Abstract
Host genetic and environmental factors including age, biological sex, diet, geographical location, microbiome composition and metabolites converge to influence innate and adaptive immune responses to vaccines. Failure to understand and account for these factors when investigating severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) vaccine efficacy may impair the development of the next generation of vaccines. Most studies aimed at identifying mechanisms of vaccine-mediated immune protection have focused on adaptive immune responses. It is well established, however, that mobilization of the innate immune response is essential to the development of effective cellular and humoral immunity. A comprehensive understanding of the innate immune response and environmental factors that contribute to the development of broad and durable cellular and humoral immune responses to SARS-CoV-2 and other vaccines requires a holistic and unbiased approach. Along with optimization of the immunogen and vectors, the development of adjuvants based on our evolving understanding of how the innate immune system shapes vaccine responses will be essential. Defining the innate immune mechanisms underlying the establishment of long-lived plasma cells and memory T cells could lead to a universal vaccine for coronaviruses, a key biomedical priority.
Similar content being viewed by others
Main
Over the past two decades, the world has experienced a substantial number of disease outbreaks from viral infections including yellow fever, dengue fever, Ebola, Zika, severe acute respiratory syndrome (SARS), Middle East respiratory syndrome and the ongoing novel severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic1. The SARS-CoV-2 pandemic has illuminated how disruptive sustained infectious disease outbreaks can be on society. Given the unpredictability of when the next pandemic will emerge, strengthening our ability to respond rapidly to any infectious disease threat is a global priority.
In what may be one of the most impressive accomplishments of modern medicine, multiple effective vaccines were developed against SARS-CoV-2 in less than 12 months after the initial outbreak. As of February 2022 ten coronavirus disease 2019 (COVID-19) vaccines are currently approved for full or emergency use authorization by the World Health Organization (https://covid19.trackvaccines.org/agency/who/). These approved vaccines span four distinct vaccine platforms, providing an additional layer of diversity in potential vaccine responses. The Moderna and Pfizer-BioNTech vaccines both use modified mRNAs to produce antigen upon vaccination2,3. Both vaccines use lipid nanoparticle-encapsulated formulations to deliver mRNA that encodes for a prefusion stabilized, full-length SARS-CoV-2 spike (S) protein. The Johnson & Johnson (Ad26 vector) and AstraZeneca AZD1222 (ChAdOx1 vector) vaccines use distinct replication-incompetent adenoviral vectors to deliver antigen (the prefusion stabilized, full-length S protein)4,5. The Sinopharm and Sinovac-CoronaVac platforms utilize β-propiolactone-inactivated virus produced in Vero E6 cells and are formulated with the adjuvant alum to promote immunogenicity6,7. Protein subunit-based platforms are in advanced stages of development, with one approved for emergency use authorization in Indonesia (Novavax8; clinical trial no. NCT04742738). Approved vaccines have efficacy rates of ~50–95%, with Moderna and Pfizer exhibiting the highest rates of short-term efficacy.
SARS-CoV-2 vaccines: what do we know about efficacy?
Vaccine-mediated protection has been associated with both humoral (neutralizing antibodies) and adaptive immune (T cells) responses to the spike (S) protein. The role of innate immunity in shaping B cell and T cell responses and directly providing protection remains underappreciated and is being explored through the development of new adjuvants. Even with the highly effective mRNA vaccines, antibody responses wane after 6 months, with binding and neutralizing antibodies estimated to have a half-life of between 52 and 68 d, respectively9,10. Interim results on the Ad26.COV2.S vaccine suggested that binding and neutralizing antibody responses may be stable through 8 months, but this study is limited by a small sample size (n = 10) and does not directly compare these responses to those generated by other vaccine platforms11.
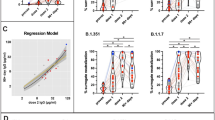
Durable antibody responses are mediated through memory B cells and long-lived plasmablasts. These subsets continuously produce neutralizing antibodies and non-neutralizing antibodies that will mobilize effector functions of innate immune cells such as antibody-dependent cellular cytotoxicity and antibody-dependent neutrophil phagocytosis12,13. This contributes to protection during acute infection and perhaps reduce the risk of reinfection. Although, on average, the vaccines induce production of sufficient antibody titers to protect against disease (particularly in people who are immunocompetent), the response to each vaccine is heterogenous, with antibody titers varying over a range of two to three orders of magnitude in adults immunized with the same vaccine. Similar variability is observed after boosting with the same or different vaccines. Notably, those who showed brisk responses after priming were also those who had the most potent responses after the boost, suggesting that the heterogeneity is due, in part, to the host environment.
Mutations within the N-terminal domain and receptor-binding domain, including a p.Glu484Lys mutation found within the B.1.351 (beta), P.1 (gamma), B.1.617.1 (kappa) and B.1.621 (mu) variants, reduce the neutralizing activity of vaccine-induced antibodies against these variants14,15,16,17. More recent studies have identified a substantial reduction in vaccine efficacy when infected with the B.1.617.2 (delta) variant, dropping to as low as 50% depending on the vaccine and population studied18. The B.1.1.529 variant (omicron) is highly infectious and appears to evade natural and perhaps vaccine-induced immunity, again raising concern about a new wave of infections that may be more resistant to vaccine-mediated protection19,20. Recent preliminary reports have suggested that the neutralizing activity of vaccine-induced antibodies is lower against omicron21,22. There may be a consensus emerging that SARS-CoV-2 is now endemic, and that we will be dealing with the emergence of new immune-evading variants for years.
Studies have identified substantial variability in the variant coverage of vaccinees among individuals who received the same vaccine. Understanding the impact of biologic and environmental/geographical factors on qualitative and quantitative features of the host environment and their impact on the innate immune response and subsequently on the breadth (variant coverage) and durability of long-lived plasmablasts as well as memory B cell and T cell responses is critical in improving our understanding and development of next-generation SARS-CoV-2 vaccines.
The global reach of SARS-CoV-2 means that the virus spreads in distinct geographical settings, each endowed with environmental (for example, microorganisms, endemic infection and coinfections) or cultural (diet) factors that might affect vaccine efficacy. These features modulate immune responses to vaccination, as has been reported for pneumococcal disease23,24, influenza25 and malaria26. Distinct viral variants are prevalent in different geographical settings, which raises challenges about vaccine coverage and vaccine durability and argues for the potential development of regional vaccines. A universal coronavirus vaccine that protects against future variants and perhaps other coronavirus strains may be necessary to fully control the ongoing pandemic and prevent future ones. Targeting the development of vaccines and their adjuvants on pathways that can trigger innate immune mechanisms of antiviral protection is one approach that can lead to universal protection. Studies of individuals who survived SARS-CoV-1 or were infected by SARS-COV-2 and then received a SARS-CoV-2 vaccine provide some proof of concept that such a vaccine may be feasible27. Comparing the innate immune response of SARS-CoV-2-infected convalescent individuals to the innate immune response triggered by the different vaccines will shed light on the innate signaling pathways that lead to long-lasting immune protection.
The success of current vaccines needs to be taken with a word of caution. The efficacy of these vaccines is assessed within a few months of receiving the full vaccine regimen, when it would be anticipated that a study participant has peak immune responses. This contrasts with most vaccines whose efficacy is monitored for years following vaccination. Although time and logistics dictated measuring SARS-CoV-2 vaccine efficacy in such a short window to prevent further uncontrolled spread and overload of healthcare systems, we will likely learn that the approaches taken initially were suboptimal; indeed, the spacing between the Pfizer (3 weeks) and Moderna (4 weeks) vaccinations is now known to have been too short28. The recent approval of booster shots for all adults in the Unites States highlights the potential for waning immunity and that initial efficacy rates for the vaccines may not hold up over time. The durability of the antibody response to SARS-CoV-2 vaccines is variable across platforms, with the adenovirus vaccines inducing lower magnitude yet more stable antibody response than the mRNA vaccines; the Moderna vaccine appears to induce more durable humoral responses than Pfizer, presumably because a higher dose and a longer time between prime and boost were used. Thus, the short-term and long-term immune responses induced by vaccination can be dependent on the vaccine platform, although several independent factors can contribute to the heterogeneity in the antibody and cellular responses. Hence, although the development of effective vaccines within a year of the outbreak was an unquestioned success, much work still needs to be done if we are to come up with a strategy that provides durable and broad protection against current and future variants. To reach this milestone, we need to decipher the mechanisms which lead to long-lived and protective immune responses in vaccination and how these are impacted by host and environmental factors.
Innate immune activation is differentially modulated by vaccine platform
In a canonical vaccine response, the initial immune response is mediated by antigen-presenting cells (APCs) at the site of immunization. These APCs encounter antigen, migrate to draining lymph nodes and present antigen to T cells and B cells. The immunological outcome of this interaction among APCs, T cells and B cells is shaped in part by the impact of the vaccine (vector, immunogen and adjuvant) and host environment on the innate immune signaling cascades (that is, toll-like receptors (TLRs), retinoic acid-inducible gene I (RIG-I)-like receptors (RLRs), DNA sensors and nucleotide oligomerization domain (NOD)-like receptors (NLRs)). These cascades trigger distinct transcriptional networks in the APCs, mainly dendritic cells (DCs) and macrophages29,30,31.
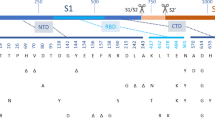
Potential and/or known sensors of SARS-CoV-2 vaccine platforms are summarized in Table 1. mRNA vaccines can activate TLR7, although current formulations used in Moderna and Pfizer are designed to minimize this activation, and melanoma-associated disease protein 5 (MDA5; which sense single-stranded RNA)32,33. The lipid nanoparticles used to deliver RNA vaccines activate TLR434,35. Adenovirus and adenoviral vector DNA are sensed by TLR932,36,37 and cyclic GMP-AMP synthase (cGAS)/stimulator of interferon genes (STING)38, while the inactivated SARS-CoV-2 virus vaccines are sensed by TLR739,40,41, MDA5 and RIG-I, which signal through mitochondrial antiviral signaling protein (MAVS)42,43,44. One report identified TLR2 as a possible sensor of the SARS-CoV-2 envelope45. Additionally, the adjuvant alum used in the inactivated viral vaccines activates the NLR family pryin domain containing 3 (NLRP3) inflammasome46,47. Activation of these pathways promotes the expression of distinct gene sets with immunomodulatory functions with TLR7 and TLR9 predominantly activating NF-κB (nuclear factor kappa-light-chain-enhancer of activated B cells), while TLR4, RIG-I, MDA5 and STING activate both NF-κB and interferon regulatory factors (IRFs). The specific transcriptional programs that are activated in innate immune cells will determine the quality and the magnitude of the adaptive immune response48. The balance of these signals shapes the differentiation of effector cells of the adaptive immune system and the degree to which any preexisting immunity is boosted. To develop an improved vaccine strategy, we will need to determine which of these pathways should be targeted, and how to target them most effectively.
APCs not only present antigen to T cells and B cells but they also express ligands for activating or inhibitory receptors present on B cells and T cells; moreover, they produce cytokines that functionally modulate the skewing of T cell responses, antibody class switching in B cells and the establishment of long-lived plasma cells and memory T cell and B cell responses49,50,51,52. The quality of the innate immune response affects all these APC–T/B cell interactions and thus shapes the degree to which an adaptive immune response will be protective and durable. It has been shown that activation of the CREB transcriptional program in innate immune cells results in the production of a network of chemokines and cytokines that recruit effector T cells and B cells producing antibody specific for the pathogen53. It is reasonable to assume that the heterogeneity in the responses to each individual vaccine is due in part to differences in innate immune responses, but this remains to be determined empirically and validated mechanistically.
Innate immunity and protective immunity: cannot have one (protective) without the other (innate)
Sensing of viral RNA and DNA by innate immune receptors including TLRs, RLRs and DNA sensors leads to the activation of immune transcription factors including IRFs and NF-κB. Activation of these transcription factors leads to production of chemokines that promote immune cell migration to the site of immunization or lymph nodes, the upregulation of viral restriction factors, and the production of immunomodulatory cytokines (Fig. 1). One way that cytokine production can directly enhance antiviral immunity and protect from viral challenge is through the upregulation of viral restriction factors downstream of cytokine signaling. This can happen in the cell producing the cytokine (autocrine) or in neighboring cells (paracrine) allowing cytokines to spread antiviral immunity to new cells. STING signaling not only mediates activation of antiviral immunity within a cell but also has been linked with ‘infectious’ spread of this antiviral immunity to neighboring cells. In this process, the dicyclic nucleotide cGAMP (an intracellular small molecule that normally activates STING signaling) is transferred from one cell to another, activating STING in the recipient cell that occurs through gap junctions54. Accordingly, spreading of cGAMP signaling is a critical driver of protective HIV-1 vaccine responses in humans and nonhuman primates53. The cGAS–cGAMP–STING axis provides another mechanism of inducing and spreading antiviral immunity in response to innate immune stimuli.
Hematopoietic and nonhematopoietic cells utilize multiple signaling cascades to recognize the presence of viral infection and modulate host responses. These innate pathways lead to the activation of key immune transcription factors driving the production of chemokines to promote migration, cytokines to activate cells and intrinsic/autocrine/paracrine induction of critical viral restriction factors that directly inhibit viral infection and function. IFITM, interferon-induced transmembrane proteins; IFIT, interferon-induced protein with tetratricopeptide repeats; APOBEC, apolipoprotein B mRNA editing enzyme, catalytic polypeptide; OAS, 2′-5′-oligoadenylate synthetase; Mx, Mx dynamin-like GTPase.
Upregulation of viral restriction factors by these nucleic acid innate sensors presumably helps by protecting immune cells from the detrimental effects of any current or future infection events. Indeed, vaccines and adjuvants modulate intrinsic innate antiviral immunity (IIAVI)55. IIAVI refers specifically to the induction of antiviral genes, and immune pathways that restrict viral replication and render cells ‘refractory’ to viral infection and persistence55,56,57. Antiviral genes promote this antiviral state by inhibiting viral entry, transcription and translation and by degrading or mutating viral nucleic acids (Fig. 1). Stromal cells, including lung epithelium58,59, also express such antiviral genes. In this respect, it has been shown that the adjuvant ASO3, when used in the trivalent influenza vaccine, induces chromatin remodeling of loci of antiviral genes and that this renders the cells refractory to viral infection55. It is noteworthy that these pathways are activated by components of SARS-CoV-2 vaccines (Table 1) suggesting that IIAVI may contribute to the effectiveness of these vaccines. To move SARS-CoV-2 vaccine development forward, we need to define the mechanisms that underlie the capacity of the different vaccine platforms to modulate the induction and long-term maintenance of IIAVI that will be critical for protecting individuals from future challenges. This is especially true for the control of viral variants as the innate immune response is not antigen specific and should not be affected by mutations in key SARS-CoV-2 antigens that impact on adaptive immune protection.
A recent study of the Pfizer-BioNTech vaccine showed that inflammatory and antiviral responses were more potently induced after the third booster dose vaccine compared to primary immunization60. In fact, there was little evidence of inflammatory transcriptional signatures after the initial dose of vaccine. Cells of the innate immune response, specifically CD14+CD16+ inflammatory monocyte frequencies, were heightened after boosting. This coincided with the augmented expression of inflammatory transcriptional signatures, including TLR pathways, and induction of antiviral and interferon (IFN) pathways; augmented inflammatory and IFN pathway activity was observed in multiple innate immune subsets including DCs and monocytes. The authors compared the 7-d post-boost signature of the Pfizer-BioNTech vaccine with signatures induced after other vaccines (including influenza, yellow fever and HIV) and found there was minimal overlap in the signatures expressed in participants who received the Pfizer-BioNTech vaccine when compared with those who received the other vaccines, although there was overlap between the other vaccine platforms. What this means for SARS-CoV-2 vaccine responses remains to be determined but it does show that the mRNA platform is inducing a different post-boost signature than what has been reported for other vaccines. The lack of a persistently induced signature of inflammation, which is a common feature of efficacy identified in other vaccines61, could explain the lack of durability of both effector arms of adaptive immunity observed with the Pfizer-BioNTech vaccine strategy and perhaps other SARS-CoV-2 vaccines.
Memory: it is not just for T cells and B cells anymore
The newly emerging field of trained immunity62, or the acquisition of ‘memory’ by innate immune cells such as monocytes/macrophages and DCs63, provides new opportunities for personalized vaccination. Previous exposure to microorganisms and pathogens can shape the innate immune response in a way that it is primed to be refractory, or susceptible, to homologous and heterologous pathogens. Activation of pattern-recognition receptors (PRRs) leads to transcriptional upregulation of target genes that result in the modulation of the epigenetic landscape of the cell64,65. Transcriptional and epigenetic changes to genetic loci involved in immune responses (IFNs, viral restriction factors and inflammatory cytokines) can result in these regions remaining in an ‘open’ (potentiated expression) or ‘closed’ (attenuated expression) chromatin state55. Accessibility of transcription factors to their cognate gene targets results in a more rapid and potent response to future stimulation to the same signals, essentially the innate immune response version of T cell and B cell memory. Importantly, trained immunity has been demonstrated to enhance immune responses in the context of vaccination, initially with Bacillus Calmette–Guérin (BCG)66,67,68 and the diphtheria, tetanus and pertussis (DTP) vaccines66 and more recently with the influenza vaccines55. Studies have been performed to identify pre-vaccination signatures that predict vaccine efficacy and outcome69,70,71,72. Other studies have linked microbiota, metabolism, epigenetics and trained immunity shedding insight into the links of these critical pathways73,74,75,76. Thus, an individual’s unique environmental and microbial exposure will define the nature of their future innate immune responses, including to vaccination, and potentially impact the efficacy of the generated immune responses. Investigating how SARS-CoV-2 vaccines are impacted by or modulate the activation and maintenance of trained immunity will substantially improve the development of the next generation of vaccines. Given the intra-platform variability in SARS-CoV-2 vaccine responses and efficacy, it is imperative that holistic approaches are used to define how individual variations in past and present microbial exposure modulate the establishment and maintenance of trained immunity and IIAVI.
Microbiome and metabolite compositions definitively shape host immune responses during vaccination
Environmental factors that can impact on immune responses to infection and vaccination include the microbiome. The microbial community can modulate immune responses during vaccination through interaction with PRRs on immune cells or indirectly via modulation of the homeostasis of immune cell subsets by metabolites (Fig. 2). This is very relevant to the vaccine efforts that are taking place to curb the SARS-CoV-2 pandemic as different vaccine platforms are being used in different geographical areas that all have distinct microbial environments. Theoretically, geographical differences in the microbiome may prove to be an important determinant of the response to SARS-CoV-2 vaccines.
Under normal homeostatic conditions, limited sensing of the microbiome and its derived metabolites promotes a tolerogenic intestinal immune response. Perturbations in the microbiome, which can be associated with diet, antibiotics or barrier disruption, lead to aberrant sensing of the microbiome and/or altered metabolite profiles that directly modulate the function of innate and adaptive immune cells within the intestines (mucosa-associated lymphoid tissue (MALT)) and draining lymph nodes (LNs). Metabolites can directly induce inflammatory signaling and cytokine production in innate immune cells (DCs, macrophages and innate lymphoid cells (ILCs)), which regulate interaction with and activation of adaptive immune cells (T cells and B cells) leading to alterations in the induction, characteristics and potentially longevity of the cellular and humoral immune responses. Mφ, macrophage; MLN, mesenteric lymph node; PSA, polysaccharide antigen A; SAA, serum amyloid protein A; TNF, tumor necrosis factor.
The gut epithelium and mucus layers provide a barrier between host tissue and this microbial community. Immune cells in the mucosa are constantly sampling antigens from the mucosal lumen and loss in mucosal barrier integrity caused by pathogenic infection, physical injury or dysregulated tissue inflammation leads to direct interaction of host immune cells in the mucosa with microbial communities. Importantly, immunocompromised individuals have ‘leaky’ gut that results from chemotherapy or opportunistic infections, which cause higher levels of microbial translocation that trigger pathogenic pro-inflammatory responses by immune cells. Of note, many of these individuals mount poor responses to SARS-CoV-2 vaccines77,78,79.
Depletion of the commensal microbiome by antibiotics impairs the response to the influenza vaccine80. Sensing of microbial-derived flagellin by TLR5 augmented antibody responses81 after influenza vaccination; the mechanisms triggered by engagement of TLR5 during the response to vaccination remain unknown. Helminths trigger perturbations in the homeostasis of effector and memory cells of the adaptive immune response as they skew the immune response toward type 2 helper T (TH2) cell responses82. Microorganisms and bacteria also impact interleukin (IL)-17-producing helper T (TH17) cell differentiation and the development of regulatory T (Treg) cells. Exploiting or controlling microbial modulation of innate and adaptive immune responses should be included in all strategies to develop improved and universal SARS-CoV-2 vaccines.
Host- and microbial-derived metabolite signatures are potent modulators of immune responses to vaccination
This systemic impact of microorganisms is also mediated by microbial production of metabolites that are produced locally in the mucosa and then taken up into systemic circulation. Many of these metabolites are important for normal cellular and tissue function, while others have detrimental impact on cellular function, including immune homeostasis and effector function. For example, some gut microorganisms (Fig. 2) produce short-chain fatty acids (SCFAs)83 such as butyrates and propionates. SCFAs are generally inhibitors of histone deacetylases (enzymes responsible for deacetylating histones leading to closed chromatin) and can regulate gene expression via activation of G-protein-coupled receptors84. SCFAs have been demonstrated to promote the chemotaxis and activation of the pro-inflammatory neutrophils85,86. SCFAs, bile acids and tryptophan metabolites mediate enhanced Treg cell differentiation by augmenting FOXP3 expression and transforming growth factor-β (TGF-β)87. Treg cells are known to be associated with the production of TGF-β and IL-10, two anti-inflammatory cytokines that have been shown to curb the response to vaccines.
The composition of mucosal and circulating metabolites is also influenced by nutrition and diet. Notably, this is true for SCFAs/butyrates88,89 and bile acids90,91,92, which have been shown to impede the response to vaccines80,93, presumably by triggering Treg cell differentiation and the production of the anti-inflammatory cytokines IL-1094,95,96 and TGF-β97,98,99. High glucose levels provide another source of immune modulation. It is well established that diabetes mellitus and elevated blood glucose are associated with poor outcomes in COVID-19100,101,102; however, the impact of high glucose levels on vaccine responses is poorly understood. Elderly individuals with diabetes had higher pneumococcal vaccine efficacy than elderly participants without diabetes103, while no differences were observed for influenza vaccine efficacy104. O-GlcNAcylation, the post-translational addition of an O-GlcNAc subunit (a derivative of glucose metabolism that rises during hyperglycemia) to serine (Ser) and threonine residues, of NF-κB c-Rel Ser350 can disturb CD4+ T cell homeostasis by enhancing type 1 helper T (TH1) cell effector function (IL-2 and IFN-γ) and suppressing Treg cells via downregulation of FOXP3, the key transcription factor regulating Treg cell development and function105,106. These studies demonstrated that O-GlcNAcylation of c-Rel regulated its binding to the promoter of target genes, a mechanism that modulated increased (more binding) and decreased (less binding) gene expression. This dysregulated balance between effector and Treg cell function could have substantial implications for the generation of immune responses during vaccination. This balance has been mostly studied in the context of tuberculosis vaccination in mice where Treg cells suppress TH1 and vaccine response107,108,109.
This capacity for metabolites to act on both the innate and adaptive arms of the immune system centrally positions them as critical modulators of systemic immune responses needed to promote vaccine efficacy. Individual variation in the levels of microorganisms and their metabolites presumably explains, in part, the multiple orders of magnitude of variation in the antibody responses to various SARS-CoV-2 vaccines. Although several groups have studied the impact of the microbiome and its metabolites on vaccine responsiveness (reviewed in refs. 110,111), we do not have a clear understanding of the impact on SARS-CoV-2 vaccine responses. The pandemic nature of SARS-CoV-2 infection provides a further impetus for considering the impact of these environmental factors on SARS-CoV-2 vaccine responses
How to overcome poor vaccine responses in immunocompromised individuals
Although our understanding of the role of the microbiome and metabolome in modulating immune function and vaccine responsiveness has improved substantially in the past few years80,112, these findings have yet to be translated into the clinical setting to enhance vaccine modalities (current studies summarized in ref. 111). With a deeper understanding of the multitude of factors that contribute to determining the efficacy of vaccines at an individual level, one can then begin to devise strategies that mitigate the heterogeneity in responses and bolster the population-level efficacy of vaccines. Some of these are clear and potentially easy solutions, others are less so and will require (again) the combined ingenuity of the biomedical research community.
There are at least two issues that might be prioritized in vaccine development now: understanding why older and/or immunocompromised individuals often demonstrate poor responses to vaccines and untangling how environmental factors (including the microbiome and its metabolites) shape these responses. There is substantial evidence that immunocompromised individuals exhibit diminished vaccine responses to SARS-CoV-2. For example, individuals with diseases such as common variable immunodeficiency113, rheumatoid arthritis and systemic lupus erythematosus exhibit lower seroconversion and antibody responses to vaccines114,115,116,117,118. This is due, at least in part, to the effects of the use of immunosuppressive drugs119,120,121,122,123. Patients with cancer124,125, in particular those with hematologic malignancies126,127,128,129, are another group of people who show less effective vaccine responses to SARS-CoV-2. As with autoimmune diseases, the reduced seroconversion rates and antibody responses are commonly linked to the use of anticancer therapies130,131,132,133. Following this same theme, patients receiving organ transplants, who need to take immunosuppressive drugs to prevent rejection, also exhibit reduced antibody responses to SARS-CoV-2 vaccination134,135,136,137,138,139,140,141,142. In addition to reduced antibody responses, patients receiving organ transplants also exhibit reduced SARS-CoV-2-specific CD4+ and CD8+ responses as measured by reduced/absent frequencies and substantially diminished ex vivo cytokine production in response to peptide stimulation134,137,138,139,142. Immunosuppressive drugs not only act directly on T cells and B cells, but they also suppress the activity of innate immune cells, which further dampens the induction of T and B cell responses. The largest challenge facing all of these vulnerable populations is that the treatments that are causing reduced efficacy are necessary for quality of life and, in many cases, survival. Simply removing individuals off of these therapies is not a viable option.
The most straightforward approach, which is currently being used, is to administer booster shots that further jump-start the immune response and boost effector and memory responses. This approach, however, is more of a bandage that does not mitigate the underlying causes of poor immunity and requires continuous administration of shots. Another critical unmet need pertains to those who are immunocompromised; such individuals are more susceptible to severe disease in addition to reduced antibody responses to vaccination. As the virus can persist and replicate much longer, the immunocompromised are assumed to be a major source for the emergence of new variants. An effective vaccine for the immunocompromised hence would go a long way to getting us to the goal of ending the SARS-CoV-2 pandemic. Using a comprehensive approach to define the immune pathways and mechanisms, including innate immune responses, that promote long-lived B cell and plasma cell memory responses and T stem cell responses, and how these are perturbed by age and/or in immunocompromised individuals, would provide the opportunity to enhance the efficacy of normal vaccine regimens and require fewer, if any, booster shots. Both the vaccine platform and the adjuvant used can be formulated to potentiate memory responses. Conceptual targets include those activating the WNT or NOTCH pathways that are associated with cell survival and stemness (that is, the development of long-term memory B cells and T cells) as well as potent activators of NF-κB and IRFs, which would prime innate immune responses and promote epigenetic modifications that could lead to long-term cell survival of antigen-specific T cells and B cells and resistance to infection.
For SARS-CoV-2 variants, utilizing vaccine platforms that incorporate multiple antigens that cover common variants, like how yearly influenza shots are handled, or using highly conserved antigen(s) unlikely to undergo immune escape are two straightforward ways to expand immune protection to include current and future variants. One critical factor to consider when addressing vaccine efficacy to variants is the differential distribution and circulation levels of SARS-CoV-2 variants based on geography. Knowing which platforms cross-neutralize which variants will be critical for ensuring that the right vaccines get to the right populations. Another critical factor that will likely need to be considered is ‘original antigenic sin’, or the effect that prior immunization or natural infection has in skewing the antigen-specific T cell and B cell response toward the original antigen, thus preventing the development of responses to new antigens and hence new variants. How innate signaling and environmental factors influence original antigenic sin and/or how manipulation of the pathways they modify might overcome this barrier is unknown and deserves to be prioritized in future studies. A more contemporary approach to overcome the issue of variant-associated reductions in vaccine efficacy relies on the identification of vaccine platforms and/or components that potently drive IIAVI in tissues like the lung. The mechanisms underlying IIAVI are not antigen specific, and hence their activation is not likely to be substantially impacted by genetic variants of SARS-CoV-2. Identifying and designing vaccines that boost IIAVI is critical for enhancing the cross-protection potential of vaccines. Immunomodulatory compounds that promote IIAVI will also be critical for future pandemic preparedness as treatment with these compounds may function as a first line of defense to limit infection/transmission until vaccines become available.
Mitigating the impact of environment factors like the microbiome and metabolome on innate and adaptive immune functions and vaccine responses will prove much more difficult than dealing with waning immunity or variants. Identifying pre-vaccination signatures and mechanisms that are associated with potent and durable vaccine responses and that are independent of the vaccine platform could provide a path to the development of interventions that can exploit or mimic such conditions. Modulation of immune responses can be achieved by combining the vaccines with immunomodulatory stimuli that are selected based on a mechanistic rationale. For example, a high level of NF-κB has been identified as a prerequisite for the induction of potent responses to 13 different vaccines61. Should these observations be confirmed, adjuvants and PRR ligands that trigger the NF-κB pathway (that is MF59, AS03, TLR ligands NOD1 and/or NOD2) might be studied as vaccine adjuvants. It has also been shown that vectors or adjuvants that trigger the cGAS–cGAMP pathway induce (1) a cascade of innate chemokines/cytokines that recruit T cells and B cells to APCs that express vaccine antigens and (2) subsequent adaptive immune pathways. Metformin and rapamycin, inhibitors of mammalian target of rapamycin signaling, have also been shown to reverse the specific immune dysfunction associated with aging and trigger higher antibody responses and improved cellular immunity to several vaccines including influenza143,144,145,146,147. Both rapamycin and metformin are well known to impact transcriptional, functional and epigenetic profiles of innate immunity triggered by the microbiome and/or metabolites148,149,150,151,152. This concept of personalized vaccination has been suggested before (topic reviewed in ref. 153) and could be a critical addition to vaccine strategies and public health policy to reach the end goal of population-level vaccine-induced immunity to SARS-CoV-2. The administration of prebiotics (nutrients which promote a healthy microbiota), probiotics (beneficial microbiota) and/or postbiotics (by-products of probiotic bacteria) could also be used to help overcome dysregulated immune responses resulting from the composition of the microbiome. Current studies of microbiota manipulation in augmenting vaccines are inconsistent, with an equivalent number of studies showing an effect as the number of studies that show no effect110,154. Differences in vaccine platforms, trial populations and the species of prebiotics or probiotics used contribute to these inconsistent findings154,155. Although such interventions could not realistically be implemented globally in the context of a pandemic, they could be used to help improve vaccine responsiveness to specific populations, including the elderly, those that are immunocompromised and those that have previously demonstrated a suboptimal vaccine response.
Conclusions
The presence of comorbidities, individual differences in microbial/metabolite composition and individual variations in the response of the innate immune system each contribute to the heterogeneity in vaccine responses. Multiple mechanisms account for how these factors shape vaccine effectiveness. For example, the capacity for microbial products and metabolites to impact epigenetics—a long-term regulator of gene expression—in both innate and adaptive immune cells strongly suggests that differences in the microbiome/metabolome between individuals can have long-lasting impacts on modulating immune responses to vaccines. The microbiome as well as the host–microbial metabolome can also impact on IIAVI and the capacity of the innate immune response to impede the initial stages of viral infection and dissemination. In the most severe of instances (Fig. 3), altered microbiome and metabolome composition substantially impairs the induction of an innate immune response by dysregulating antigen processing/presentation and the expression of cytokines, preventing the acquisition of T cell and B cell effector and memory function, and altering the magnitude and isotype class switching of antibodies.
Environmental factors can lead to alterations in the microbiome and metabolite composition, which are sensed by the innate immune system. This can lead to impaired innate immune function, disrupted antigen presentation and improper skewing of adaptive (T cell and B cell) responses. In this environment of dysregulated adaptive immune activation, antigen-specific responses induced during vaccination are functionally altered compared to those generated under homeostatic conditions. This can not only impact generating a protective vaccine response but also directly modulates the induction of memory responses, which substantially dictates the longevity of vaccine-induced immunity. In this way, differences in the combination of environmental factors between individuals can substantially impact on both the primary generation of a vaccine response and how long an individual is protected from future infection. AA, amino acid; Teff cells, effector T cells; TFH cells, follicular helper T cells; TM cells, memory T cells; GC, germinal center.
Targeting the innate immune system as a means to improve SARS-CoV-2 vaccine efficacy is a promising avenue of investigation. The innate immune system functions as the foundation for the induction of adaptive immunity: processing and presenting the vaccine antigen to T cells and B cells, and producing the chemokines and cytokines required to bring lymphocytes into sites of immunization and skew appropriate T cell and B cell responses. Thus, the innate immune system modulates the quantity (magnitude) and the quality (that is, B cell maturation and antibody isotype switching) of vaccine responses. Additionally, both molecular and cellular innate immune pathways are highly conserved across the human population, and hence easier to target therapeutically, at least as compared to the far more heterogenous adaptive immune response, where factors such as HLA background and prior exposures can result in highly variable vaccine outcomes. Using different adjuvants or altering the immunogenic component(s) of vaccines (for example, the CpG motifs in DNA vaccines) represents the best way of directly targeting innate immunity during vaccination. Targeting conserved innate pathways upstream of adaptive immunity might prove to be an effective and scalable way to generate improved vaccines. To further define the complex impact of innate signaling on adaptive immunity and vaccine effectiveness, well-powered controlled clinical studies of potential vaccine adjuvants are needed. Now that we have effective options, the field can afford to slow down and begin the hard work of determining how host environmental factors, different adjuvants and different vaccine platforms compare head-to-head. Recent efforts to improve such comparative research in vaccines is moving the field forward but we are still ‘playing catch-up’. If we are to ever achieve the goal of a vaccine that protects an overwhelming majority of the human population against existing and yet-to-emerge variants, we need to get ahead of SARS-CoV-2 instead of chasing it.
If the factors discussed in this Perspective and specifically their impact on innate immunity are not accounted for when evaluating the efficacy of vaccines, it could lead to disparate healthcare outcomes that may impact substantially on vulnerable and/or underserved populations. Ensuring effective vaccines are administered to all regardless of age, preexisting conditions, biological sex, race/ethnicity or geographical locale is the only way to contain and potentially eliminate SARS-CoV-2. Further research and the implementation of controlled clinical studies are needed to investigate how individual differences in innate immune function, resulting from vaccine-associated factors as well as host and environmental factors, mechanistically modulate cellular and humoral immune responses to SARS-CoV-2 vaccination. Studying innate immunity as part of a holistic approach integrating all the components that can impact on vaccine responses is a pathway to developing a more universal and effective vaccine to help end this deadly pandemic.
References
Trovato, M., Sartorius, R., D’Apice, L., Manco, R. & De Berardinis, P. Viral emerging diseases: challenges in developing vaccination strategies. Front. Immunol. 11, 2130 (2020).
Baden, L. R. et al. Efficacy and safety of the mRNA-1273 SARS-CoV-2 vaccine. N. Engl. J. Med. 384, 403–416 (2021).
Polack, F. P. et al. Safety and efficacy of the BNT162b2 mRNA COVID-19 vaccine. N. Engl. J. Med. 383, 2603–2615 (2020).
Sadoff, J. et al. Safety and efficacy of single-dose Ad26.COV2.S vaccine against COVID-19. N. Engl. J. Med. 384, 2187–2201 (2021).
Voysey, M. et al. Safety and efficacy of the ChAdOx1 nCoV-19 vaccine (AZD1222) against SARS-CoV-2: an interim analysis of four randomised controlled trials in Brazil, South Africa and the UK. Lancet 397, 99–111 (2021).
Zhang, Y. et al. Safety, tolerability and immunogenicity of an inactivated SARS-CoV-2 vaccine in healthy adults aged 18–59 years: a randomised, double-blind, placebo-controlled, phase 1/2 clinical trial. Lancet Infect. Dis. 21, 181–192 (2021).
Xia, S. et al. Safety and immunogenicity of an inactivated SARS-CoV-2 vaccine, BBIBP-CorV: a randomised, double-blind, placebo-controlled, phase 1/2 trial. Lancet Infect. Dis. 21, 39–51 (2021).
Heath, P. T. et al. Safety and efficacy of NVX-CoV2373 COVID-19 vaccine. N. Engl. J. Med. 385, 1172–1183 (2021).
Doria-Rose, N. et al. Antibody persistence through 6 months after the second dose of mRNA-1273 vaccine for COVID-19. N. Engl. J. Med. 384, 2259–2261 (2021).
Suthar, M. S. et al. Durability of immune responses to the BNT162b2mRNA vaccine. Med 3, 25–27 (2022).
Barouch, D. H. et al. Durable humoral and cellular immune responses 8 months after Ad26.COV2.S vaccination. N. Engl. J. Med. 385, 951–953 (2021).
Kaplonek, P. et al. Subtle immunological differences in mRNA-1273 and BNT162b2 COVID-19 vaccine induced Fc-functional profiles. Preprint at bioRxiv https://doi.org/10.1101/2021.08.31.458247 (2021).
Tauzin, A. et al. A single dose of the SARS-CoV-2 vaccine BNT162b2 elicits Fc-mediated antibody effector functions and T cell responses. Cell Host Microbe 29, 1137–1150 (2021).
Pegu, A. et al. Durability of mRNA-1273-induced antibodies againstSARS-CoV-2 variants. Science 373, 1372–1377 (2021).
Edara, V. V. et al. Infection and vaccine-induced neutralizing-antibody responses to the SARS-CoV-2 B.1.617 variants. N. Engl. J. Med. 385, 664–666 (2021).
Edara, V. V. et al. Infection- and vaccine-induced antibody binding and neutralization of the B.1.351 SARS-CoV-2 variant. Cell Host Microbe 29, 516–521 (2021).
Messali, S. et al. A cluster of the new SARS-CoV-2 B.1.621 lineage in Italy and sensitivity of the viral isolate to the BNT162b2 vaccine. J. Med. Virol. https://doi.org/10.1002/jmv.27247 (2021).
Mostaghimi, D., Valdez, C. N., Larson, H. T., Kalinich, C. C. & Iwasaki, A. Prevention of host-to-host transmission by SARS-CoV-2 vaccines. Lancet Infect. Dis. https://doi.org/10.1016/S1473-3099(21)00472-2 (2021).
Karim, S. S. A. & Karim, Q. A. Omicron SARS-CoV-2 variant: a new chapter in the COVID-19 pandemic. Lancet https://doi.org/10.1016/S0140-6736(21)02758-6 (2021).
Torjesen, I. COVID-19: omicron may be more transmissible than other variants and partly resistant to existing vaccines, scientists fear. BMJ 375, n2943 (2021).
Khoury, D. et al. SARS-CoV-2 omicron has extensive but incomplete escape of Pfizer BNT162b2 elicited neutralization and requires ACE2 for infection. Preprint at https://www.medrxiv.org/content/10.1101/2021.12.08.21267417 (2021).
Widera, A. W. M. et al. Reduced neutralization of SARS-CoV-2 omicron variant by vaccine sera and monoclonal antibodies. Preprint at https://www.medrxiv.org/content/10.1101/2021.12.07.21267432 (2021).
Choe, Y. J., Blatt, D. B., Lee, H. J. & Choi, E. H. Associations between geographic region and immune response variations to pneumococcal conjugate vaccines in clinical trials: a systematic review and meta-analysis. Int. J. Infect. Dis. 92, 261–268 (2020).
Gjini, E. Geographic variation in pneumococcal vaccine efficacy estimated from dynamic modeling of epidemiological data post-PCV7. Sci. Rep. 7, 1–16 (2017).
Fink, A. L. et al. Regional differences in vaccine uptake and serological responses to vaccine and circulating strains of H1N1 viruses among patients with confirmed influenza. J. Clin. Virol. Plus https://doi.org/10.1016/j.jcvp.2021.100034 (2021).
Bei, A. K. et al. Functional analysis reveals geographical variation in inhibitory immune responses against a polymorphic malaria antigen. J. Infect. Dis. 216, 267–275 (2017).
Tan, C. W. et al. Pan-sarbecovirus neutralizing antibodies in BNT162b2-immunized SARS-CoV-1 survivors. N. Engl. J. Med. 385, 1401–1406 (2021).
Payne, R. P. et al. Immunogenicity of standard and extended dosing intervals of BNT162b2 mRNA vaccine. Cell 184, 5699–5714 (2021).
Kwissa, M., Nakaya, H. I., Oluoch, H. & Pulendran, B. Distinct TLR adjuvants differentially stimulate systemic and local innate immune responses in nonhuman primates. Blood 119, 2044–2055 (2012).
Agrawal, S. et al. Cutting edge: different Toll-like receptor agonists instruct dendritic cells to induce distinct TH responses via differential modulation of extracellular signal-regulated kinase-mitogen-activated protein kinase and c-Fos. J. Immunol. 171, 4984–4989 (2003).
Arunachalam, P. S. et al. Adjuvanting a subunit COVID-19 vaccine to induce protective immunity. Nature 594, 253–258 (2021).
Teijaro, J. R. & Farber, D. L. COVID-19 vaccines: modes of immune activation and future challenges. Nat. Rev. Immunol. 21, 195–197 (2021).
Pardi, N., Hogan, M. J., Porter, F. W. & Weissman, D. mRNA vaccines—a new era in vaccinology. Nat. Rev. Drug Discov. 17, 261–279 (2018).
Zhang, H. et al. Delivery of mRNA vaccine with a lipid-like material potentiates antitumor efficacy through Toll-like receptor 4 signaling. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2005191118 (2021).
Kedmi, R., Ben-Arie, N. & Peer, D. The systemic toxicity of positively charged lipid nanoparticles and the role of Toll-like receptor 4 in immune activation. Biomaterials 31, 6867–6875 (2010).
Zhu, J., Huang, X. & Yang, Y. The TLR9–MyD88 pathway is critical for adaptive immune responses to adeno-associated virus gene therapy vectors in mice. J. Clin. Invest. 119, 2388–2398 (2009).
Appledorn, D. M. et al. Adenovirus vector-induced innate inflammatory mediators, MAPK signaling, as well as adaptive immune responses are dependent upon both TLR2 and TLR9 in vivo. J. Immunol. 181, 2134–2144 (2008).
Lam, E., Stein, S. & Falck-Pedersen, E. Adenovirus detection by the cGAS/STING/TBK1 DNA sensing cascade. J. Virol. 88, 974–981 (2014).
Asano, T. et al. X-linked recessive TLR7 deficiency in ~1% of men under 60 years old with life-threatening COVID-19. Sci. Immunol. https://doi.org/10.1126/sciimmunol.abl4348 (2021).
Salvi, V. et al. SARS-CoV-2-associated ssRNAs activate inflammation and immunity via TLR7/8. JCI Insight https://doi.org/10.1172/jci.insight.150542 (2021).
Fallerini, C. et al. Association of Toll-like receptor 7 variants with life-threatening COVID-19 disease in males: findings from a nested case-control study. Elife https://doi.org/10.7554/eLife.67569 (2021).
Yang, D. M., Geng, T. T., Harrison, A. G. & Wang, P. H. Differential roles of RIG-I like receptors in SARS-CoV-2 infection. Mil. Med. Res. 8, 49 (2021).
Kouwaki, T., Nishimura, T., Wang, G. & Oshiumi, H. RIG-I-like receptor-mediated recognition of viral genomic RNA of severe acute respiratory syndrome coronavirus 2 and viral escape from the host innate immune responses. Front. Immunol. 12, 700926 (2021).
Yin, X. et al. MDA5 governs the innate immune response to SARS-CoV-2 in lung epithelial cells. Cell Rep. 34, 108628 (2021).
Zheng, M. et al. TLR2 senses the SARS-CoV-2 envelope protein to produce inflammatory cytokines. Nat. Immunol. 22, 829–838 (2021).
Franchi, L. & Nunez, G. The Nlrp3 inflammasome is critical for aluminium hydroxide-mediated IL-1β secretion but dispensable for adjuvant activity. Eur. J. Immunol. 38, 2085–2089 (2008).
Eisenbarth, S. C., Colegio, O. R., O’Connor, W., Sutterwala, F. S. & Flavell, R. A. Crucial role for the Nalp3 inflammasome in the immunostimulatory properties of aluminium adjuvants. Nature 453, 1122–1126 (2008).
Pulendran, B. & Ahmed, R. Translating innate immunity into immunological memory: implications for vaccine development. Cell 124, 849–863 (2006).
Ueno, H., Schmitt, N., Palucka, A. K. & Banchereau, J. Dendritic cells and humoral immunity in humans. Immunol. Cell Biol. 88, 376–380 (2010).
Diebold, S. S. Determination of T cell fate by dendritic cells. Immunol. Cell Biol. 86, 389–397 (2008).
Steinman, R. M. & Hemmi, H. Dendritic cells: translating innate to adaptive immunity. Curr. Top. Microbiol. Immunol. 311, 17–58 (2006).
Jego, G., Pascual, V., Palucka, A. K. & Banchereau, J. Dendritic cells control B cell growth and differentiation. Curr. Dir. Autoimmun. 8, 124–139 (2005).
Tomalka, J. A. et al. The transcription factor CREB1 is a mechanistic driver of immunogenicity and reduced HIV-1 acquisition following ALVAC vaccination. Nat. Immunol. 22, 1294–1305 (2021).
Ablasser, A. et al. Cell intrinsic immunity spreads to bystander cells via the intercellular transfer of cGAMP. Nature 503, 530–534 (2013).
Wimmers, F. et al. The single-cell epigenomic and transcriptional landscape of immunity to influenza vaccination. Cell 184, 3915–3935 (2021).
Kawamura, T., Ogawa, Y., Aoki, R. & Shimada, S. Innate and intrinsic antiviral immunity in skin. J. Dermatol. Sci. 75, 159–166 (2014).
Yan, N. & Chen, Z. J. Intrinsic antiviral immunity. Nat. Immunol. 13, 214–222 (2012).
Whitsett, J. A. & Alenghat, T. Respiratory epithelial cells orchestrate pulmonary innate immunity. Nat. Immunol. 16, 27–35 (2015).
Bals, R. & Hiemstra, P. S. Innate immunity in the lung: how epithelial cells fight against respiratory pathogens. Eur. Respir. J. 23, 327–333 (2004).
Arunachalam, P. S. et al. Systems vaccinology of the BNT162b2 mRNA vaccine in humans. Nature 596, 410–416 (2021).
Fourati, S. et al. An innate immune activation state prior to vaccination predicts responsiveness to multiple vaccines. Preprint at bioRxiv https://doi.org/10.1101/2021.09.26.461847 (2021).
Netea, M. G. et al. Defining trained immunity and its role in health and disease. Nat. Rev. Immunol. 20, 375–388 (2020).
Divangahi, M. et al. Trained immunity, tolerance, priming and differentiation: distinct immunological processes. Nat. Immunol. 22, 2–6 (2021).
Owen, A. M., Fults, J. B., Patil, N. K., Hernandez, A. & Bohannon, J. K. TLR agonists as mediators of trained immunity: mechanistic insight and immunotherapeutic potential to combat infection. Front. Immunol. 11, 622614 (2020).
van der Heijden, C. et al. Epigenetics and trained immunity. Antioxid. Redox Signal. 29, 1023–1040 (2018).
Stevens, N. E. et al. Immunisation with the BCG and DTPw vaccines induces different programs of trained immunity in mice. Vaccine https://doi.org/10.1016/j.vaccine.2021.03.084 (2021).
Vierboom, M. P. M. et al. Stronger induction of trained immunity by mucosal BCG or MTBVAC vaccination compared to standard intradermal vaccination. Cell Rep. Med. 2, 100185 (2021).
Arts, R. J. W. et al. BCG vaccination protects against experimental viral infection in humans through the induction of cytokines associated with trained immunity. Cell Host Microbe 23, 89–100 (2018).
Kardava, L. et al. Pre-vaccination and early B cell signatures predict antibody response to SARS-CoV-2 mRNA vaccine. Preprint at medRxiv https://doi.org/10.1101/2021.07.06.21259528 (2021).
Blanc, F. et al. Influence of genetics and the pre-vaccination blood transcriptome on the variability of antibody levels after vaccination against Mycoplasma hyopneumoniae in pigs. Genet. Sel. Evol. 53, 24 (2021).
Shannon, C. P. et al. Multi-omic data integration allows baseline immune signatures to predict hepatitis B vaccine response in a small cohort. Front. Immunol. 11, 578801 (2020).
Fourati, S. et al. Pre-vaccination inflammation and B cell signalling predict age-related hyporesponse to hepatitis B vaccination. Nat. Commun. 7, 10369 (2016).
Su, H. et al. miR-9-5p regulates immunometabolic and epigenetic pathways in beta-glucan-trained immunity via IDH3α. JCI Insight https://doi.org/10.1172/jci.insight.144260 (2021).
Lynch, S. V. & Vercelli, D. Microbiota, epigenetics and trained immunity. Convergent drivers and mediators of the asthma trajectory from pregnancy to childhood. Am. J. Respir. Crit. Care Med. 203, 802–808 (2021).
Fanucchi, S., Dominguez-Andres, J., Joosten, L. A. B., Netea, M. G. & Mhlanga, M. M. The intersection of epigenetics and metabolism in trained immunity. Immunity 54, 32–43 (2021).
Riksen, N. P. & Netea, M. G. Immunometabolic control of trained immunity. Mol. Aspects Med. 77, 100897 (2021).
Liu, C. et al. A retrospective analysis of COVID-19 mRNA vaccine breakthrough infections—risk factors and vaccine effectiveness. Preprint at https://www.medrxiv.org/content/10.1101/2021.10.05.21264583 (2021).
Lederer, K. et al. Germinal center responses to SARS-CoV-2 mRNA vaccines in healthy and immunocompromised individuals. Preprint at https://www.medrxiv.org/content/10.1101/2021.09.16.21263686 (2021).
Gavriatopoulou, M. et al. Poor neutralizing antibody responses in 106 patients with WM after vaccination against SARS-CoV-2: a prospective study. Blood Adv. https://doi.org/10.1182/bloodadvances.2021005444 (2021).
Hagan, T. et al. Antibiotics-driven gut microbiome perturbation alters immunity to vaccines in humans. Cell 178, 1313–1328 (2019).
Oh, J. Z. et al. TLR5-mediated sensing of gut microbiota is necessary for antibody responses to seasonal influenza vaccination. Immunity 41, 478–492 (2014).
Nutman, T. B. Looking beyond the induction of TH2 responses to explain immunomodulation by helminths. Parasite Immunol. 37, 304–313 (2015).
Krautkramer, K. A., Fan, J. & Bäckhed, F. Gut microbial metabolites as multi-kingdom intermediates. Nat. Rev. Microbiol. 19, 77–94 (2021).
Sun, M. et al. Microbiota-derived short-chain fatty acids promote TH1 cell IL-10 production to maintain intestinal homeostasis. Nat. Commun. 9, 3555 (2018).
Le Poul, E. et al. Functional characterization of human receptors for short-chain fatty acids and their role in polymorphonuclear cell activation. J. Biol. Chem. 278, 25481–25489 (2003).
Vinolo, M. A. et al. SCFAs induce mouse neutrophil chemotaxis through the GPR43 receptor. PLoS ONE 6, e21205 (2011).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Shimizu, H. et al. Dietary short-chain fatty acid intake improves the hepatic metabolic condition via FFAR3. Sci. Rep. 9, 16574 (2019).
Bach Knudsen, K. E. et al. Impact of diet-modulated butyrate production on intestinal barrier function and inflammation. Nutrients https://doi.org/10.3390/nu10101499 (2018).
Nakade, Y. et al. Characteristics of bile acid composition in high-fat-diet-induced nonalcoholic fatty liver disease in obese diabetic rats. PLoS ONE 16, e0247303 (2021).
Trefflich, I. et al. Associations between dietary patterns and bile acids—results from a cross-sectional study in vegans and omnivores. Nutrients https://doi.org/10.3390/nu12010047 (2019).
Just, S. et al. The gut microbiota drives the impact of bile acids and fat source in diet on mouse metabolism. Microbiome 6, 134 (2018).
Pelletier, A. N. et al. A prevaccination validated network that drives the breadth of the protective neutralizing antibody response following dengue vaccine TV003 immunization. Preprint at https://www.medrxiv.org/content/10.1101/2021.09.25.21264123 (2021).
Xu, M. et al. c-MAF-dependent regulatory T cells mediate immunological tolerance to a gut pathobiont. Nature 554, 373–377 (2018).
Chaudhry, A. et al. Interleukin-10 signaling in regulatory T cells is required for suppression of TH17 cell-mediated inflammation. Immunity 34, 566–578 (2011).
Rubtsov, Y. P. et al. Regulatory T cell-derived interleukin 10 limits inflammation at environmental interfaces. Immunity 28, 546–558 (2008).
Zemmour, D. et al. Single-cell gene expression reveals a landscape of regulatory T cell phenotypes shaped by the TCR. Nat. Immunol. 19, 291–301 (2018).
Li, M. O., Wan, Y. Y. & Flavell, R. A. T cell-produced transforming growth factor-β1 controls T cell tolerance and regulates TH1 and TH17 cell differentiation. Immunity 26, 579–591 (2007).
Marie, J. C., Letterio, J. J., Gavin, M. & Rudensky, A. Y. TGF-β1 maintains suppressor function and Foxp3 expression in CD4+CD25+ regulatory T cells. J. Exp. Med. 201, 1061–1067 (2005).
Logette, E. et al. A machine-generated view of the role of blood glucose levels in the severity of COVID-19. Front. Public Health 9, 695139 (2021).
Codo, A. C. et al. Elevated glucose levels favor SARS-CoV-2 infection and monocyte response through a HIF-1α/glycolysis-dependent axis. Cell Metab. 32, 437–446 (2020).
Caballero, A. E. et al. COVID-19 in people living with diabetes: an international consensus. J. Diabetes Complications 34, 107671 (2020).
Huijts, S. M., van Werkhoven, C. H., Bolkenbaas, M., Grobbee, D. E. & Bonten, M. J. M. Post-hoc analysis of a randomized controlled trial: diabetes mellitus modifies the efficacy of the 13-valent pneumococcal conjugate vaccine in elderly. Vaccine 35, 4444–4449 (2017).
McElhaney, J. E. et al. Predictors of the antibody response to influenza vaccination in older adults with type 2 diabetes. BMJ Open Diabetes Res. Care 3, e000140 (2015).
de Jesus, T. J. et al. Negative regulation of FOXP3 expression by c-Rel O-GlcNAcylation. Glycobiology 31, 812–826 (2021).
Ramakrishnan, P. et al. Activation of the transcriptional function of the NF-κB protein c-Rel by O-GlcNAc glycosylation. Sci. Signal. 6, ra75 (2013).
Ho, P., Wei, X. & Seah, G. T. Regulatory T cells induced by Mycobacterium chelonae sensitization influence murine responses to Bacille Calmette-Guerin. J. Leukoc. Biol. 88, 1073–1080 (2010).
Jaron, B., Maranghi, E., Leclerc, C. & Majlessi, L. Effect of attenuation of Treg during BCG immunization on anti-mycobacterial TH1 responses and protection against Mycobacterium tuberculosis. PLoS ONE 3, e2833 (2008).
Moore, A. C. et al. Anti-CD25 antibody enhancement of vaccine-induced immunogenicity: increased durable cellular immunity with reduced immunodominance. J. Immunol. 175, 7264–7273 (2005).
Lynn, D. J., Benson, S. C., Lynn, M. A. & Pulendran, B. Modulation of immune responses to vaccination by the microbiota: implications and potential mechanisms. Nat. Rev. Immunol. https://doi.org/10.1038/s41577-021-00554-7 (2021).
de Jong, S. E., Olin, A. & Pulendran, B. The impact of the microbiome on immunity to vaccination in humans. Cell Host Microbe 28, 169–179 (2020).
Li, S. et al. Metabolic phenotypes of response to vaccination in humans. Cell 169, 862–877 (2017).
Arroyo-Sanchez, D. et al. Immunogenicity of anti-SARS-CoV-2 vaccines incommon variable immunodeficiency. J. Clin. Immunol. https://doi.org/10.1007/s10875-021-01174-5 (2021).
Simader, E. et al. Importance of the second SARS-CoV-2 vaccination dose for achieving serological response in patients with rheumatoid arthritis and seronegative spondyloarthritis. Ann. Rheum. Dis. https://doi.org/10.1136/annrheumdis-2021-221347 (2021).
Ferri, C. et al. Impaired immunogenicity to COVID-19 vaccines in autoimmune systemic diseases. High prevalence of non-response in different patients’ subgroups. J. Autoimmun. 125, 102744 (2021).
Picchianti Diamanti, A. et al. Severe acute respiratory syndrome coronavirus-2 infection and autoimmunity 1 year later: the era of vaccines. Front. Immunol. 12, 708848 (2021).
Rubbert-Roth, A. et al. Anti-SARS-CoV-2 mRNA vaccine in patients with rheumatoid arthritis. Lancet Rheumatol. 3, e470–e472 (2021).
Geisen, U. M. et al. Immunogenicity and safety of anti-SARS-CoV-2 mRNA vaccines in patients with chronic inflammatory conditions and immunosuppressive therapy in a monocentric cohort. Ann. Rheum. Dis. 80, 1306–1311 (2021).
Iancovici, L. et al. Rheumatoid arthritis patients treated with Janus kinase inhibitors show reduced humoral immune responses following BNT162b2 vaccination. Rheumatology (Oxford) https://doi.org/10.1093/rheumatology/keab879 (2021).
Jena, A. et al. Response to SARS-CoV-2 vaccination in immune mediated inflammatory diseases: Systematic review and meta-analysis. Autoimmun. Rev. https://doi.org/10.1016/j.autrev.2021.102927 (2021).
Deepak, P. et al. Effect of immunosuppression on the immunogenicity of mRNA vaccines to SARS-CoV-2: a prospective cohort study. Ann. Intern. Med. 174, 1572–1585 (2021).
Ammitzboll, C. et al. Impaired antibody response to the BNT162b2 messenger RNA coronavirus disease 2019 vaccine in patients with systemic lupus erythematosus and rheumatoid arthritis. ACR Open Rheumatol. 3, 622–628 (2021).
Haberman, R. H. et al. Methotrexate hampers immunogenicity to BNT162b2 mRNA COVID-19 vaccine in immune-mediated inflammatory disease. Ann. Rheum. Dis. 80, 1339–1344 (2021).
Naranbhai, V. et al. Immunogenicity and reactogenicity of SARS-CoV-2 vaccines in patients With cancer: The CANVAX Cohort Study. J. Clin. Oncol. https://doi.org/10.1200/JCO.21.01891 (2021).
Linardou, H. et al. Responses to SARS-CoV-2 vaccination in patients with cancer (ReCOVer study): a prospective cohort study of the Hellenic Cooperative Oncology Group. Cancers (Basel) https://doi.org/10.3390/cancers13184621 (2021).
Figueiredo, J. C. et al. Longitudinal SARS-CoV-2 mRNA vaccine-induced humoral immune responses in cancer patients. Cancer Res. https://doi.org/10.1158/0008-5472.CAN-21-3554 (2021).
Corti, C. et al. Seroconversion rate after vaccination against COVID-19 in cancer patients—a systematic review. Ann. Oncol. https://doi.org/10.1016/j.annonc.2021.10.014 (2021).
Addeo, A. et al. Immunogenicity of SARS-CoV-2 messenger RNA vaccines in patients with cancer. Cancer Cell 39, 1091–1098 (2021).
Thakkar, A. et al. Seroconversion rates following COVID-19 vaccination among patients with cancer. Cancer Cell 39, 1081–1090 (2021).
Peeters, M. et al. Reduced humoral immune response after BNT162b2 coronavirus disease 2019 messenger RNA vaccination in cancer patients under antineoplastic treatment. ESMO Open 6, 100274 (2021).
Ligumsky, H. et al. Immunogenicity and safety of the BNT162b2 mRNA COVID-19 vaccine among actively treated cancer patients. J. Natl Cancer Inst. https://doi.org/10.1093/jnci/djab174 (2021).
Agbarya, A. et al. Efficacy of the mRNA-based BNT162b2 COVID-19 vaccine in patients with solid malignancies treated with anti-neoplastic drugs. Cancers (Basel) https://doi.org/10.3390/cancers13164191 (2021).
Massarweh, A. et al. Evaluation of seropositivity following BNT162b2 messenger RNA vaccination for SARS-CoV-2 in patients undergoing treatment for cancer. JAMA Oncol. 7, 1133–1140 (2021).
Schramm, R. et al. Poor humoral and T cell response to two-dose SARS-CoV-2 messenger RNA vaccine BNT162b2 in cardiothoracic transplant recipients. Clin. Res. Cardiol. 110, 1142–1149 (2021).
Holden, I. K. et al. Immunogenicity of SARS-CoV-2 mRNA vaccine in solid organ transplant recipients. J. Intern. Med. 290, 1264–1267 (2021).
Mazzola, A. et al. Poor antibody response after two doses of SARS-CoV-2 vaccine in transplant recipients. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciab580 (2021).
Havlin, J. et al. Immunogenicity of BNT162b2 mRNA COVID-19 vaccine and SARS-CoV-2 infection in lung transplant recipients. J. Heart Lung Transplant. 40, 754–758 (2021).
Sattler, A. et al. Impaired humoral and cellular immunity after SARS-CoV-2 BNT162b2 (tozinameran) prime-boost vaccination in kidney transplant recipients. J. Clin. Invest. https://doi.org/10.1172/JCI150175 (2021).
Cucchiari, D. et al. Cellular and humoral response after mRNA-1273 SARS-CoV-2 vaccine in kidney transplant recipients. Am. J. Transplant. 21, 2727–2739 (2021).
Boyarsky, B. J. et al. Antibody response to two-dose SARS-CoV-2 mRNA vaccine series in solid organ transplant recipients. JAMA 325, 2204–2206 (2021).
Marinaki, S. et al. Immunogenicity of SARS-CoV-2 BNT162b2 vaccine in solid organ transplant recipients. Am. J. Transplant 21, 2913–2915 (2021).
Chavarot, N. et al. Poor anti-SARS-CoV-2 humoral and T cell responses after 2 injections of mRNA vaccine in kdney transplant recipients treated with belatacept. Transplantation 105, e94–e95 (2021).
Mannick, J. B. et al. TORC1 inhibition enhances immune function and reduces infections in the elderly. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.aaq1564 (2018).
Diaz, A. et al. Metformin improves in vivo and in vitro B cell function in individuals with obesity and type 2 diabetes. Vaccine 35, 2694–2700 (2017).
Mannick, J. B. et al. mTOR inhibition improves immune function in the elderly. Sci. Transl. Med. 6, 268ra179 (2014).
Jagannath, C. & Bakhru, P. Rapamycin-induced enhancement of vaccine efficacy in mice. Methods Mol. Biol. 821, 295–303 (2012).
Pearce, E. L. et al. Enhancing CD8 T cell memory by modulating fatty acid metabolism. Nature 460, 103–107 (2009).
Yang, C. & Malarkannan, S. Transcriptional regulation of NK cell development by mTOR complexes. Front. Cell Dev. Biol. 8, 566090 (2020).
Xiao, Y. et al. Metformin promotes innate immunity through a conserved PMK-1/p38 MAPK pathway. Virulence 11, 39–48 (2020).
Stathopoulou, C., Nikoleri, D. & Bertsias, G. Immunometabolism: an overview and therapeutic prospects in autoimmune diseases. Immunotherapy 11, 813–829 (2019).
Boor, P. P., Metselaar, H. J., Mancham, S., van der Laan, L. J. & Kwekkeboom, J. Rapamycin has suppressive and stimulatory effects on human plasmacytoid dendritic cell functions. Clin. Exp. Immunol. 174, 389–401 (2013).
Saemann, M. D., Haidinger, M., Hecking, M., Horl, W. H. & Weichhart, T. The multifunctional role of mTOR in innate immunity: implications for transplant immunity. Am. J. Transplant. 9, 2655–2661 (2009).
Kennedy, R. B., Ovsyannikova, I. G., Lambert, N. D., Haralambieva, I. H. & Poland, G. A. The personal touch: strategies toward personalized vaccines and predicting immune responses to them. Expert Rev. Vaccines 13, 657–669 (2014).
Zimmermann, P. & Curtis, N. The influence of probiotics on vaccine responses—a systematic review. Vaccine 36, 207–213 (2018).
Peroni, D. G. & Morelli, L. Probiotics as adjuvants in vaccine strategy: is there more room for improvement? Vaccines (Basel) https://doi.org/10.3390/vaccines9080811 (2021).
Acknowledgements
We acknowledge S. Shukla and S. Bal for contributions to the generation of figures 1-3 created with BioRender.com. This project was supported for R.P.S and J.A.T by NCI/NHI U54CA260563 and NIAID/NIH U19AI128910. This project has been funded for M.S.S. in part with federal funds from the National Institute of Allergy and Infectious Diseases (NIAID; R56AI147623, U19AI057266, U19AI090023 and 1U54CA260563 to Emory University) National Institutes of Health (NIH), Department of Health and Human Services (HHSN272201400004C and 75N93021C00017; NIAID Centers of Excellence for Influenza Research and Surveillance) and NIH P51 OD011132 to Emory University. This work was also supported in part by the Emory Executive Vice President for Health Affairs Synergy Fund award (75D30121C10084), COVID-Catalyst-I3 funds from the Woodruff Health Sciences Center and Emory School of Medicine, the Pediatric Research Alliance Center for Childhood Infections and Vaccines and Children’s Healthcare of Atlanta, and Woodruff Health Sciences Center 2020 COVID-19 CURE award.
Author information
Authors and Affiliations
Contributions
J.A.T. conceptually developed and wrote the article. M.S.S. and S.G.D. contributed to writing and editing. R.P.S. conceptually developed, wrote and edited the article.
Corresponding author
Ethics declarations
Competing interests
M.S.S. serves on the advisory board for Moderna. The other authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Katie Flanagan and Sabra Klein for their contribution to the peer review of this work. Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tomalka, J.A., Suthar, M.S., Deeks, S.G. et al. Fighting the SARS-CoV-2 pandemic requires a global approach to understanding the heterogeneity of vaccine responses. Nat Immunol 23, 360–370 (2022). https://doi.org/10.1038/s41590-022-01130-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-022-01130-4
This article is cited by
-
Human memory T cell dynamics after aluminum-adjuvanted inactivated whole-virion SARS-CoV-2 vaccination
Scientific Reports (2023)
-
Binding and inactivation of human coronaviruses, including SARS-CoV-2, onto purified clinoptilolite-tuff
Scientific Reports (2023)
-
Endothelium dysfunction and thrombosis in COVID-19 with type 2 diabetes
Endocrine (2023)
-
Immune response in COVID-19: what is next?
Cell Death & Differentiation (2022)