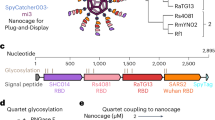
Abstract
The SARS-CoV-2 virus emerged in December 2019 and has caused a worldwide pandemic due to the lack of any pre-existing immunity. Accurate serology testing is urgently needed to help diagnose infection, determine past exposure of populations and assess the response to a future vaccine. The landscape of antibody responses to SARS-CoV-2 is unknown. In this study, we utilized the luciferase immunoprecipitation system to assess the antibody responses to 15 different SARS-CoV-2 antigens in patients with COVID-19. We identified new targets of the immune response to SARS-CoV-2 and show that nucleocapsid, open reading frame (ORF)8 and ORF3b elicit the strongest specific antibody responses. ORF8 and ORF3b antibodies, taken together as a cluster of points, identified 96.5% of COVID-19 samples at early and late time points of disease with 99.5% specificity. Our findings could be used to develop second-generation diagnostic tests to improve serological assays for COVID-19 and are important in understanding pathogenicity.
Similar content being viewed by others
Main
Serological testing is important for the diagnosis and sero-epidemiology of SARS-CoV-2 virus infection and vaccines. Several serology tests are currently in use that primarily assess Spike (S) and on occasion, nucleocapsid (N) antibodies1,2. Seroconversion to S and N generally occurs in the second or third week of illness and is not suitable for diagnosis of acute disease and these assays have suboptimal sensitivity and specificity. Moreover, most vaccines in development against COVID-19 target the S protein to elicit neutralizing antibodies to block infection3, as the S protein contains the receptor binding domain, which is critical for viral entry4. However, it is unclear whether neutralizing antibodies to S protein are the major contributor to a protective immune response5. SARS-CoV-2 has at least four structural proteins: S, envelope (E), membrane (M) and N. In addition, the SARS-CoV-2 genome encodes for 23 putative nonstructural proteins6. ORF1ab encodes for a large polyprotein that is proteolytically cleaved into 16 nonstructural proteins (NSP1–16). ORF3a, ORF3b, ORF6, ORF7a, ORF7b, ORF8 and ORF10 may encode for proteins but their functions are yet to be determined.
Encouragingly, most patients with SARS-CoV-2 develop robust T and B cellular responses, along with neutralizing and high titer S1-specific antibody responses, which has formed the basis of serological diagnosis and vaccine development7,8,9. In our previous study we found that neutralizing antibodies eventually develop by 28 d after infection10; however, it is reported by some studies that a proportion of patients with confirmed COVID-19, determined by PCR with reverse transcription (RT–PCR), do not develop robust antibody responses5. Therefore the possibility of low or no antibody responses by traditional serological approaches may lead to an underestimation of asymptomatic and mild infection and threaten the success of a potential vaccine that targets S alone. Therefore, a broader landscape of antibody responses to a range of viral proteins needs to be assessed to better detect the immunogenicity of SARS-CoV-2 infection and understand pathogenesis and immunity.
Accurate SARS-CoV-2 serology is currently a global challenge and the antibody landscape is yet to be determined8,11,12. The luciferase immunoprecipitation system (LIPS) assay allows for a comprehensive understanding of the antibody immune response by the expression of any protein or antigen as a recombinant Renilla luciferase (Ruc)-antigen fusion13. These (Ruc)-antigen fusions are then pulled down with immune serum. This is an advantage when commercial recombinant proteins are limited, especially in the early days of a pandemic.
In this study, we used LIPS to initially assess the antibody responses to a panel of 15 SARS-CoV-2 antigens, representing the structural and nonstructural viral proteins in 15 patients with COVID-19 and 15 pre-pandemic negative controls. Antibodies to the four structural proteins (S, N, M and E), three S subunits (S1, S2 and S2′), the seven available ORFs (ORF3a, ORF3b, ORF6, ORF7a, ORF7b, ORF8 and ORF10) and one relevant NSP within ORF1ab (NSP1) were tested14. A cutoff calculated as the mean of the results on the negative plasma samples plus 3 × s.d. allowed selection of sensitive and specific tests15,16,17. We then further validated our selected assays on a larger panel of sera totaling 84 plasma samples from patients with COVID-19 from early (n = 51, <14 d post-symptom onset) and late time points (n = 33, >14 d post-symptom onset) and 176 negative pre-pandemic controls to assess assay performances of relevant N, ORF3b and ORF8 antigenic targets. Finally, sequential samples from patients were tested to determine the kinetics of the antibody response.
Results
Antibody responses to structural proteins in patients with COVID-19
First, the antibody responses to the four structural proteins of SARS-CoV-2 were assessed by LIPS (Supplementary Table 1). We detected significantly higher antibody responses to the full S, N and M proteins in patients with COVID-19 (Supplementary Table 2) compared to negative controls (P < 0.0001, P = 0.023 and P = 0.0116 respectively; Fig. 1a–c), whereas E antibodies were comparable to negative controls (P = 0.0591; Fig. 1d).
a–d, Antibodies against the four SARS-CoV-2 structural proteins S (a), N (b), M (c) and E (d) were measured by LIPS from patients with COVID-19 and age-matched negative controls. e, Antibodies against the S subunits S1, S2 and S2′ by LIPS from patients with COVID-19 and negative controls. The cutoff value is shown by the dotted line and was based on the mean plus 3 × s.d. of the negative control group. Blue dots represent data points of patients with COVID-19 before day 14. f, Full S, S1, S2 and S2′ antibody LIPS responses in patients with COVID-19 with low MN titers (<160) versus high MN titers (≥160). g, Full S antibody LIPS responses in patients with COVID-19 with low ELISA S IgG responses (<1) versus high ELISA IgG responses (≥1). All data represent mean ± s.d. and individual responses (n = 15). Experiments were repeated twice. Two-sided P values were calculated using the Mann–Whitney U-test. Background values were subtracted. Asterisks indicates statistical significance between patients with COVID-19 versus negative controls or between patients grouped by MN > 160 versus MN < 160 or ELISA optical density (OD) > 1 versus OD < 1. NS, P = 0.0591, *P < 0.05, **P < 0.01.
The S protein has multiple stages of maturation, which may present unique antigens. The trimer S protein is cleaved into S1, containing the receptor binding domain and S2 subunits6,18 and S2 is further cleaved into S2′ to form the viral fusion peptide19. The S1 subunit showed significantly higher antibodies in patients with COVID-19 than controls (5,191 ± 1,469 luminometer units (LU) versus 4,003 ± 1,062 LU, P = 0.0288; Fig. 1e). There was no difference in the levels of antibodies to S2 in the LIPS assay between patients with COVID-19 and negative controls (P = 0.5683), whereas antibodies to the S2′ cleaved subunit were significantly higher in patients with COVID-19 (P = 0.0391 for S2′; Fig. 1e). We detected significant differences in the magnitude of responses by LIPS between patients and controls for S, S1 and S2′, but these antigens show very low sensitivity levels (33%, 13.3% and 0% respectively; Supplementary Table 3), especially for sera collected early (<14 d) after disease onset (indicated by blue circles in Fig. 1a,e).
The S protein also elicits non-neutralizing antibodies targeted to conserved epitopes8, and among our cohort an absence of in vitro neutralization has been observed for some patients in the first 2 weeks after onset of illness (Supplementary Table 2). Patients with higher SARS-CoV-2 micro-neutralization (MN) titers (>160 reciprocal serum dilution) also had higher antibody responses toward full S by LIPS, but not for S1, S2 and S2′ subunits (Fig. 1f, P = 0.0049). Traditional full-S ELISA correlated with LIPS LU responses (R2 = 0.5289) (Fig. 1g and Extended Data Fig. 1a,b). Patients with COVID-19 also had significantly higher N-specific IgG by ELISA (Extended Data Fig. 1c); however, N-specific IgG by ELISA did not correlate strongly with LIPS results (Extended Data Fig. 1d). Pre-adsorption of patient plasma with the commercial N protein led to a significant reduction of LIPS signal by 88% indicating that the N protein and our R-Luc N antigen both detect N-specific antibodies (Extended Data Fig. 1e).
Antibody responses of patients with COVID-19 are directed toward unique ORFs
We next investigated the presence of antibodies specific to previously uncharacterized ORFs to SARS-CoV-2. We used LIPS to detect antibodies specific to NSP1, ORF3a, ORF3b, ORF6, ORF7a, ORF7b, ORF8 and ORF10 (Fig. 2). We expressed only the NSP1 as a representative antigen of ORF1ab due to its extended length (>21 kb)20. We detected significantly higher antibody levels in the 15 patients with COVID-19 toward NSP1, ORF3a, ORF3b, ORF7a, ORF7b and ORF8 (P < 0.0001, P = 0.0302, P < 0.0001, P < 0.0001, P = 0.0019 and P < 0.0001, respectively; Fig. 2a–g). The largest difference in mean antibody signals among the ORFs between COVID-19 samples and negative controls was observed for ORF3b (7,712 ± 2,947 LU versus 3,599 ± 1,029 LU) and ORF8 (16,933 ± 7,489 LU versus 5,440 ± 1,096 LU) (Fig. 2c,g). There was no significant difference between patients with COVID-19 and negative controls for ORF6 and ORF10 antigens (Fig. 2d,h). Furthermore, there was no difference in the magnitude of responses between males and females or between patients aged below and above 60 years of age (P = 0.6289, P = 0.9363; Extended Data Fig. 2a,b).
a–h, Antibodies against NSP1 (a) (in ORF1ab) and other ORFs (ORF3a (b), ORF3b (c), ORF6 (d), ORF7a (e), ORF7b (f), ORF8 (g) and ORF10 (h)) were measured by LIPS to cover all the ORFs of the virus. i–l, Cumulative antibody LIPS levels to the SARS-CoV-2 antigens in patients with COVID-19 and healthy controls for 11 relevant antigens (N, M, S, S1, S2′, NSP1, ORF3a, ORF3b, ORF7a, ORF7b and ORF10) (i), three most sensitive antigens (N, ORF3b and ORF8) (j), selective antigens (ORF3b and ORF8) (k) and three S-relevant antigens (S, S1 and S2′) (l). The cutoff value is shown by the dotted line and was based on the mean + (3 × s.d.) of the negative control group. Blue dots represents data points of patients with COVID-19 before day 14. Experiments were repeated twice. Two-sided P values were calculated using the Mann–Whitney U-test. Asterisks denote statistical significance between patients with COVID-19 versus negative controls. *P < 0.05, **P < 0.01, ***P < 0.005, ****P < 0.0001. All data represent the mean ± s.d. and individual responses (n = 15); background values were subtracted.
The ability of each antigenic target to correctly identify patients with COVID-19 was determined using cutoffs calculated as the mean of the negative population plus 3 × s.d. (dotted line on graphs and values given in Supplementary Table 3)15,16,17. The cutoff values of the 11 relevant tests (excluding the nonsignificant E, S2, ORF6 and ORF10 tests) allowed the calculation of assay performances (specificity, sensitivity, positive predictive value (PPV) and negative predictive value (NPV) (Supplementary Table 3). All 11 antigens (S, S1, S2′, N, M, NSP1, ORF3a, ORF3b, ORF7a, ORF7b and ORF8) showed high specificity (absence of false positives in the negative population). On the other hand, eight antigens (M, S, S1, S2′, NSP1, ORF3a, ORF7a and ORF7b) showed a low (<75%) sensitivity (false negatives of patient samples), hence being insufficient to correctly identify all the patients with COVID-19 and limiting their use for SARS-CoV-2 diagnostics (Supplementary Table 3). Notably, only the N, ORF3b and ORF8 antigens showed high sensitivity levels of 93.3%, 86.6% and 100%, respectively (Supplementary Table 3).
Combined antigen test panels as a potential diagnostic tool for COVID-19
As many SARS-CoV-2 LIPS tests showed low sensitivity, we then used an approach based on combining LIPS antibody LU signals for the 11 separate SARS-CoV-2 antigen tests15,16,17. This approach of combining the LU values as a sum from the 11 LIPS tests efficiently detected 100% of the 15 patients with COVID-19 even at early time points (Fig. 2i; blue dots for ≤14 d). Further, combination of only three LIPS tests N + ORF3b + ORF8 antibodies also had a 100% sensitivity and specificity (Fig. 2j and Supplementary Table 3). As ORF3b and ORF8 showed the lowest homology to previous SARS-CoV among all the viral proteins6 we also looked separately at the combination of responses toward ORF3b and ORF8 (Fig. 2k). Again, the ORF3b + ORF8 sum showed all patients with COVID-19 had a total score above the cutoff and all negative controls had a sum below, with all early time points being correctly detected (Fig. 2k). Alternatively, the combination of the three relevant S protein LIPS tests (S, S1 and S2′), decreased the sensitivity of the LIPS test to 26.6% as only four of the patients with COVID-19 had a total combined LU above the cutoff (Fig. 2l), with all samples from early time points (day 4 to day 14) under the cutoff value (blue dots, Fig. 2l). Hence, early time points samples were mis-identified (below the cutoff) using both full S and S sums (Figs. 1a and 2l), but were correctly identified as positive using both the N + ORF3b + ORF8 and the ORF3b + ORF8 sums (Fig. 2j,k). Therefore, the combinational use of ORF3b and ORF8 tests alone could be sufficient to detect individuals exposed to COVID-19 at any time point of infection.
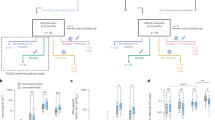
Validation of LIPS in a second larger cohort and two-dimensional plane for diagnosis by ORF3b and ORF8
To validate these findings, we further increased our sample size to 84 COVID-19 plasmas and 176 negatives for N (Fig. 3a), ORF3b (Fig. 3b) and ORF8 (Fig. 3c) antibody responses. We observed high specificity (>95%) for all three tests as above (Fig. 3a–c,g) and N (83.3% sensitivity) and ORF8 (84.5% sensitivity) showed the highest sensitivity (Fig. 3g), while the sum of N + ORF3b + 8 increased the sensitivity to 87.5% (Fig. 3a–e,g). Representation of the ORF3b and ORF8 antibody values in a two-dimensional plane (x, y) allowed a clearer visualization and definition of the COVID-19 (red) and negative cohorts (gray) (Fig. 3f). Indeed, by introducing an optimal discriminant line (the line parallel to the vector (0.2185; 0.5927) in the plane), we could decipher clearly positive and negative populations. Only 1 patient out of 176 negative controls was an outlier and all other negative patients were clearly identified as negative with a 99.5% specificity. Meanwhile, the sensitivity was also increased to 96.5% for the cluster analysis versus 81% for the sum of ORF3b + ORF8 (Fig. 3f,g). This geometric interpretation allows a highly accurate diagnosis of COVID-19 and emphasizes the diagnostic relevance of these two antibody responses, ORF3b and ORF8 to be considered together.
a–c, Antibodies against the three relevant SARS-CoV-2 antigens N (a), ORF3b (b) and ORF8 (c), were measured by LIPS from COVID-19 samples (n = 84) and negative controls (n = 176). d,e, Sum of antibody LIPS levels to N + ORF3b + ORF8 (d) and ORF3b + ORF8 (e) SARS-CoV-2 antigens in patients with COVID-19 and negative controls. Data represents the mean ± s.d. and individual responses. The cut-off value is shown by the dotted line and was based on the mean + 3 × s.d. of the negative control. Background values were subtracted. Experiments were repeated twice. f, Cluster of points representing each patient with COVID-19 in red and negative controls in gray in the plane (ORF3b, ORF8). The equation of the line is 0.2187 x + 0.5927 y = 4,643.19. g, Sensitivity and specificity performances of the N, ORF3b, ORF8 and their sums (from a–e) and cluster analysis (from f). The contour red line shows the highest sensitivity and specificity. Two-sided P values were calculated using the Mann–Whitney U-test. Asterisks denote statistical significance between patients with COVID-19 versus negative controls. ****P < 0.0001.
Early diagnostic use of ORF3b + ORF8 specific antibodies by LIPS
One of the main challenges of the serologic diagnosis of SARS-CoV-2 is identification of infection at early time points. The assessment of the N, ORF3b and ORF8 tests from day 0 to day 14 samples revealed that 25.5%, 37.3% and 19.6% of the samples, respectively were mis-identified (below the cutoff; Fig. 4a and Extended Data Fig. 3). However, using the ORF3b + ORF8 sum improved the diagnosis of these early-time-point samples with a sensitivity of 86.4%, allowing the correct detection of 44 out of 51 samples within 14 d of symptom onset (Fig. 4a–c).
a–c, Sensitivity performances of N, ORF3b, ORF8 and their sums for early time points before day 14. The contour red line shows the highest sensitivity and specificity (a). Detection of early-time-point patients only (≤day 14) with the sum of three most sensitive antigens (N + ORF3b + ORF8) (b) and ORF3b + ORF8 (c). d, Longitudinal time point patient (n = 14) responses for N (red), ORF3b (pink), ORF8 (green) and S (blue). The x axis represents time point (to scale) after onset of symptoms in days for each patient. On the y axis, each unit represents 103 for ORF3b, ORF8 and S and 106 for N (origin shifted for the sake of clarity). e, Linear regression of full S LU versus ORF8 LU of patients with COVID-19 (from a) (R2 = 0.66902, P < 0.0001, two-sided P value). f, Fold changes from earliest available acute time point sample (from a) to convalescence (14–30 d post-symptom onset) and long-term memory (>31 d post-symptom onset) of S, N, ORF8 and ORF3b LU. *P < 0.05 shows statistical significance by one-sample Student’s t-test versus a hypothetical value of one fold change. g, Percentages of OC43, 229E and NL63 S IgG in the negative cohort. ELISAs were performed on n = 176 COVID-19 negative control cohort samples. Positivity in HCoV ELISA was defined as OD > 1 and negativity as OD < 1.
Characterization of longitudinal antibody responses of S, N, ORF3b and ORF8
Longitudinal studies enable the assessment of kinetics of antibody responses, especially antibody waning. Longitudinal and paired samples from two to four time points were collected from 14 patients for longitudinal measurement of S, N, ORF3b and ORF8 antibody responses. The antibody responses to N, S, ORF3b and ORF8 are well maintained over time (even up to day 100) (Fig. 4d). Of note, among the four antibodies tested, ORF8 and S antibodies followed a very similar trend in all patients (Fig. 4d, blue (S) and green lines (ORF8)). Indeed, the kinetics of the ORF8- and S-specific antibody responses by LIPS significantly correlated (R2 = 0.66902, P < 0.0001) (Fig. 4e), whereas the other antibody responses did not.
The fold changes (Fig. 4f) of antibody levels from acute (<14 d after symptom onset) to convalescent (day 14–30) and long-term memory (day > 31) responses were determined from longitudinal samples (Fig. 4d). The N response is significantly increased at long-term memory (P = 0.0359), whereas the S, ORF3b and ORF8 all show a similar trend of maintained responses with a fold change close to one (Fig. 4f). Of note, ORF8 and ORF3b responses are the most stable across patients (with the narrowest s.d.) over time, making them ideal markers of acute and past infection.
Lack of cross-reactivity of existing human coronavirus responses for N, ORF3b and ORF8 by LIPS
Owing to the high prevalence of human coronavirus (HCoV) in the community, we assessed the positive responses to β-HCoV (OC43) and α-HCoV (229E and NL63) S antibodies in our negative cohort. We found that 89.6% of our negative cohort was positive for OC43 S in ELISA (OD > 1.00; Fig. 4g), while these samples are true negatives in our SARS-CoV-2 LIPS assays with comparable responses to OC43 responders and nonresponders (Figs. 3a–c and 4g). The same trend was observed for α-HCoV-229E and NL63 with a lower prevalence (21% and 19% S IgG positive respectively; Fig. 4g). Similarly, the common β-HCoV HKU1 was found to be highly prevalent in the general adult population (98% positive for HKU1 S IgG) in previous studies9. When we separated our negative cohort (n = 176) into donors with either high or low S OC43, 229E or NL63 IgG responses, we found no differences between these groups for N, ORF3b and ORF8 responses by LIPS (Extended Data Fig. 4). Moreover, the structural proteins of SARS-CoV-2 and other common HCoV only share 18 to 40% amino acid homology (Supplementary Table 4). These results clearly demonstrate the high specificity of our N, ORF3b and ORF8 LIPS tests despite the high prevalence of endemic HCoVs in most samples.
Immune dominance of N-specific antibodies in patients with COVID-19
A comparison of the global SARS-CoV-2 antibody responses from the initial cohort of 15 patients with COVID-19 revealed that anti-N antibodies dominate the humoral response detected by LIPS (Fig. 5a). To further investigate specificity of responses versus negative controls, we subtracted the mean LU levels of negative controls from the mean LU of patients infected with COVID-19 for each test and compared this difference across each antigen (Fig. 5b). We found that this magnitude of the difference was significantly greater for the N-specific antibody response compared to all the other relevant antibody responses (M, S, S1, S2′, NSP1, ORF3a, ORF3b, ORF7a, ORF7b and ORF8) (P < 0.0001 by one-way analysis of variance (ANOVA); Fig. 5b), highlighting a possible dominance and specificity of N across the SARS-CoV-2 humoral immune responses. Besides N, ORF8 and ORF3b also seem to be important antigenic targets with significantly higher responses above the cutoff (Fig. 3b,c). Multiple comparison analysis revealed that ORF8 was significantly increased compared to all other antigens (P < 0.0001 versus ten remaining antigens: M, S, S1, S2′, NSP1, ORF3a, ORF3b, ORF7a and ORF7b; Fig. 5b,c). Meanwhile results for ORF3b were significant against the remaining eight antigens (M, S, S1, S2′, NSP1, ORF3a, ORF3b and ORF7b), excluding ORF7a (Fig. 5c). NSP1 and ORF7a also seem to be interesting targets, though little is known about their function. Of note, ORF7a was the fourth immunogenic target after ORF3b (Fig. 5c) and responses against additional NSPs within ORF1ab remain to be determined.
a, Global individual immune responses detected by LIPS in the 15 patients with COVID-19 for 11 relevant antigens. b,c, Differences of antibody response means between COVID-19 and negative control populations for 11 relevant antigens (b) and as a pie chart excluding N (c). Data for b,c are derived from Figs. 1 and 2. Each symbol represents the mean ± s.d. of the differences between COVID-19 (n = 15) and negative groups for the 11 relevant antibody levels in LU. Data shown in b were analyzed by one-way ANOVA with Tukey’s multiple comparisons test. Owing to the differences in scale between N and all other ten antigens, a second ordinary one-way ANOVA with Tukey’s multiple comparisons test was performed excluding N to determine the dominance among the remaining ten antigens. ****P < 0.0001 versus all other antigens unless otherwise indicated. Background values were subtracted. Experiments were repeated twice.
Discussion
SARS-CoV-2 antibody testing is a major component for diagnosis of recent and past COVID-19 infection21. Antibody tests are important for determining infection attack rates in the population, population immunity and to inform vaccine development. We report the detection of antibody responses directed against an extensive spectrum of 15 SARS-CoV-2 antigens to identify new and unique antigenic targets of the humoral immune response of patients with COVID-19. Using LIPS technology with crude lysates from transfected cells, we screened all structural proteins along with all ORFs of the SARS-CoV-2 virus (NSP1 only for ORF1ab). In our panel, 11 antigens (the structural proteins full S, S1, S2′, N and M and the ORFs: NSP1, ORF3a, ORF3b, ORF7a, ORF7b and ORF10) showed elevated antibody responses in patients with COVID-19. Among them, the S structural protein is responsible for viral entry and is widely used as a diagnostic marker of infection. In the S subunits S1, S2 and S2′, only antibodies to S1 and S2′ were elevated in patients with COVID-19 by our LIPS test. The trimer S conformation and maturation of viral particles by the cleavage of S2 during virus endocytosis to form the S2′ fusion peptide may explain the difference in antigenicity between S2 and S2′. Patients with higher MN titers had correspondingly higher levels of full-S LIPS and ELISA results, establishing consistency among assays for the full-S protein. While we detected significant differences in the magnitude of responses by LIPS between patients and controls for S, S1 and S2′, these antigens did not show high sensitivity levels, especially for plasma collected early post-disease onset, making these results insufficient for early diagnostic use. As expected, N showed elevated antibody responses, better performing as a diagnostic tool than the other structural proteins22.
We next cloned all the available ORFs of the virus ORF1ab (as NSP1 only), ORF3a, ORF3b, ORF6, ORF7a, ORF7b, ORF8 and ORF10, to acquire more extensive information of the unique immunogenic targets of the virus beyond known structural proteins. We found that although NSP1, ORF3a, ORF7a and ORF7b elicited high antibody levels in the COVID-19 cohort, their performance was insufficient for use in diagnostics, unlike ORF3b and ORF8.
Single-antibody LIPS tests resulted in a high proportion of false negatives at early time points of infection (day 0 to day 14), but our data reveal that the combinational use of ORF3b, ORF8 and N is a high-performing marker of infection at early and late time points. Therefore, the combination of multiple antigens beyond S could be the basis for supplementary serological tests useful to determine SARS-CoV-2 exposure to overcome false-negative results.
NSP1 and ORF7a antibodies could be of interest in understanding the physiopathology of COVID-19. Bioinformatic predictions showed that they could be involved in suppressing the antiviral host response6,19. On the other hand, E, S2, ORF6 and ORF10 antibodies did not show elevated levels in patients with COVID-19 in our assay, consistent with the findings of Wang et al. by microarray23.
ORF3b and ORF8 are the least identical proteins to SARS-CoV6 and homologous proteins do not exist in other strains of HCoV other than sarbecoviruses. However, very little is known about their function and expression in SARS-CoV-2. Previous reports found the ORF3b of SARS-CoV plays an important role in the interaction with the innate immune system through inhibition of type 1 interferon synthesis24. In SARS-CoV, ORF8 has been shown to accumulate in the endoplasmic reticulum and mediate cell death by autophagy25. Previously, ORF8 was reported to have a strong association with the S protein in SARS-CoV, while also inhibiting expression of E protein26, which may account for the correlation of the antibody response kinetics observed for these two proteins in our longitudinal samples. Notably, recent findings report that SARS-CoV-2 utilizes ORF8 to alter the expression of major histocompatibility complex I to evade immune surveillance27. A deletion of ORF8 has been reported in a few patients with COVID-19 from Singapore28; however, this lineage has not continued or been maintained in Singapore and other countries29.
Our study includes 37 longitudinal samples from 14 patients, which are necessary to assess kinetics of antibody responses. Testing these longitudinal samples allowed us to determine that ORF3b and ORF8 antibodies are stable over time, highlighting their importance for early rapid diagnosis of patients with COVID-19. The levels of these antibodies were stable up to day 100 post-symptom onset and further studies on a potential waning at later time points are currently under investigation in our group.
Endemic HCoVs are ubiquitous and sequence homology with other human coronaviruses, such as α-HCoV strains: 229E and NL63 and β-HCoV: HKU1 and OC43, could result in the detection of pre-existing cross-reactive antibodies, reducing the specificity of serological assays. We showed herein, that despite the high sero-positivity rate for OC43, 229E and NL63 in our negative control cohort, there is an absence of cross-reactivity with our N, ORF8 and ORF3b antibody LIPS tests, which is crucial for their wide use for diagnosis. Our results need to be confirmed in cohorts from different continents to account for antibody responses across different ethnicities and viral strains. The LIPS platform allowed broad antibody screening to many antigens, but a translation of the assay into a simpler setup such as ELISA is required for the use in large-scale diagnostics, particularly in resource-poor settings.
In conclusion, our results provide insights into the overall spectrum of antibody responses associated with COVID-19, as patients produce antibodies to structural and nonstructural proteins beyond S. There is a need for improvement of current sero-diagnostic tests for detecting infection early after onset of symptoms and for confirmatory assays when existing S and N protein-specific antibodies by ELISA are positive. We have identified ORF8 antibodies as a major marker of acute, convalescent and long-term antibody response to SARS-CoV-2 infection because of its immunodominance and specificity, in addition to N protein. Moreover, the combined use of ORF3b and ORF8 provides a highly sensitive and specific method for the detection of patients with COVID-19, both early and later in infection. Despite the high prevalence of common-cold HCoVs, we have demonstrated the specificity of our assay to SARS-CoV-2. Further investigation on the protective potential of antibodies to these nonstructural targets are needed. Such information will help prioritize antigen targets for vaccine development, monoclonal antibody reagents and most importantly, detection of early responses to infection by standardized immunoassays.
Methods
Patients and samples collection
Our study enrolled a total of 61 adult patients based on recruitment of available patients with RT–PCR-confirmed COVID-19 infection in Hong Kong and used 84 plasma samples from these 61 patients with COVID-19, including 14 individuals with 2–4 sampling time points and 51 early-time-point samples (≤day 14). Sample day was defined as day post-symptom onset. The COVID-19 patient study was approved by the Institutional Review Board of the Hong Kong West Cluster of the Hospital Authority of Hong Kong (approval number: UW20-169) and all patients provided informed consent.
To first determine the LIPS assay for patients and controls (Figs. 1 and 2), an initial cohort of 15 patients with COVID-19 (minimum age of 37 years old, maximum age of 73 years old, mean of 58.6 years old) (Supplementary Table 2) was tested versus a set of 15 plasma samples from age- and sex-matched negative healthy controls collected before the COVID-19 pandemic (minimum age of 37 years old, maximum age of 70 years old, mean of 50.5 years old). To increase sample size and validate findings on a second set of samples (Figs. 3 and 4), a larger panel of 176 additional negative control plasma samples from healthy subjects (minimum age of 16 years old, maximum age of 65 years old) also collected before the COVID-19 pandemic were tested. The negative control plasma was from Hong Kong blood donors collected from June to August 2017 (before the emergence of COVID-19) and was recently used as a control cohort for neutralization assays10. The collection of negative control blood donors was approved by the Institutional Review Board of The Hong Kong University and the Hong Kong Island West Cluster of Hospitals (reference no. UW16-254). Plasma samples were collected from heparinized blood and heat-inactivated before experimental use at 56 °C for 30 min.
SARS-CoV-2 gene cloning
On the basis of previous studies describing the structure of the SARS-CoV-2 genome6,20, an extensive panel of 12 proteins (S, E, M, N, NSP1, ORF3a, ORF3b, ORF6, ORF7a, ORF7b, ORF8 and ORF10) was chosen for antibody testing by LIPS (Supplementary Table 1). Primers for the amplification of SARS-CoV-2 proteins were designed (Supplementary Table 5). RT–PCR was performed using RNA extracted from SARS-CoV-2 virus strain βCoV/Hong Kong/VM20001061/2020 grown in Vero E6 cells to amplify target genes corresponding to structural and nonstructural proteins of the virus (Supplementary Table 1)19 using Platinum SuperScriptIII One Step RT–PCR system. The bands were then extracted using QIAGEN gel extraction kit (QIAGEN) and digested with BamHI and NotI or KpnI-HF and XhoI (New England Biolabs). Extracted products were ligated using T4 DNA ligase (New England Biolabs) into the pREN2 plasmid (from P. Burbelo, National Institute of Dental and Craniofacial Research, National Institutes of Health). Plasmids were transformed using DH10B competent cells and purified using PureYield Plasmid mid-prep system (Promega). Constructs were confirmed by Sanger Sequencing (3730xl DNA Analyzer Applied Biosystems).
SARS-CoV-2 (Ruc)-antigen expression
Constructs with pREN2-Renilla luciferase plasmid containing the SARS-CoV-2 antigen of interest were transfected into Cos1 cells using Fugene 6 (Promega) as per manufacturer’s instructions. Cells were collected 48 h later, lysed and sonicated and (Ruc)-antigen yields were measured using a luminometer plate reader (PerkinElmer) according to the protocol of Burbelo et al.14. Each Ruc-fusion antigen is tested for its LU yield after production in Cos1 cells. The LU was then standardized to 107 LU per antigen in each well before each LIPS assay, therefore different yields during transfection for recovery of luciferase-tagged proteins is controlled in the assay and plasma samples run against the same amount of each antigen.
Measurement of antibody responses using LIPS
The LIPS assays were performed following the protocol of Burbelo et al.14 with the following modifications. Briefly, (Ruc)-antigen (at an equal concentration for each antigen at 107 per well) and plasma (heat-inactivated and diluted 1:100) were incubated for 2 h with shaking at 800 r.p.m. Ultralink protein A/G beads were added to the (Ruc)-antigen and serum mixture in a 96-deep-well polypropylene microtiter plate and incubated for 2 h with shaking at 800 r.p.m. The entire volume was then transferred into HTS plates and washed as previously described. The plate was read using QUANTI-Luc Gold substrate (Invivogen) as per manufacturer’s instructions on a Wallac MicroBeta JET luminometer 1450 LSC & Luminescence counter and its software for analysis (PerkinElmer). Experimental controls include no-plasma blank wells with (Ruc)-antigens and negative control serum from age-matched noninfected patient plasma collected before the COVID-19 pandemic. The background corresponds to the LU signal from each Ruc-fusion antigen with protein A/G and substrate with no plasma.
ELISA
ELISA assays were performed with the available SARS-CoV-2 proteins S (S1 + S2) and N, as well as HCoV-OC43, NL63 and 229E S (SinoBiological). Briefly, recombinant S and N proteins were coated on 96‐well flat‐bottom immunosorbent plates (Nunc Immuno MaxiSorp) at a concentration of 500 ng ml−1, in 100 μl of coating buffer (PBS with 53% Na2CO3 and 42% NaHCO3, pH 9.6) at 4 °C overnight. An additional plate coated with a nonspecific protein (blocking buffer, PBS with 5% FBS) was used to measure the background binding of each plasma sample. Following FBS blocking and thorough washing, diluted plasma samples (1:100) were bound for 2 h, further washed and then detected by an anti‐human IgG secondary antibody labeled with HRP (Invitrogen) at 1:5,000 dilution in dilution buffer (1% FBS, 0.05% Tween in PBS).
N adsorption
Plasma samples (n = 3) were incubated on 5 μg ml−1 N protein coated 96-round-well plate with shaking at 800 r.p.m. at 37 °C for 2 h twice. N LIPS was then performed as described above.
Micro-neutralization assay
Plasma samples were diluted in serial twofold dilutions commencing with a dilution of 1:10 and mixed with equal volumes of SARS-CoV-2 at a dose of 200 tissue culture infective doses 50% determined by Vero E6 cells respectively. After 1 h of incubation at 37 °C, 35 µl of the virus–serum mixture was added in quadruplicate to Vero or Vero E6 cell monolayers in 96-well microtiter plates. After 1 h of adsorption, the virus–serum mixture was removed and replaced with 150 µl of virus growth medium in each well. The plates were incubated for 3 d at 37 °C in 5% CO2 in a humidified incubator. Cytopathic effect was observed at day 3 post-inoculation. The highest plasma dilution protecting 50% of the replicate wells was denoted as the neutralizing antibody titer. A virus back-titration of the input virus was included in each batch of tests.
Multiple alignments of coronaviruses
Multiple amino acid alignments of structural proteins from HKU1 (Genbank AY597011.2), HCoV-229E (Genbank AF304460.1), HCoV-OC43 (Genbank AY391777.1) and HCoV-NL63 (Genbank AY567487.2) were compared versus SARS-CoV-2 (Genbank MN908947) using CLUSTAL 2.1.
ORF3b–ORF8 cluster of points
The ORF3b and ORF8 full dataset has been treated through the free software ConTeXt, with LuaMetaTeXengine (v.2020.05.18) developed by H. Hagen (http://www.pragma-ade.nl), which uses TeX, Metapost and Lua to obtain the two-dimensional cloud shown in Fig. 3f. The red line (equation 0.2185ORF3b + 0.5927ORF8 = 4643.1972) allows the most accurate discrimination between negative controls (gray dots) and positive patient (red dots) populations.
Statistics and reproducibility
GraphPad Prism v.8 software was used for statistical analysis. Antibody levels are presented as the geometric mean ± s.d. For the calculation of sensitivity (percentage of true positives, absence of true negatives in the positive population) and specificity (percentage of true negatives, absence of false positives in the negative population), cutoff limits for each antigen were derived from the mean value plus 3 × s.d. of the negative controls. The PPV (n of true positives / (true + false positives)) and NPV (n of true negatives / (true + false negatives)) were also determined (Supplementary Table 3). Nonparametric two-sided Mann–Whitney U-tests were used to compare antibody levels between COVID-19 and negative groups. In Fig. 5b, an ordinary one-way ANOVA with Tukey’s multiple comparisons test was performed. Due to the differences in scale between N and all other ten antigens, a second ordinary one-way ANOVA with Tukey’s multiple comparisons test was performed excluding N to determine the dominance among the remaining ten antigens. All experiments were repeated twice independently.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Amino acid sequences for structural and ORF1ab protein were obtained from Genbank for HKU1 (Genbank AY597011.2), HCoV-229E (Genbank AF304460.1), HCoV-OC43 (Genbank AY391777.1) and HCoV-NL63 (Genbank AY567487.2) and SARS-CoV-2 (Genbank MN908947) viruses (Supplementary Table 4). SARS-CoV-2 ORF nucleotide sequences from Genbank and primers designed for pREN2 cloning are described in Supplementary Table 5. Raw LIPS LU and ELISA IgG responses are shown in the manuscript (Figs. 1–5 and Extended Data Figs. 1–4). The data that support the findings of this study are available from the corresponding authors upon request.
Change history
27 August 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Yong, S. E. F. et al. Connecting clusters of COVID-19: an epidemiological and serological investigation. Lancet Infect. Dis. 20, 809–815 (2020).
Weissleder, R., Lee, H., Ko, J. & Pittet, M. J. COVID-19 diagnostics in context. Sci.Transl. Med. 12, eabc1931 (2020).
Paules, C. I., Marston, H. D. & Fauci, A. S. Coronavirus infections—more than just the common cold. JAMA 323, 707–708 (2020).
Yuan, M. et al. A highly conserved cryptic epitope in the receptor-binding domains of SARS-CoV-2 and SARS-CoV. Science 368, 630–633 (2020).
Wu, F. et al. Neutralizing antibody responses to SARS-CoV-2 in a COVID-19 recovered patient cohort and their implications. Preprint at medRxiv https://doi.org/10.1101/2020.03.30.20047365 (2020).
Chan, J. F. et al. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg. Microbes Infect. 9, 221–236 (2020).
Thevarajan, I. et al. Breadth of concomitant immune responses prior to patient recovery: a case report of non-severe COVID-19. Nat. Med. 26, 453–455 (2020).
Lv, H. et al. Cross-reactive antibody response between SARS-CoV-2 and SARS-CoV infections. Cell Rep. 31, 107725 (2020).
Juno, J. A. et al. Humoral and circulating follicular helper T cell responses in recovered patients with COVID-19. Preprint at Nat. Med. https://doi.org/10.1038/s41591-020-0995-0 (2020).
Perera, R. A. et al. Serological assays for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), March 2020. Euro Surveill. https://doi.org/10.2807/1560-7917.ES.2020.25.16.2000421 (2020).
Zhang, W. et al. Molecular and serological investigation of 2019-nCoV infected patients: implication of multiple shedding routes. Emerg. Microbes Infect. 9, 386–389 (2020).
Okba, N. M. A. et al. Severe acute respiratory syndrome coronavirus 2−specific antibody responses in coronavirus disease patients. Emerg. Infect. Dis. https://doi.org/10.3201/eid2607.200841 (2020).
Burbelo, P. D., Goldman, R. & Mattson, T. L. A simplified immunoprecipitation method for quantitatively measuring antibody responses in clinical sera samples by using mammalian-produced Renilla luciferase-antigen fusion proteins. BMC Biotechnol. 5, 22 (2005).
Burbelo, P. D., Ching, K. C., Klimavicz, C. M. & Iadarola, M. J. Antibody profiling by luciferase immunoprecipitation systems (LIPS). J. Vis. Exp. 7, 1549 (2009).
Burbelo, P. D. et al. Serological diagnosis of pulmonary Mycobacterium tuberculosis infection by LIPS using a multiple antigen mixture. BMC Microbiol. 15, 205 (2015).
Frey, A., Di Canzio, J. & Zurakowski, D. A statistically defined endpoint titer determination method for immunoassays. J. Immunol. Methods 221, 35–41 (1998).
Burbelo, P. D., Chaturvedi, A., Notkins, A. L. & Gunti, S. Luciferase-based detection of antibodies for the diagnosis of HPV-associated head and neck squamous cell carcinoma. Diagnostics (Basel) 9, 89 (2019).
Walls, A. C. et al. Structure, function, and antigenicity of the SARS-CoV-2 Spike glycoprotein. Cell 181, 281–292 (2020).
Zhang, C. et al. Protein structure and sequence reanalysis of 2019-nCoV genome refutes snakes as its intermediate host and the unique similarity between its spike protein insertions and HIV-1. J. Proteome Res. 19, 1351–1360 (2020).
Wu, F. et al. A new coronavirus associated with human respiratory disease in China. Nature 579, 265–269 (2020).
Wang, W. et al. Detection of SARS-CoV-2 in different types of clinical specimens. JAMA 323, 1843–1844 (2020).
Zeng, W. et al. Biochemical characterization of SARS-CoV-2 nucleocapsid protein. Biochem. Biophys. Res. Commun. 527, 618–623 (2020).
Hongye Wang, X. H. et al. SARS-CoV-2 proteome microarray for mapping COVID-19 antibody interactions at amino acid resolution. Preprint at bioRxiv https://doi.org/10.1101/2020.03.26.994756 (2020).
Shi, C. S., Nabar, N. R., Huang, N. N. & Kehrl, J. H. SARS-coronavirus open reading frame-8b triggers intracellular stress pathways and activates NLRP3 inflammasomes. Cell Death Discov. 5, 101 (2019).
McBride, R. & Fielding, B. C. The role of severe acute respiratory syndrome (SARS)-coronavirus accessory proteins in virus pathogenesis. Viruses 4, 2902–2923 (2012).
Keng, C. T. et al. The human severe acute respiratory syndrome coronavirus (SARS-CoV) 8b protein is distinct from its counterpart in animal SARS-CoV and down-regulates the expression of the envelope protein in infected cells. Virology 354, 132–142 (2006).
Zhang, Y. et al. The ORF8 protein of SARS-CoV-2 mediates immune evasion through potently downregulating MHC-I. Preprint at bioRXiv https://doi.org/10.1101/2020.05.24.111823 (2020).
Su, Y. C. F. et al. Discovery and genomic characterization of a 382-nucleotide deletion in ORF7b and ORF8 during the early evolution of SARS-CoV-2. Mbio https://doi.org/10.1128/mBio.01610-20 (2020).
GISAID 2008–2020. EpiCoV. https://www.epicov.org/epi3/frontend#190d47 (2020).
Acknowledgements
This study was partly supported by the Research Grants Scheme (GRF 17113718) and Health and Medical Research Fund (HMRF COVID190115). We thank L.-F. Wang (Duke NUS, Singapore) for technical advice for establishing the LIPS assay. We thank J.T. Wu and K. Leung for providing access to the blood donor plasma as controls. The authors thank O. Kavian and H. Hagen for their help with the ConTeXt software used in Figs. 3f and 4d. This project was supported by the Hong Kong Health and Medical Research Fund (COVID190115, from S.A.V.) and General Research Fund (17113718, from S.A.V.). The study of the patients with COVID-19 was approved by the Institutional Review Board of the Hong Kong West Cluster of the Hospital Authority of Hong Kong (approval no. UW20-169). The collection of plasma from blood donors serving as controls was approved by the Institutional Review Board of The Hong Kong University and the Hong Kong Island West Cluster (reference no. UW16-254).
Author information
Authors and Affiliations
Contributions
A.H., N.K. and C.A.C. performed experiments. A.W.H.C., D.K.W.C., C.K.P.M., O.T.Y.T., Y.C.Y. and R.A.P.M.P. supplied specialist reagents and samples. A.H., N.K., L.L.M.P., J.S.M.P. and S.A.V. designed the study, interpreted results and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.H., N.K., L.L.M.P., J.S.M.P. and S.A.V. have filed a patent application (US 63/016,898) on the basis of the invention described in this study.
Additional information
Peer review information Peer reviewer reports are available. Jamie D. K. Wilson was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 S and N IgG responses by ELISA and LIPS.
(a) full S IgG ELISA for COVID-19 patients and healthy controls. (b) Pearson Spearman correlation of full S ELISA and LIPS of COVID-19 patients (R2=0.5289,P=0.021). (c) N IgG ELISA for COVID-19 patients and healthy controls. (d) Pearson Spearman correlation of N ELISA and LIPS of COVID-19 patients (R2=0.01579, P=0.6554). Data represents the mean ± stdev, and individual responses (n=15), background values were subtracted. (e) N pre- and post-adsorption LIPS Antibody results of COVID-19 patients (n=3). Experiments were repeated twice. Two-sided P values were calculated using the Mann-Whitney U test. * shows statistical significance between COVID-19 patients versus healthy controls. *P<0.05, **P<0.01, ***P<0.01.
Extended Data Fig. 2 SARS-CoV-2 antibody levels by LIPS according to gender and age.
Antibodies against SARS-CoV-2 antigens were measured by LIPS in COVID-19 patients and stratified by gender (a) and age (<60, and >60 years of age) (b). Data represents the mean ± stdev, and individual COVID-19 responses (n=15 COVID-19 patients from the initial cohort), background values were subtracted. Experiments were repeated twice. Two-sided P values were calculated using the Mann-Whitney U test. * shows statistical significance between COVID-19 patients versus healthy controls. *P<0.05, **P<0.01, ***P<0.005.
Extended Data Fig. 3 Detection of early time-points patients only (≤ day 14) for N, ORF3b and ORF8.
LIPS results from with N (a), ORF3b (b), ORF8 LIPS tests (c) and N+ORF8 (d). Data represents individual responses (n=51). The cutoff value of the sums is shown by the dotted line and was based on the mean plus three std-dev of the negative control group. Experiments were repeated twice.
Extended Data Fig. 4 Alpha-and Beta-HCoV positive samples do not cross-react with N, ORF3b and ORF8 SARS-CoV-2 LIPS assays.
OC43, 229E, and NL63 Spike ELISAs were performed on n=176 COVID-19 negative control cohort samples (Negative pre-pandemic controls used in this study). Graphs (a–c) represent the LIPS results for SARS-CoV-2 N, ORF3b and ORF8 according to the positivity/negativity for OC43 Spike by ELISA in the COVID-19 negative sample cohort. Graphs (d–f) represent the LIPS results for N, ORF3b and ORF8 according positivity/negativity for 229E Spike by ELISA. Graphs (g–i) represent the LIPS results for SARS-CoV-2 N, ORF3b and ORF8 according to the positivity/negativity for NL63 Spike by ELISA in the COVID-19 negative sample cohort. The dotted-line represents the cut-offs used in the main figures of the paper (Fig. 3). Two-sided P values were calculated using the Mann-Whitney U test. Ns means no significant difference.
Supplementary information
Rights and permissions
About this article
Cite this article
Hachim, A., Kavian, N., Cohen, C.A. et al. ORF8 and ORF3b antibodies are accurate serological markers of early and late SARS-CoV-2 infection. Nat Immunol 21, 1293–1301 (2020). https://doi.org/10.1038/s41590-020-0773-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0773-7
This article is cited by
-
Positive selection underlies repeated knockout of ORF8 in SARS-CoV-2 evolution
Nature Communications (2024)
-
Humoral immunity for durable control of SARS-CoV-2 and its variants
Inflammation and Regeneration (2023)
-
An interferon-integrated mucosal vaccine provides pan-sarbecovirus protection in small animal models
Nature Communications (2023)
-
The molecular mechanism of SARS-CoV-2 evading host antiviral innate immunity
Virology Journal (2022)
-
SARS-CoV-2 Delta variant isolates from vaccinated individuals
BMC Genomics (2022)