Abstract
Plasmodium parasite–specific antibodies are critical for protection against malaria, yet the development of long-lived and effective humoral immunity against Plasmodium takes many years and multiple rounds of infection and cure. Here, we report that the rapid development of short-lived plasmablasts during experimental malaria unexpectedly hindered parasite control by impeding germinal center responses. Metabolic hyperactivity of plasmablasts resulted in nutrient deprivation of the germinal center reaction, limiting the generation of memory B cell and long-lived plasma cell responses. Therapeutic administration of a single amino acid to experimentally infected mice was sufficient to overcome the metabolic constraints imposed by plasmablasts and enhanced parasite clearance and the formation of protective humoral immune memory responses. Thus, our studies not only challenge the current model describing the role and function of blood-stage Plasmodium-induced plasmablasts but they also reveal new targets and strategies to improve anti-Plasmodium humoral immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
References
Nutt, S. L., Hodgkin, P. D., Tarlinton, D. M. & Corcoran, L. M. The generation of antibody-secreting plasma cells. Nat. Rev. Immunol. 15, 160–171 (2015).
Inoue, T., Moran, I., Shinnakasu, R., Phan, T. G. & Kurosaki, T. Generation of memory B cells and their reactivation. Immunol. Rev. 283, 138–149 (2018).
Martin, F., Oliver, A. M. & Kearney, J. F. Marginal zone and B1 B cells unite in the early response against T-independent blood-borne particulate antigens. Immunity 14, 617–629 (2001).
Racine, R., Chatterjee, M. & Winslow, G. M. CD11c expression identifies a population of extrafollicular antigen-specific splenic plasmablasts responsible for CD4 T-independent antibody responses during intracellular bacterial infection. J. Immunol. 181, 1375–1385 (2008).
Slifka, M. K., Antia, R., Whitmire, J. K. & Ahmed, R. Humoral immunity due to long-lived plasma cells. Immunity 8, 363–372 (1998).
World Malaria Report 2018 (World Health Organization, 2018); https://www.who.int/malaria/publications/world-malaria-report-2018/en/
Cohen, S., McGregor, I. A. & Carrington, S. Gamma-globulin and acquired immunity to human malaria. Nature 192, 733–737 (1961).
Changrob, S. et al. Persistence of long-lived memory B cells specific to Duffy binding protein in individuals exposed to Plasmodium vivax. Sci. Rep. 8, 8347 (2018).
Longley, R. J. et al. Naturally acquired antibody responses to more than 300 Plasmodium vivax proteins in three geographic regions. PLoS Negl. Trop. Dis. 11, e0005888 (2017).
Ndungu, F. M. et al. Long-lived Plasmodium falciparum specific memory B cells in naturally exposed Swedish travelers. Eur. J. Immunol. 43, 2919–2929 (2013).
Crompton, P. D. et al. A prospective analysis of the Ab response to Plasmodium falciparum before and after a malaria season by protein microarray. Proc. Natl Acad. Sci. USA 107, 6958–6963 (2010).
Tran, T. M. et al. An intensive longitudinal cohort study of Malian children and adults reveals no evidence of acquired immunity to Plasmodium falciparum infection. Clin. Infect. Dis. 57, 40–47 (2013).
Obeng-Adjei, N. et al. Circulating Th1-cell-type Tfh cells that exhibit impaired B cell help are preferentially activated during acute malaria in children. Cell Rep. 13, 425–439 (2015).
Kurup, S. P. et al. Regulatory T cells impede acute and long-term immunity to blood-stage malaria through CTLA-4. Nat. Med. 23, 1220–1225 (2017).
Walther, M. et al. Upregulation of TGF-β, FOXP3, and CD4+CD25+ regulatory T cells correlates with more rapid parasite growth in human malaria infection. Immunity 23, 287–296 (2005).
Weiss, G. E. et al. Atypical memory B cells are greatly expanded in individuals living in a malaria-endemic area. J. Immunol. 183, 2176–2182 (2009).
Obeng-Adjei, N. et al. Malaria-induced interferon-γ drives the expansion of Tbethi atypical memory B cells. PLoS Pathog. 13, e1006576 (2017).
Portugal, S. et al. Malaria-associated atypical memory B cells exhibit markedly reduced B cell receptor signaling and effector function. Elife 4, e07218 (2015).
Sullivan, R. T. et al. FCRL5 delineates functionally impaired memory B cells associated with Plasmodium falciparum exposure. PLoS Pathog. 11, e1004894 (2015).
Ryg-Cornejo, V. et al. Severe malaria infections impair germinal center responses by inhibiting T follicular helper cell differentiation. Cell Rep. 14, 68–81 (2016).
Keitany, G. J. et al. Blood stage malaria disrupts humoral immunity to the pre-erythrocytic stage circumsporozoite protein. Cell Rep. 17, 3193–3205 (2016).
Butler, N. S. et al. Therapeutic blockade of PD-L1 and LAG-3 rapidly clears established blood-stage Plasmodium infection. Nat. Immunol. 13, 188–195 (2011).
Horne-Debets, J. M. et al. PD-1 dependent exhaustion of CD8+ T cells drives chronic malaria. Cell Rep. 5, 1204–1213 (2013).
Guthmiller, J. J., Graham, A. C., Zander, R. A., Pope, R. L. & Butler, N. S. Cutting edge: IL-10 is essential for the generation of germinal center B cell responses and anti-Plasmodium humoral immunity. J. Immunol. 198, 617–622 (2017).
Krishnamurty, A. T. et al. Somatically hypermutated plasmodium-specific IgM+ memory B cells are rapid, plastic, early responders upon malaria rechallenge. Immunity 45, 402–414 (2016).
Shaffer, A. L. et al. Blimp-1 orchestrates plasma cell differentiation by extinguishing the mature B cell gene expression program. Immunity 17, 51–62 (2002).
William, J., Euler, C. & Shlomchik, M. J. Short-lived plasmablasts dominate the early spontaneous rheumatoid factor response: differentiation pathways, hypermutating cell types, and affinity maturation outside the germinal center. J. Immunol. 174, 6879–6887 (2005).
Tellier, J. et al. Blimp-1 controls plasma cell function through the regulation of immunoglobulin secretion and the unfolded protein response. Nat. Immunol. 17, 323–330 (2016).
Meding, S. J., Cheng, S. C., Simon-Haarhaus, B. & Langhorne, J. Role of gamma interferon during infection with Plasmodium chabaudi chabaudi. Infect. Immun. 58, 3671–3678 (1990).
Su, Z. & Stevenson, M. M. Central role of endogenous gamma interferon in protective immunity against blood-stage Plasmodium chabaudi AS infection. Infect. Immun. 68, 4399–4406 (2000).
Reimold, A. M. et al. Plasma cell differentiation requires the transcription factor XBP-1. Nature 412, 300–307 (2001).
Boothby, M. & Rickert, R. C. Metabolic regulation of the immune humoral response. Immunity 46, 743–755 (2017).
Cantor, J. et al. CD98hc facilitates B cell proliferation and adaptive humoral immunity. Nat. Immunol. 10, 412–419 (2009).
Cordy, R. J. et al. Distinct amino acid and lipid perturbations characterize acute versus chronic malaria. JCI Insight 4, e125156 (2019).
Suan, D. et al. CCR6 defines memory B cell precursors in mouse and human germinal centers, revealing light-zone location and predominant low antigen affinity. Immunity 47, 1142–1153.e4 (2017).
Alugupalli, K. R. et al. B1b lymphocytes confer T cell-independent long-lasting immunity. Immunity 21, 379–390 (2004).
Haas, K. M., Poe, J. C., Steeber, D. A. & Tedder, T. F. B-1a and B-1b cells exhibit distinct developmental requirements and have unique functional roles in innate and adaptive immunity to S. pneumoniae. Immunity 23, 7–18 (2005).
Pelletier, N. et al. Plasma cells negatively regulate the follicular helper T cell program. Nat. Immunol. 11, 1110–1118 (2010).
Khan, A. R. et al. PD-L1hi B cells are critical regulators of humoral immunity. Nat. Commun. 6, 5997 (2015).
Matsumoto, M. et al. Interleukin-10-producing plasmablasts exert regulatory function in autoimmune inflammation. Immunity 41, 1040–1051 (2014).
Clark, E. H. et al. Plasmodium falciparum malaria in the Peruvian Amazon, a region of low transmission, is associated with immunologic memory. Infect. Immun. 80, 1583–1592 (2012).
Labak, C. M. et al. Glucose transport: meeting the metabolic demands of cancer, and applications in glioblastoma treatment. Am. J. Cancer Res. 6, 1599–1608 (2016).
Lam, W. Y. et al. Metabolic and transcriptional modules independently diversify plasma cell lifespan and function. Cell Rep. 24, 2479–2492.e6 (2018).
Chapuis, N., Poulain, L., Birsen, R., Tamburini, J. & Bouscary, D. Rationale for targeting deregulated metabolic pathways as a therapeutic strategy in acute myeloid leukemia. Front. Oncol. 9, 405 (2019).
Zhu, X., Pan, Y., Li, Y., Cui, L. & Cao, Y. Supplement of l-arg improves protective immunity during early-stage Plasmodium yoelii 17XL infection. Parasite Immunol. 34, 412–420 (2012).
Gordon, E. B. et al. Targeting glutamine metabolism rescues mice from late-stage cerebral malaria. Proc. Natl Acad. Sci. USA 112, 13075–13080 (2015).
Miyakoda, M. et al. Malaria-specific and nonspecific activation of CD8+ T cells during blood stage of Plasmodium berghei infection. J. Immunol. 181, 1420–1428 (2008).
McCarthy, J. S. et al. A pilot randomised trial of induced blood-stage Plasmodium falciparum infections in healthy volunteers for testing efficacy of new antimalarial drugs. PLoS ONE 6, e21914 (2011).
Rockett, R. J. et al. A real-time, quantitative PCR method using hydrolysis probes for the monitoring of Plasmodium falciparum load in experimentally infected human volunteers. Malar. J. 10, 48 (2011).
Collins, K. A. et al. A controlled human malaria infection model enabling evaluation of transmission-blocking interventions. J. Clin. Invest. 128, 1551–1562 (2018).
Montes de Oca, M. et al. Type I interferons regulate immune responses in humans with blood-stage Plasmodium falciparum infection. Cell Rep. 17, 399–412 (2016).
Malleret, B. et al. A rapid and robust tri-color flow cytometry assay for monitoring malaria parasite development. Sci. Rep. 1, 118 (2011).
Villarino, N. F. et al. Composition of the gut microbiota modulates the severity of malaria. Proc. Natl Acad. Sci. USA 113, 2235–2240 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-Seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Gray, L. R. et al. Hepatic mitochondrial pyruvate carrier 1 is required for efficient regulation of gluconeogenesis and whole-body glucose homeostasis. Cell Metab. 22, 669–681 (2015).
Acknowledgements
We thank members of the University of Iowa Butler laboratory for assistance and helpful discussions, F. Lund (University of Alabama, Birmingham) for the μS–/– mice, T. Honjo (Kyoto University) for the Aicda–/– mice, T. Waldschmidt (University of Iowa) for the clone MR-1 antibody, J. Harty and S. Perlman (University of Iowa) for critical reading of the manuscript and helpful discussions, A. Pewa, E. Taylor and A. Rauckhorst (University of Iowa) for assistance with the metabolic measurements, G. Beuttener and B. Wagner (University of Iowa) for the metabolic flux assays, and members of the University of Iowa Flow Cytometry Facility for cell sorting. The research reported in this publication was supported by the NCI (grant number P30CA086862) and the National Center for Research Resources of the NIH (grant number S10OD016199). J.J.G. was supported by a Predoctoral Fellowship from the American Heart Association (grant number 16PRE27660002). F.A.S. was supported by the NIH (grant number T32 AI007485). K.J.R. was supported by the NIH (T32 GM067795). W.J.M. was supported by the NIH (grant numbers AI134733 and AI139902). H.-H.X. was supported by the NIH (grant numbers AI121080 and AI139874) and the Veteran Affairs BLR&D Merit Review Program (BX002903). C.R.E. was supported by an NHMRC Senior Research Fellowship (grant number 1154265) and an NHMRC Program grant (grant number 1132975). J.S.M. was supported by an NHMRC Program grant (grant number 1132975) and an NHMRC Practitioner Fellowship (grant number 1135955). M.J.B. was supported by an NHMRC Career Development Fellowship (grant number 1141632) and an NHMRC Project Grant (grant number 1141278). N.S.B. was supported by the NIH (grant numbers AI125446, AI127481 and AI139902). We thank the participants involved in the malaria VISs, Q-Pharm staff and the Medicine for Malaria Venture for funding the clinical trials.
Author information
Authors and Affiliations
Contributions
R.V., J.J.G. and N.S.B. conceptualized the project. J.J.G. performed the foundational studies and characterized the phenotype of plasmablasts in mice. R.V. characterized the transcriptome of the plasmablasts and conceived and executed the adoptive transfer studies, chimeric studies, metabolism analyses and glutamine supplementation experiments. W.J.M., K.J.R., F.L. and H.-H.X. provided critical reagents and technical assistance. C.R.E., J.S.M. and M.J.B. supervised the clinical studies. N.S.B. supervised the experimental rodent studies. R.V., J.J.G., H.-H.X., C.R.E., J.S.M., M.J.B. and N.S.B. designed the experiments. R.V., J.J.G., A.J.S., F.A.S., R.L.P., D.A., J.-A.C. and F.d.L.R. performed the experiments. R.V., J.J.G., A.J.S., F.A.S., R.R.S., J.-A.C., F.d.L.R., L.W., C.R.E., J.S.M., M.J.B. and N.S.B. analyzed the data. R.V. and R.R.S. performed the bioinformatics analyses. R.V. and J.J.G. made the figures. R.V. wrote the original draft. J.J.G., A.J.S., C.R.E., J.S.M., M.J.B. and N.S.B. reviewed and edited the manuscript and figures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information L. A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Experimental malaria expands extrafollicular plasmablast populations.
a, Gating strategy for identifying plasmablasts (CD138hiIgDneg), activated (CD138loIgDneg) and resting B cells (CD138loIgDhi). b, kinetics of PB in blood. Data are means ± s.d, representative of n = 2 biologically independent experiments with similar results using n = 3 mice/time point. c, CD138hiIgDneg plasmablasts were sort-purified from Py-infected mice on day 10 p.i., cultured for 20 hours and parasite lysate-specific IgM and IgG secreting ASCs were detected. Representative wells of ELISPOT assay. d, Relative CD19 expression by CD138hiIgDneg plasmablasts on days 7, 10, and 14 post Py infection. Data are representative of n > 5 experiments with similar results. e, BrdU incorporation in CD138hiIgDneg plasmablasts was assessed on day 10 p.i. Histogram represents BrdU staining, solid gray histogram is isotype (mouse IgG1). Data are representative of n = 2 experiments with similar results using n = 8 mice. f, Forward scatter and side scatter of CD138hiIgDneg plasmablasts examined on day 10 p.i. Data are representative of n > 5 experiments with similar results. g, Blimp-1/eYFP reporter mice were infected with Py. CD138hi Blimp-1/eYFP+ cells in bone marrow from naïve (left panel) or day 21 infected mice (right panel). Data are representative of n = 2 independent experiments with similar results using n = 4 mice/group. h,i, CD21 and CD23 expression by B cells in a naïve mouse (h) and day 10 Py-infected mouse (i) showing plasmablasts (green box) activated (blue box) and resting (red box) B cells. Data are representative of n > 5 experiments with similar results using n = 3 mice/group. j, Representative plots of the frequency and total numbers of GC B cells (GL7+CD95+) among plasmablasts, activated and resting B cell populations on day 10 p.i. Data are means ± s.e.m., representative of n = 3 experiments with similar results using n = 5 (Day 0 Total B cells) and n = 4 mice (each remaining group).
Extended Data Fig. 2 Developmental abrogation of blood stage Plasmodium infection-induced plasmablast responses.
a, Experimental design for adoptive transfers. b, Experimental design for Rosa26-ERT2/Cre × Prdm1fl/fl (CD45.2) : μMT bone marrow chimeric system. Eight weeks after engraftment, mice were infected with 1 × 106 Py and then treated with either corn oil or tamoxifen on days 4, 5, and 6 p.i. c, Gating strategy of TFH cells. d, Kinetics of parasite burden in Py-infected wild-type mice treated with tamoxifen or corn oil on days 4, 5, and 6 p.i. Data are means ± s.e.m., pooled from n = 2 biologically independent experiments with n = 6 mice/group. e, Experimental design for the three-way mixed bone marrow chimera. f, Eight weeks after engraftment, mice were infected with Py, treated with either corn oil or tamoxifen on days 4, 5, and 6 p.i. and the relative proportions (pie diagram) of Prdm1fl/fl (CD45.2) and wild-type (CD45.1) cells in the GC B cell compartment was analyzed. Data are means ± s.e.m., pooled from n = 2 biologically independent experiments with n = 4 mice/group. g, Evaluation of TH1 responses in PBS and tamoxifen-treated mixed bone marrow chimeric mice (as shown in b). Data are means ± s.d. representative of n = 2 biologically independent experiments with similar results using n = 5 (corn oil) and n = 4 mice (tamoxifen). Data in f,g were analyzed using two-tailed Mann-Whitney. Symbols and symbols represent individual mice.
Extended Data Fig. 3 Deletion of blood stage Plasmodium infection-induced plasmablast responses.
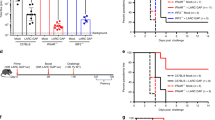
a, Experimental design for generating CD138-DTR chimeras. Eight weeks after engraftment, mice were infected with 1 × 106 Py and on days 5 and 7 p.i. treated with either DTx or PBS to delete plasmablasts. Data are means ± s.d., representative of n = 2 independent experiments with similar results using n = 4 mice/group. b-d, Py-infected wild-type mice were treated with either DTx or PBS on days 5 and 7 p.i. Kinetics of parasite burden (b), representative plots and summary data of GC B cells (c) and GC-Tfh cells (d) on 21 p.i. Data are means ± s.d., representative of n = 2 biologically independent experiments with similar results using n = 4 (PBS) and n = 3 mice (DTx). e-g, CD138-DTR chimeric mice were infected with 1 × 106 Py, plasmablasts were deleted with DTx and mice were subsequently treated with either MR-1 (anti-CD40L) or hamster IgG on days 8–11 p.i. Representative plots and summary data of GC B (e) and GC-TFH (f) cells as measured on day 21 p.i. and kinetics of parasite burden (g). Data are means ± s.e.m., pooled from n = 2 biologically independent experiments with n = 6 mice/group. Symbols in c-f represent individual mice and data were analyzed using two-tailed Mann-Whitney.
Extended Data Fig. 4 Differentially expressed genes among splenic B cell populations.
a, Venn diagram showing differentially expressed genes assessed by RNA-seq among the three splenic B cell populations on day 10 p.i. Respective cell types were sort-purified from n = 4 Py-infected mice. Data were obtained from one RNA-Seq experiment. Two-tailed ANOVA was used for identifying differentially expressed genes. b, Heat map showing the relative expression of all annotated genes assessed using RNA-Seq. c, Heat map showing the relative expression of genes involved in the unfolded protein response pathway (UPR).
Extended Data Fig. 5 L-glutamine enhances GC responses during experimental malaria.
a, L-glut concentrations in the spleens of naive and Py-infected wild-type mice on day 5 p.i. Data are means ± s.d. representative of n = 2 biologically independent experiments with similar results using n = 3 mice/group analyzed by a two-tailed unpaired t test (DF = 4; t = 5.933). b, c, Kinetics of parasite burden in mice treated with either L-alanine (b, H2O + L-ala), L-valine (c, H2O + L-val) or water starting on day 0 p.i. Data are means ± s.d., representative of n = 2 independent experiments with similar results using n = 3 mice/group. d–f, Py-infected wild-type mice were treated with L-glutamine (H2O + L-glut) or water starting day 0 p.i. and subsequently treated with MR-1 (anti-CD40L) or Hamster IgG antibody on days 8–11 p.i. Kinetics of parasite burden (d) and frequency of GC B cells (e) and GC-Tfh cells (f) on day 21 p.i. Data in d are means ± s.e.m, pooled from n = 2 independent experiments with n = 7 mice/group. Data in e,f are representative of n = 2 biologically independent experiments with similar results using n = 3 mice/group. g-m, Py-infected wild-type mice were treated with L-glutamine (H2O + L-glut) or water starting day 0 p.i. Kinetics of GC B cells (g), class-switched GC B cells (h), plasmablasts (i), and GC-TFH-like cells (j). Data in g-j are means ± s.d., representative of n = 2 biologically independent experiments with similar results using n = 6 mice/group analyzed by two-tailed Mann-Whitney. k, gMFI of BCL6 on GC-TFH-like cells. Data are means ± s.d., representative of n = 2 biologically independent experiments with similar results using n = 3 mice/group analyzed by two tailed unpaired t test (DF = 4; t = 1.257). Number of TH1 cells (l, Ly6C+CXCR3+IFNg+) and gMFI of CD80 and CD86 expression on splenic dendritic cells (m, MHCII+CD11c+) on day 10 p.i. Data in l,m are means ± s.e.m, pooled from n = 2 biologically independent experiments with n = 6 (H2O) and n = 7 mice (H2O + L-glut) analyzed by two-tailed Mann-Whitney. n,o, Kinetics of parasite burden (n) and area under curve as a measure of total parasite biomass (o) in Py-infected wild-type mice treated with L-glutamine (H2O + L-glut) starting on day 6 p.i. Data in n are means ± s.d., representative of n = 2 biologically independent experiments with similar results using n = 4 mice/group. Data in n analyzed using two-way ANOVA with Sidak’s multiple comparison (DF = 5; F = 5.728). Data in o analyzed with two-tailed Mann Whitney. p,q, Kinetics of parasite burden (p) and area under curve (q) as a measure of total parasite biomass in Py-infected wild-type mice treated with L-glutamine (H2O + L-glut) starting day on 10 p.i. Data in p,q are means ± s.d., representative of n = 2 independent experiments with similar results using n = 4 mice/group analyzed by two-tailed Mann Whitney. Symbols in a, l-q represent individual mice.
Extended Data Fig. 6 Experimental design for treating CD138-DTR: WT chimeras with L-glutamine.
Eight weeks after engraftment, mice were infected with Py and treated with L-glut or water starting on day 0 p.i. Mice were subsequently treated with either DTx or PBS on days 5 and 7 p.i. to delete plasmablasts.
Extended Data Fig. 7 Post-GC administration of L-glutamine does not appreciably enhance LLPC and MBC responses.
a, Gating strategy for long lived plasma cells (LLPCs) in the bone marrow of Py-infected mice. b, Numbers of LLPCs in the bone marrow on day 60 p.i. from Py-infected mice treated with either L-glutamine (H2O + L-glut) or water (H2O). Data are means ± s.e.m., pooled from n = 2 biologically independent experiments with n = 7 mice/group analyzed by two-tailed Mann Whitney. c,d, Py-infected wild-type mice were treated with either L-glutamine (H2O + L-glut) or water (H2O) at indicated time points p.i. and analyzed on day 30 p.i. Numbers of splenic memory B cells (c, CCR6+CD38+) and representative ELISPOT wells and summary data (d) demonstrating number of antibody-secreting LLPCs in the bone marrow. Data in c,d are means ± s.e.m, pooled from n = 2 biologically independent experiments with n = 5 (d, days 6–10) and n = 6 mice (remaining groups) analyzed by two-tailed Mann Whitney. Symbols represent individual mice.
Extended Data Fig. 8 Experimental design for the volunteer infection study.
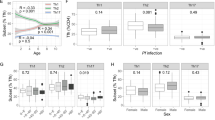
Two thousand eight hundred viable P. falciparum infected RBCs were intravenously into malaria naïve healthy volunteers (n = 36 men, n = 4 women). On day 8 p.i., volunteers received anti-malarial drug treatment. On days 0, 8 (treatment day), 14/15 and 27/28/36 (end-of-study, EOS), blood samples were collected and plasmablasts were assessed by flow cytometry.
Supplementary information
Supplementary Tables
Supplementary Tables 1–6.
Source data
Source Data Fig. 1
Statistical Source Data.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 6
Statistical Source Data.
Source Data Fig. 7
Statistical Source Data.
Source Data Fig. 8
Statistical Source Data.
Source Data Extended Data Fig. 1
Statistical Source Data.
Source Data Extended Data Fig. 2
Statistical Source Data.
Source Data Extended Data Fig. 3
Statistical Source Data.
Source Data Extended Data Fig. 5
Statistical Source Data.
Source Data Extended Data Fig. 7
Statistical Source Data.
Rights and permissions
About this article
Cite this article
Vijay, R., Guthmiller, J.J., Sturtz, A.J. et al. Infection-induced plasmablasts are a nutrient sink that impairs humoral immunity to malaria. Nat Immunol 21, 790–801 (2020). https://doi.org/10.1038/s41590-020-0678-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0678-5
This article is cited by
-
A candidate antibody drug for prevention of malaria
Nature Medicine (2024)
-
Single cell transcriptomics shows that malaria promotes unique regulatory responses across multiple immune cell subsets
Nature Communications (2023)
-
Dysregulated expression of amino-acid and glucose transporters on circulating plasma cells in septic shock patients: a preliminary study
Intensive Care Medicine Experimental (2022)
-
Supplying the trip to antibody production—nutrients, signaling, and the programming of cellular metabolism in the mature B lineage
Cellular & Molecular Immunology (2022)
-
Oncogenic RAS commandeers amino acid sensing machinery to aberrantly activate mTORC1 in multiple myeloma
Nature Communications (2022)