Abstract
Multimodal T cell profiling can enable more precise characterization of elusive cell states underlying disease. Here, we integrated single-cell RNA and surface protein data from 500,089 memory T cells to define 31 cell states from 259 individuals in a Peruvian tuberculosis (TB) progression cohort. At immune steady state >4 years after infection and disease resolution, we found that, after accounting for significant effects of age, sex, season and genetic ancestry on T cell composition, a polyfunctional type 17 helper T (TH17) cell–like effector state was reduced in abundance and function in individuals who previously progressed from Mycobacterium tuberculosis (M.tb) infection to active TB disease. These cells are capable of responding to M.tb peptides. Deconvoluting this state—uniquely identifiable with multimodal analysis—from public data demonstrated that its depletion may precede and persist beyond active disease. Our study demonstrates the power of integrative multimodal single-cell profiling to define cell states relevant to disease and other traits.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Single-cell sequencing data are available on dbGaP; count matrices and metadata are available on the GEO (GSE158769). Single-cell expression and cluster abundance data can also be explored in an interactive browser at https://immunogenomics.io/tbru/. Genotype data are available on dbGaP under accession code phs002025.v1.p1.
Code availability
Custom analysis scripts are available on GitHub (https://github.com/immunogenomics/TB_Tcell_CITEseq).
References
Nathan, A., Baglaenko, Y., Fonseka, C. Y., Beynor, J. I. & Raychaudhuri, S. Multimodal single-cell approaches shed light on T cell heterogeneity. Curr. Opin. Immunol. 61, 17–25 (2019).
Spitzer, M. H. & Nolan, G. P. Mass cytometry: single cells, many features. Cell 165, 780–791 (2016).
Peterson, V. M. et al. Multiplexed quantification of proteins and transcripts in single cells. Nat. Biotechnol. 35, 936–939 (2017).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Carr, E. J. et al. The cellular composition of the human immune system is shaped by age and cohabitation. Nat. Immunol. 17, 461–468 (2016).
Brodin, P. et al. Variation in the human immune system is largely driven by non-heritable influences. Cell 160, 37–47 (2015).
Houben, R. M. & Dodd, P. J. The global burden of latent tuberculosis infection: a reestimation using mathematical modelling. PLoS Med. 13, e1002152 (2016).
World Health Organization. Global Tuberculosis Report 2020. (WHO, 2020); https://www.who.int/tb/publications/global_report/en/
Verver, S. et al. Rate of reinfection tuberculosis after successful treatment is higher than rate of new tuberculosis. Am. J. Respir. Crit. Care Med. 171, 1430–1435 (2005).
Day, C. L. et al. PD-1 expression on Mycobacterium tuberculosis-specific CD4 T cells is associated with bacterial load in human tuberculosis. Front. Immunol. 9, 1995 (2018).
Adekambi, T. et al. Distinct effector memory CD4+ T cell signatures in latent Mycobacterium tuberculosis infection, BCG vaccination and clinically resolved tuberculosis. PLoS ONE 7, e36046 (2012).
Arrigucci, R. et al. Active tuberculosis is characterized by highly differentiated effector memory TH1 cells. Front. Immunol. 9, 2127 (2018).
Lindestam Arlehamn, C. S. et al. Memory T cells in latent Mycobacterium tuberculosis infection are directed against three antigenic islands and largely contained in a CXCR3+CCR6+ TH1 subset. PLoS Pathog. 9, e1003130 (2013).
Scriba, T. J. et al. Distinct, specific IL-17- and IL-22-producing CD4+ T cell subsets contribute to the human anti-mycobacterial immune response. J. Immunol. 180, 1961–1970 (2008).
Coulter, F. et al. IL-17 production from T helper 17, mucosal-associated invariant T, and γδ cells in tuberculosis infection and disease. Front. Immunol. 8, 1252 (2017).
Burel, J. G. et al. Transcriptomic analysis of CD4+ T cells reveals novel immune signatures of latent tuberculosis. J. Immunol. 200, 3283–3290 (2018).
Perreau, M. et al. Lack of Mycobacterium tuberculosis–specific interleukin-17A-producing CD4+ T cells in active disease. Eur. J. Immunol. 43, 939–948 (2013).
Becerra, M. C. et al. Transmissibility and potential for disease progression of drug resistant Mycobacterium tuberculosis: prospective cohort study. BMJ 367, l5894 (2019).
Zumla, A., Raviglione, M., Hafner, C. & von Reyn, C. F. Tuberculosis. N. Engl. J. Med. 368, 745–755 (2013).
Fontenot, J. D., Gavin, M. A. & Rudensky, A. Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 4, 330–336 (2003).
Liu, W. et al. CD127 expression inversely correlates with FoxP3 and suppressive function of human CD4+ Treg cells. J. Exp. Med. 203, 1701–1711 (2006).
Gutierrez-Arcelus, M. et al. Lymphocyte innateness defined by transcriptional states reflects a balance between proliferation and effector functions. Nat. Commun. 10, 687 (2019).
Korsunsky, I. et al. Fast, sensitive, and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Savage, A. K. et al. The transcription factor PLZF directs the effector program of the NKT cell lineage. Immunity 29, 391–403 (2008).
Suliman, S. et al. Peripheral blood mucosal-associated invariant T cells in tuberculosis patients and healthy Mycobacterium tuberculosis-exposed controls. J. Infect. Dis. 222, 995–1007 (2020).
Sallusto, F., Lenig, D., Förster, R., Lipp, M. & Lanzavecchia, A. Two subsets of memory T lymphocytes with distinct homing potentials and effector functions. Nature 401, 708–712 (1999).
Joeckel, L. T. & Bird, P. I. Are all granzymes cytotoxic in vivo? Biol. Chem. 395, 181–202 (2014).
Maecker, H. T., McCoy, J. P. & Nussenblatt, R. Standardizing immunophenotyping for the Human Immunology Project. Nat. Rev. Immunol. 12, 191–200 (2012).
Acosta-Rodriguez, E. V. et al. Surface phenotype and antigenic specificity of human interleukin 17–producing T helper memory cells. Nat. Immunol. 8, 639–646 (2007).
Szabo, S. J. et al. A novel transcription factor, T-bet, directs TH1 lineage commitment. Cell 100, 665–669 (2000).
Sallusto, F., Zielinski, C. E. & Lanzavecchia, A. Human TH17 subsets. Eur. J. Immunol. 42, 2215–2220 (2012).
Wambre, E. et al. A phenotypically and functionally distinct human TH2 cell subpopulation is associated with allergic disorders. Sci. Transl. Med. 9, eaam9171 (2017).
Kleinewietfeld, M. et al. CCR6 expression defines regulatory effector/memory-like cells within the CD25+CD4+ T cell subset. Blood 105, 2877–2886 (2005).
Lee, J. et al. Enrichment of human CCR6+ regulatory T cells with superior suppressive activity in oral cancer. J. Immunol. 199, 467–476 (2017).
Fonseka, C. Y. et al. Mixed-effects association of single cells identifies an expanded effector CD4+ T cell subset in rheumatoid arthritis. Sci. Transl. Med. 10, eaaq0305 (2018).
Kaczorowski, K. J. et al. Continuous immunotypes describe human immune variation and predict diverse responses. Proc. Natl Acad. Sci. USA 114, E6097–E6106 (2017).
Hashimoto, K. et al. Single-cell transcriptomics reveals expansion of cytotoxic CD4 T cells in supercentenarians. Proc. Natl Acad. Sci. USA 116, 24242–24251 (2019).
Amadori, A. et al. Genetic control of the CD4/CD8 T cell ratio in humans. Nat. Med. 1, 1279–1283 (1995).
Kissick, H. T. et al. Androgens alter T cell immunity by inhibiting T-helper 1 differentiation. Proc. Natl Acad. Sci. USA 111, 9887–9892 (2014).
Ter Horst, R. et al. Host and environmental factors influencing individual human cytokine responses. Cell 167, 1111–1124 (2016).
Nedelec, Y. et al. Genetic ancestry and natural selection drive population differences in immune responses to pathogens. Cell 167, 657–669 (2016).
Berry, M. P. et al. An interferon-inducible neutrophil-driven blood transcriptional signature in human tuberculosis. Nature 466, 973–977 (2010).
Scriba, T. J. et al. Sequential inflammatory processes define human progression from M. tuberculosis infection to tuberculosis disease. PLoS Pathog. 13, e1006687 (2017).
Fritsch, R. D. et al. Stepwise differentiation of CD4 memory T cells defined by expression of CCR7 and CD27. J. Immunol. 175, 6489–6497 (2005).
Coquet, J. M. et al. The CD27 and CD70 costimulatory pathway inhibits effector function of T helper 17 cells and attenuates associated autoimmunity. Immunity 38, 53–65 (2013).
MA State Public Health Laboratory Tuberculosis Report. (Massachusetts Department of Public Health, 2020).
Arlehamn, C. S., McKinney, D. M. & Carpenter, C. A quantitative analysis of complexity of human pathogen-specific CD4 T cell responses in healthy M. tuberculosis-infected South Africans. PLoS Pathog. 12, e1005760 (2016).
Christophersen, A. et al. Distinct phenotype of CD4+ T cells driving celiac disease identified in multiple autoimmune conditions. Nat. Med. 25, 734–737 (2019).
Zhang, F. et al. Defining inflammatory cell states in rheumatoid arthritis joint synovial tissues by integrating single-cell transcriptomics and mass cytometry. Nat. Immunol. 20, 928–942 (2019).
Smillie, C. S. et al. Intra- and inter-cellular rewiring of the human colon during ulcerative colitis. Cell 178, 714–730 (2019).
Boisson-Dupuis, S. et al. Inherited and acquired immunodeficiencies underlying tuberculosis in childhood. Immunol. Rev. 264, 103–120 (2015).
Pai, M. et al. Tuberculosis. Nat. Rev. Dis. Primers 2, 16076 (2016).
Bustamante, J., Boisson-Dupuis, S., Abel, L. & Casanova, J. L. Mendelian susceptibility to mycobacterial disease: genetic, immunological and clinical features of inborn errors of IFN-γ immunity. Semin. Immunol. 26, 454–470 (2014).
Okada, S. et al. Impairment of immunity to Candida and Mycobacterium in humans with bi-allelic RORC mutations. Science 349, 606–613 (2015).
Luo, Y. et al. Early progression to active tuberculosis is a highly heritable trait driven by 3q23 in Peruvians. Nat. Commun. 10, 3765 (2019).
Cosmi, L. et al. Human interleukin 17–producing cells originate from a CD161+CD4+ T cell precursor. J. Exp. Med. 205, 1903–1916 (2008).
Bengsch, B. et al. Human TH17 cells express high levels of enzymatically active dipeptidylpeptidase IV (CD26). J. Immunol. 188, 5438–5447 (2012).
Morimoto, C. & Schlossman, S. F. The structure and function of CD26 in the T cell immune response. Immunol. Rev. 161, 55–70 (1998).
Fergusson, J. R. et al. CD161 defines a transcriptional and functional phenotype across distinct human T cell lineages. Cell Rep. 9, 1075–1088 (2014).
Yamazaki, T. et al. CCR6 regulates the migration of inflammatory and regulatory T cells. J. Immunol. 181, 8391–8401 (2008).
Scriba, T. J. et al. Differential recognition of Mycobacterium tuberculosis-specific epitopes as a function of tuberculosis disease history. Am. J. Respir. Crit. Care Med. 196, 772–781 (2017).
Chowdhury, R. et al. A multi-cohort study of the immune factors associated with M. tuberculosis infection outcomes. Nature 560, 644–648 (2018).
Wang, M. et al. Genetic polymorphisms of IL-17A, IL-17F, TLR4 and miR-146a in association with the risk of pulmonary tuberculosis. Sci. Rep. 6, 28586 (2016).
Khader, S. A. et al. IL-23 and IL-17 in the establishment of protective pulmonary CD4+ T cell responses after vaccination and during Mycobacterium tuberculosis challenge. Nat. Immunol. 8, 369–377 (2007).
Arlehamn, C. L. et al. Transcriptional profile of tuberculosis antigen-specific T cells reveals novel multifunctional features. J. Immunol. 193, 2931–2940 (2014).
Shanmugasundaram, U. et al. Pulmonary Mycobacterium tuberculosis control associates with CXCR3- and CCR6-expressing antigen-specific Th1 and Th17 cell recruitment. JCI Insight 5, 137858 (2020).
Darrah, P. A. et al. Prevention of tuberculosis in macaques after intravenous BCG immunization. Nature 577, 95–102 (2020).
Lindenstrom, T. et al. Vaccine-induced TH17 cells are maintained long-term postvaccination as a distinct and phenotypically stable memory subset. Infect. Immun. 80, 3533–3544 (2012).
Fares, A. Seasonality of tuberculosis. J. Glob. Infect. Dis. 3, 46–55 (2011).
Moody, D. B. et al. CD1b-mediated T cell recognition of a glycolipid antigen generated from mycobacterial lipid and host carbohydrate during infection. J. Exp. Med. 192, 965–976 (2000).
Thornton, T. et al. Estimating kinship in admixed populations. Am. J. Hum. Genet. 91, 122–138 (2012).
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Reich, D. et al. Reconstructing Native American population history. Nature 488, 370–374 (2012).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
O’Connell, J. et al. A general approach for haplotype phasing across the full spectrum of relatedness. PLoS Genet. 10, e1004234 (2014).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Kang, H. M. et al. Multiplexed droplet single-cell RNA-sequencing using natural genetic variation. Nat. Biotechnol. 36, 89–94 (2018).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
McInnes, L. & Healy, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at arXiv https://arxiv.org/abs/1802.03426 (2018).
Gonzalez, I., Déjean, S., Martin, P. G. P. & Baccini, A. CCA: an R package to extend canonical correlation analysis. J. Stat. Softw. https://doi.org/10.18637/jss.v023.i12 (2007).
Dobin, A., Davis, C. A. & Schlesinger, F. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Friedman, J. H., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. https://doi.org/10.18637/jss.v033.i01 (2010).
Kuznetsova, A., Brockhoff, P. B. & Christensen, R. H. B. lmerTest Package: tests in linear mixed-effects models. J. Stat. Softw. https://doi.org/10.18637/jss.v082.i13 (2017).
Acknowledgements
We thank K. Slowikowski for building an interactive data browser. This work is supported in part by funding from the National Institutes of Health (U19 AI111224 to S.R., M.B.M. and D.B.M.; UH2AR067677 to S.R.; T32 HG002295 to A.N.; T32 AR007530 to A.N.; U01 HG009379 to S.R.; U19 AI111211 to J.D.E.; and R01AI049313 to D.B.M.).
Author information
Authors and Affiliations
Contributions
S.R., D.B.M. and M.B.M. conceptualized and designed the study. A.N. and S.R. designed the statistical and computational strategy and analyzed the data. K.I., S.A. and Y.L. conducted additional statistical analyses. J.I.B., Y.B., S.S., C.S.L.A., J.D.E., A.N., I.V.R., D.B.M. and S.R. designed the immunoprofiling strategy. J.I.B., Y.B. and S.S. conducted all immunoprofiling experiments. J.J., L.L. and M.B.M. recruited, clinically phenotyped and obtained blood samples from human participants. I.V.R., K.L. and R.C. organized processing, transportation and management of PBMCs. C.-C.H., Z.Z. and M.B.M. curated and analyzed clinical phenotype data. A.N. and S.R. wrote the initial manuscript. All authors contributed to the writing and editing of the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Immunology thanks Thomas Scriba and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CITE-seq cell and feature quality.
a, In silico memory T cell gating. Each cell is plotted based on its normalized surface expression of CD3 and CD45RO, measured through CITE-seq. Gates are demarcated with red dashed lines. Red cells were removed. Counts represent the number of cells in each quadrant. b, UMAP representation of gated cells. Red cells were gated out in (a) and cells clustering with them in the UMAP (shown in black in (a) and (b)) were also removed. c, Normalized CD3 and CD45RO surface protein expression. d, Number of cells per sample after QC, stratified by TB progression status. P value is from a two-sided t test comparing mean cell counts in each group. e, Pearson correlation coefficient (r) was calculated between normalized mRNA and surface protein expression for each marker across cells passing QC. r is plotted against average normalized mRNA expression for each protein. f, Each cell is plotted based on normalized expression of each marker in surface protein and mRNA, both measured through CITE-seq, with density contours. We fit a best-fit line (in blue) with a linear model.
Extended Data Fig. 2 Comparing proportions of eight major T cell states between flow cytometry and CITE-seq.
a, Average percents per population in CITE-seq vs. flow cytometry. Gates in (b). Dashed line indicates the identity line. b, For each population, proportions plotted across 259 donors. Flow cytometry gating occurred after gating T cells. CITE-seq gating occurred after isolation of memory T cells. The dashed line indicates the identity line, and we calculated Pearson correlation coefficients (r) for each state.
Extended Data Fig. 3 Multimodal integration with canonical correlation analysis.
a, Correlations for the top 20 canonical dimensions used in downstream analysis. Bars represent the Pearson correlation between mRNA and protein projections for each dimension. b, Marker correlation with canonical variates (CVs). Each marker is plotted based on its mRNA and protein correlation with CV1 (left) or CV2 (right). c, Innateness scores. UMAP is colored based on a gene expression-derived cytotoxicity score defined in Gutierrez-Arcelus, et al. d, Correlation between innateness score and CV1. Each cell is plotted based on its innateness score from (c) and its CV1 projection, and we report the Pearson correlation coefficient.
Extended Data Fig. 4 Single-cell expression of surface proteins, measured with CITE-seq.
Each cell is colored according to its expression of each protein and plotted in UMAP space. Colors are scaled independently for each marker from minimum (blue) to maximum (yellow) expression.
Extended Data Fig. 5 Technical replicate consistency.
For each of the 31 multimodal clusters, we plotted its proportion in replicate 1 and in replicate 2 of each donor. We calculated the Pearson correlation coefficients (r) for each cluster.
Extended Data Fig. 6 Effects of donor covariates on memory T cell states.
Effects of age, sex, winter blood draw and proportion of European genetic ancestry in (a) model correcting for technical covariates (# UMIs/cell, % MT UMIs/cell), donor, batch and TB disease status, and (b) full model with TB disease status, age, sex, winter blood draw, proportion of European genetic ancestry, technical covariates, donor and batch. For all, n = 271 samples from 259 independent donors. For each cluster, data are presented as the MASC OR of a cell being in each cluster given the contrast covariate (95% CI error bars) and the -log(LRT p value) of the association. The dashed horizontal line corresponds to a Bonferroni p-value threshold of 0.05/31. Labeled clusters are significant at this threshold.
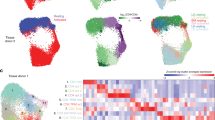
Extended Data Fig. 7 Unimodal clusters and associations with TB disease progression.
a–c, mRNA clusters. d–f, protein clusters. a and d, UMAPs colored by unimodal clusters. Clusters boxed in red are CD4+, purple are mixed CD4+ and CD8+, blue are CD8+, and green are CD4-CD8-. b and e, Expression of major lineage-defining surface markers measured through CITE-seq. The UMAPs are colored by the expression of five markers measured through CITE-seq. Colors are scaled independently for each marker from minimum (blue) to maximum (yellow) expression. c and f, Heatmap of overlap between unimodal and multimodal clusters. Colors indicate the proportion of the multimodal cluster (column) overlapping with the unimodal cluster (row). g, Associations between TB disease status and unimodal protein clusters. For each cluster, the data are presented as MASC ORs of a cell being in each cluster for cases vs. controls (95% CI error bars), and the -log(LRT p value) of the association (n = 271 samples from 259 independent donors). The dashed horizontal line corresponds to a Bonferroni p-value threshold of 0.05/40. Labeled clusters are significant at this threshold. h, Abundance of C-20 in 128 cases and 131 controls. P value is from an LRT with 1 d.f. Boxplots show the median (vertical bar), 25th and 75th percentiles (lower and upper bounds of box, respectively), and 1.5xIQR (or minimum/maximum if they lie within that range; end of whiskers).
Extended Data Fig. 8 Cell state signature extrapolation in public data.
a, Correlation between actual and predicted C-12 proportion, per sample in memory T cell CITE-seq study (n = 271, black) and bulk PBMC RNA-seq (n = 15, blue). Line represents the identity line and we calculated the Pearson correlation coefficients (r) for T cell samples and bulk PBMC samples separately. Predicted (b) C-12 or (c) C-11 (TH2, negative control) proportion in 3 categories of donors from Berry, et al. d, Predicted C-12 proportion in active donors at 3 time points during anti-mycobacterial treatment (0, 2, 12 months) and uninfected controls. In b–d, we calculated p values with a two-sided t test. e, Histogram of two-sided t test p values from 1,000 trials of downsampling LIMAA cohort to 7 cases and 12 controls (as in Berry, et al.) and comparing the average C-12 proportion in cases vs controls. Dashed line is the significance threshold of p = 0.05 (power = 0.15). f, Predicted C-12 proportion in active cases and latent controls at 2 pre-disease-progression time points in Scriba, et al. g, Predicted C-12 proportion in active cases and latent controls in pre- and post-disease cohorts. Pre-disease data are aggregated across 2 time points. In f and g, p-values are from a one-sided t-test (Satterthwaite’s d.f. method) of the beta estimate for TB progression status in the linear mixed model. h, Concordance of Pearson correlations between each cluster’s proportion and the C-12 score or the C-12 cluster’s proportion. Pearson correlation coefficients were computed for each cluster across 271 memory T cell samples. Each point represents one of the 31 clusters, and the dashed line is the identity line. All boxplots show the median (vertical bar), 25th and 75th percentiles (lower and upper bounds of box, respectively), and 1.5xIQR (or minimum/maximum if they lie within that range; end of whiskers).
Extended Data Fig. 9 Cytokine production in Boston donors.
Bars represent the mean and error bars show standard error of the mean across 5 Boston donors unascertained for TB. a, Per-donor percent of cells producing each cytokine in gated populations. b, Per-donor percent of total cytokine-producing memory CD4 + T cells in each gated population.
Extended Data Fig. 10 M.tb antigen-specific response in CD4+CD26+CD161+CCR6+ memory T cells.
a, Biaxial plots showing representative gating of CD3+CD4+CD45RO+CD161+CCR6+CD26+ cells in a Boston donor. b, Intracellular staining for IL-17A and IFNγ in a Boston control donor and two Peruvian TB cohort donors after either no peptide stimulation (control) or stimulation with the MTB300 megapool. c, IL-17A or IFNγ response to MTB300 stimulation in all CD3 + T cells from either Boston control donors (n = 2) or Peruvian TB cohort donors (n = 6). Each point corresponds to the percent of cells producing IL-17A or IFNγ from one donor, measured with intracellular cytokine staining. Lines connect measurements from the same donor before and after stimulation with MTB300 peptide megapool. Boxplots show the median (vertical bar), 25th and 75th percentiles (lower and upper bounds of box, respectively), and minimum/maximum (end of whiskers). P values are from a two-sided Wilcoxon signed-rank test comparing donors before and after antigen stimulation.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3.
Supplementary Tables
Supplementary Tables 1–10
Supplementary Data
Weights for each gene in linear mixed model estimating C-12 proportion, trained on T cell and PBMC data.
Rights and permissions
About this article
Cite this article
Nathan, A., Beynor, J.I., Baglaenko, Y. et al. Multimodally profiling memory T cells from a tuberculosis cohort identifies cell state associations with demographics, environment and disease. Nat Immunol 22, 781–793 (2021). https://doi.org/10.1038/s41590-021-00933-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-021-00933-1
This article is cited by
-
The genetic basis of autoimmunity seen through the lens of T cell functional traits
Nature Communications (2024)
-
Multimodal and spatially resolved profiling identifies distinct patterns of T cell infiltration in nodal B cell lymphoma entities
Nature Cell Biology (2024)
-
Intricacies of TGF-β signaling in Treg and Th17 cell biology
Cellular & Molecular Immunology (2023)
-
Th17.1 cell driven sarcoidosis-like inflammation after anti-BCMA CAR T cells in multiple myeloma
Leukemia (2023)
-
A statistical framework for differential pseudotime analysis with multiple single-cell RNA-seq samples
Nature Communications (2023)