining LN after influenza infection. J. Exp. Med. 217, e20192197 (2020).
Klicznik, M. M. et al. Human CD4+CD103+ cutaneous resident memory T cells are found in the circulation of healthy individuals. Sci. Immunol. 4, eaav8995 (2019).
Masopust, D. et al. Dynamic T cell migration program provides resident memory within intestinal epithelium. J. Exp. Med. 207, 553–564 (2010).
Oh, D. Y. & Fong, L. Cytotoxic CD4+ T cells in cancer: expanding the immune effector toolbox. Immunity 54, 2701–2711 (2021).
Cheroutre, H. & Husain, M. M. CD4 CTL: living up to the challenge. Semin. Immunol. 25, 273–281 (2013).
Delacher, M. et al. Single-cell chromatin accessibility landscape identifies tissue repair program in human regulatory T cells. Immunity 54, 702–720.e17 (2021).
Durand, A. et al. Profiling the lymphoid-resident T cell pool reveals modulation by age and microbiota. Nat. Commun. 9, 68 (2018).
Zaid, A. et al. Persistence of skin-resident memory T cells within an epidermal niche. Proc. Natl Acad. Sci. USA 111, 5307–5312 (2014).
Acknowledgements
We thank the Flow Cytometry Unit and Bioresources Facility at Peter Doherty Institute (University of Melbourne) for technical assistance. This work was supported by a Howard Hughes Medical Institute and Bill and Melinda Gates International Research Scholarship OPP1175796, National Health and Medical Research Council (NHMRC) AP1113293 to F.RC. and L.K.M. S.L.P. was supported by a NHMRC EL1 Investigator Grant GNT1175626. N.G.Z. was supported by FAPESP (BEPE 2019/12431-2). A.T.S. was supported by the National Institutes of Health (U01CA260852 and UM1HG012076), the Parker Institute for Cancer Immunotherapy and a Pew-Stewart Scholars for Cancer Research Award. L.K.M is a Senior Medical Research Fellow supported by the Sylvia and Charles Viertel Charitable Foundation.
Author information
Authors and Affiliations
Contributions
R.F., T.N.B., L.C.G., S.D., A.O., S.L.P., M.E., S.K.S., S.N.C., N.M.Z., N.G.Z., F.A.B., C.A.L., K.M. and A.Z. performed experiments and analyzed data; S.N.M., T.P.S., A.T.S. and L.K.M. provided supervision; R.F., T.N.B., S.L.P. and L.K.M. contributed to experimental design; R.F., T.N.B., F.R.C. and L.K.M. prepared the manuscript; A.T.S., F.R.C. and L.K.M. provided funding; F.R.C. and L.K.M. led the research program.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests. A.T.S. is a founder of Immunai and Cartography Biosciences and receives unrelated research funding from Merck Research Laboratories, Allogene Therapeutics, and Arsenal Biosciences, L.K.M. receives unrelated research funding from Pfizer.
Peer review
Peer review information
Nature Immunology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Distribution and phenotype of memory T cells in the epithelia.
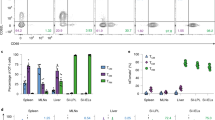
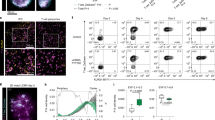
(a-d) Microscopy showing CD4+ SMARTA and CD8+ P14 cells in the small intestine (SI) 7 and 30 d.p.i. with LCMV (IEL cells highlighted by teal (CD4+) and yellow (CD8+) circles) (a), kinetics of CD4+ SMARTA and CD8+ P14 cells (**P=0.0079, 0.0043, 0.0079 respectively, two-tailed Mann-Whitney’s test) (b), CD69 and CD103 expression in SI-IEL at 7, 14 or 30 d.p.i. (***P=0.007, <0.0001 respectively, multiple paired t-tests) (c) and representative flow plots 30 d.p.i. (d). (e-i) CD69 and CD103 expression in CD4+ gDT-II and CD8+ gBT-I cells from skin at 7, 14 or 30d post-HSV infection (***P=0.0028, <0.0001 respectively, multiple paired t-tests) (e) and representative flow plots in skin 30 d.p.i. (f), representative histograms of Runx3 expression in spleen or skin 30 d.p.i. (g), representative histograms of ThPOK and Runx3 expression (h) and Runx3 expression (gMFI) in spleen 7 d.p.i. normalized to naïve (i). (j) Correlation plots of Runx1 or Runx3 and CD103 gMFI on endogenous Foxp3−CD69+CD4+ and CD69+CD8+ isolated from skin 14 d.p.i. with HSV (linear regression line with 95% confidence interval, P showing slope is non-zero). (k, l) Runx1 and Runx3 expression in CD8+ gBT-I cells (k) and CD4+ gDT-II cells (l) in vitro prior to transfer into recipient mice following CRISPR-Cas9 mediated editing of CD19 (Ctrl), Runx1 (sgRunx1), or Runx3 (sgRunx3). (m) Log2-fold change of sgCtrl and sgRunx1 CD4+ SMARTA cells (as in k, **P=0.002, paired t-test) in the spleen and SI-IEL >20d post-LCMV infection. Data are representative of 2 independent experiments with (a) n=3, (b-i) n=5, (j) 1 experiment, n=5, (k) 2 independent biological replicates, (m) pooled from 2 independent experiments, with n=5 mice. Symbols represent (c, e, j, m) mice, (b, i) mean; error bars indicate SEM. Box plots represent the median, interquartile range, and minimum/maximum whiskers.
Extended Data Fig. 2 Ectopic Runx3 enhances CD8+ TRM cell formation in the epithelia.
(a, b) Schematic and frequency of CD8+ P14 cells transduced with control (CD8-Ctrl) or Runx3 (CD8-Runx3) retroviruses in the spleen, skin and SI-IEL 14d post-LCMV infection and DNFB skin treatment (a) and enumeration of total or CD69+CD103+ cells in the spleen, skin (**P=0.0022, two-tailed paired t-test) and SI-IEL (***P=0.0002 and 0.0007 respectively) (b). Data pooled from (a, b) 2 independent experiments, n=4 mice. Symbols represent individual mice; bars represent mean; error bars indicate SEM.
Extended Data Fig. 3 Runx3 increases CD4+ TRM cell formation in the epithelia.
(a-c) Schematic and frequency of CD4+ gDT-II cells transduced with control (CD4-Ctrl) or Runx3 (CD4-Runx3) retroviruses in the spleen and skin (a), epidermis/dermis proportion (**P=0.0048, two-tailed paired t-test) (b) and CD69 and CD103 expression in the epidermis (**P=0.0021, two-tailed paired t-test) and dermis (***P=0.0001, two-tailed paired t-test) (c) 14d post-HSV infection. (d-f) Schematic (d), frequency of CD4+ SMARTA cells transduced with control (CD4-Ctrl) or Runx3 (CD4-Runx3) retroviruses (e) and number of total and CD69+CD103+ cells in spleen, skin (*P=0.043, two-tailed paired t-test) and SI-IEL (*P=0.0432 and 0.0332) (f) 14d post-LCMV infection in mice treated on the skin with DNFB. (g, h) Representative histograms of Runx3 (g) and expression (MFI) by endogenous CD4+, CD8+ and CD4+CD8αα+ T cells (h) in the spleen and SI-IEL >90d post-LCMV infection. (i) Runx3 expression by endogenous CD8+ T cells (CD8-Endog) and transduced Ctrl- (CD4-Ctrl) or Runx3-transduced (CD4-Runx3) CD4+ gDT-II (spleen and skin) or CD4+ SMARTA (SI-IEL) cells as in (a-f) 14 d.p.i. with HSV or LCMV respectively. Data pooled from (a-c) 2 independent experiments with n=10 (d-f) 2 independent experiments, n=5 or representative of (g, h) 2 independent experiments, n=5 and (i) 2 independent experiments, n=3 mice. Symbols represent individual mice; bars indicate mean; error bars indicate SEM.
Extended Data Fig. 4 Runx3 induces a CD8+ TRM cell-like signature in CD4+ T cells.
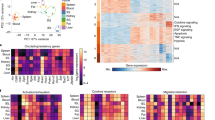
(a) Heatmap showing expression of the top TRM cell DEGs (derived from the comparison between skin CD8+ TRM cells vs spleen CD8+ TCIRC cells from GSE70813) in CD4-Runx3 and CD8-Ctrl samples relative to CD4-Ctrl cells, by CD8+ gBT-I cells transduced with control (CD8-Ctrl) and CD4+ gDT-II cells transduced with control (CD4-Ctrl) or Runx3-expressing (CD4-Runx3) retroviruses isolated from the skin 14 d.p.i. (columns represents independent samples, color scale is based on Z-score). (b) GSEA in CD8-Ctrl vs CD4-Ctrl (top) or CD4-Runx3 vs CD8-Ctrl cells (bottom) for the top 100 DEGs in skin CD8+ TRM vs splenic CD8+ TCIRC cells. (c) Scatter plot showing DEGs between skin CD8+ TRM vs splenic CD8+ TCIRC cells and CD8-Ctrl vs CD4-Ctrl or CD8-Ctrl vs CD4-Runx3 (dots represent up- (red) or down-regulated (blue) genes in skin CD8+ TRM vs splenic CD8+ TCIRC cells, quadrants represent up- (red) or down-regulated (blue) genes in CD8-Ctrl vs CD4-Ctrl or CD8-Ctrl vs CD4-Runx3 comparisons). (d) Scatter plots of standardized log fold-change of CD8-Ctrl vs CD4-Ctrl (x-axis) and CD4-Runx3 vs CD4-Ctrl (y-axis; orange dots highlight top skin TRM cell genes (left panel), blue and red dots highlight top 200 down- (middle panel) and up-regulated (right panel) genes in skin TRM cells) (d). Data pooled from 2 independent experiments, n=2 biological replicates pooled from 10 mice. Green line represents least-squares regression line fitted to orange points.
Extended Data Fig. 5 Runx3 promotes epigenetic remodeling in CD4+ T cells.
(a) Scatter plots showing peak accessibility changes in CD8-Ctrl cells treated with TGF-β vs Ctrl and CD4-Ctrl (top) or CD4-Runx3 (bottom) treated with TGF-β vs Ctrl. (b) Scatter plots showing peak accessibility changes in CD8-Runx3 vs CD8-Ctrl cells and CD4-Runx3 vs CD4-Ctrl treated with TGF-β (bottom) or untreated (top; orange dots highlight TGF-β regulated genes). (c) Itga1, Cd244, S1pr1 and Ly6c genome tracks (height normalized) in CD4+ gDT-II and CD8+ gBT-I cells transduced with control (CD4-Ctrl and CD8-Ctrl) or Runx3-expressing (CD4-Runx3 and CD8-Runx3) retroviruses cultured ± TGF-β for 48 hours. Data representative of 2 independent experiments, with n=2 technical replicates.
Extended Data Fig. 6 Runx3 induces expression of TGF-β-regulated genes in CD4+ T cells.
(a) Heatmap showing the top TGF-β regulated genes (derived from the comparison between skin wild-type (WT) CD8+ T cells vs Tgfbr2−/− CD8+ T cells 14d post-HSV infection from GSE178769) in CD4-Runx3 and CD8-Ctrl samples relative to CD4-Ctrl expression, by CD8+ gBT-I cells transduced with control (CD8-Ctrl) and CD4+ gDT-II cells transduced with either control (CD4-Ctrl) or Runx3-expressing (CD4-Runx3) retroviruses isolated from the skin 14d post-HSV infection (columns represents independent samples, color scale is based on Z-score). (b) GSEA in CD8-Ctrl vs CD4-Ctrl (top) or CD4-Runx3 vs CD8-Ctrl cells (bottom) for the top 100 DEGs in skin WT vs Tgfbr2−/− cells. (c) Scatter plot showing DEGs between WT vs Tgfbr2−/− cells and CD8-Ctrl vs CD4-Ctrl (left) or CD8-Ctrl vs CD4-Runx3 (right, dots represent up- (red) and down-regulated (blue) genes in WT vs Tgfbr2−/− cells comparison, quadrants represents up- (red) and down-regulated (blue) genes in CD8-Ctrl vs CD4-Ctrl or CD8-Ctrl vs CD4-Runx3 comparisons). (d) Scatter plots of standardized log-fold changes of CD8-Ctrl vs CD4-Ctrl (x-axis) and CD4-Runx3 vs CD4-Ctrl (y-axis) for all genes (orange dots highlight top TGF-β regulated genes (left panel), blue and red dots highlight top 200 down-regulated (middle panel) and up-regulated (right panel) genes in WT cells). Data pooled from 2 independent experiments, n=2 biological replicates pooled from 10 mice. Green line represents least-squares regression line fitted to orange dots.
Extended Data Fig. 7 Runx3 promotes enhanced local immune protection by skin CD4+ T cells.
(a) Intracellular cytokine production of IFNγ and TNFα by CD4+ gDT-II cells transduced with either control (CD4-Ctrl) or Runx3-expressing (CD4-Runx3) retroviruses 4 hours after PMA+ionomycin stimulation. (b, c) Viral titer (*P=0.0111, two-tailed unpaired t-test) 6d post-HSV infection (b), and enumeration of transduced cells, inflammatory monocytes, NK cells, CD4+ T cells, CD8+ T cells, neutrophils, macrophages, B cells and dendritic cells 3 d.p.i (*P=0.0455, 0.0301 and 0.021 respectively, two-tailed unpaired t-test) (c) in the skin of C57BL/6 mice transferred i.d. with CD4+ gDT-II cells transduced with control (CD4-Ctrl) or Runx3-expressing (CD4-Runx3) retroviruses 14d prior to HSV infection. Data pooled from 2 independent experiments with (a) n=2 biological replicates, (b, c) n=5 mice. Symbols represent (a) biological replicates or (b, c) mice. Bars represent mean; error bars indicate SEM. Box plots show the median, interquartile range and minimum/maximum whiskers.
Supplementary information
Supplementary Information
Supplementary methods, references and gating strategy.
Supplementary Video 1
CD4+ T cells from uGFP-reporter mice were transduced with an empty-Ametrine-encoding retrovirus (CD4-Ctrl) and CD4+ T cells from ubiTomato-reporter mice were transduced with a Runx3-GFP-encoding retrovirus (CD4-Runx3). Transduced cells were cotransferred i.d. into recipient mice and intravital two-photon microscopy performed at day 14 post-i.d. transfer. SHG..
Supplementary Table 1
List of antibodies.
Source data
Source Data Fig. 1
Source Data Fig. 1.
Source Data Fig. 2
Source Data Fig. 2.
Source Data Fig. 5
Source Data Fig. 5.
Source Data Fig. 6
Source Data Fig. 6.
Source Data Fig. 7
Source Data Fig. 7.
Source Data Extended Data Fig. 1
Source Data Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Source Data Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Source Data Extended Data Fig. 3.
Source Data Extended Data Fig. 7
Source Data Extended Data Fig. 7.
Rights and permissions
About this article
Cite this article
Fonseca, R., Burn, T.N., Gandolfo, L.C. et al. Runx3 drives a CD8+ T cell tissue residency program that is absent in CD4+ T cells. Nat Immunol 23, 1236–1245 (2022). https://doi.org/10.1038/s41590-022-01273-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-022-01273-4
This article is cited by
-
RUNX3 pathway signature predicts clinical benefits of immune checkpoint inhibition plus tyrosine kinase inhibition in advanced renal cell carcinoma
BMC Urology (2024)
-
Tissue-resident memory T cells: decoding intra-organ diversity with a gut perspective
Inflammation and Regeneration (2024)
-
Tissue-Resident Memory T Cells in Allergy
Clinical Reviews in Allergy & Immunology (2024)
-
The expanding Pandora’s toolbox of CD8+T cell: from transcriptional control to metabolic firing
Journal of Translational Medicine (2023)
-
CD4+ T cell memory
Nature Immunology (2023)