Abstract
Multiple sclerosis (MS) is characterized by pathological inflammation that results from the recruitment of lymphoid and myeloid immune cells from the blood into the brain. Due to subset heterogeneity, defining the functional roles of the various cell subsets in acute and chronic stages of MS has been challenging. Here, we used index and transcriptional single-cell sorting to characterize the mononuclear phagocytes that infiltrate the central nervous system from the periphery in mice with experimentally induced autoimmune encephalomyelitis, a model of MS. We identified eight monocyte and three dendritic cell subsets at acute and chronic disease stages in which the defined transcriptional programs pointed toward distinct functions. Monocyte-specific cell ablation identified Cxcl10+ and Saa3+ monocytic subsets with a pathogenic potential. Transfer experiments with different monocyte and precursor subsets indicated that these Cxcl10+ and Saa3+ pathogenic cells were not derived from Ly6C+ monocytes but from early myeloid cell progenitors. These results suggest that blocking specific pathogenic monocytic subsets, including Cxcl10+ and Saa3+ monocytes, could be used for targeted therapeutic interventions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data generated during this study have been deposited in Gene Expression Omnibus with the accession code GSE144317.
Code availability
Scripts and auxiliary data needed to reconstruct analysis files will be made available by request.
Change history
24 June 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41590-020-0742-1
08 June 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41590-020-0722-5
References
Geissmann, F., Jung, S. & Littman, D. R. Blood monocytes consist of two principal subsets with distinct migratory properties. Immunity 19, 71–82 (2003).
Carlin, L. M. et al. Nr4a1-dependent Ly6Clow monocytes monitor endothelial cells and orchestrate their disposal. Cell 153, 362–375 (2013).
Mildner, A., Yona, S. & Jung, S. A close encounter of the third kind: monocyte-derived cells. Adv. Immunol. 120, 69–103 (2013).
Mildner, A. et al. Genomic characterization of murine monocytes reveals C/EBPβ transcription factor dependence of Ly6C− cells. Immunity 46, 849–862.e7 (2017).
Menezes, S. et al. The heterogeneity of Ly6Chi monocytes controls their differentiation into iNOS+ macrophages or monocyte-derived dendritic cells. Immunity 45, 1205–1218 (2016).
Yáñez, A. et al. Granulocyte-monocyte progenitors and monocyte-dendritic cell progenitors independently produce functionally distinct monocytes. Immunity 47, 890–902.e4 (2017).
Liu, Z. et al. Fate mapping via Ms4a3-expression history traces monocyte-derived cells. Cell 178, 1509–1525.e19 (2019).
Mildner, A. et al. CCR2+Ly-6Chi monocytes are crucial for the effector phase of autoimmunity in the central nervous system. Brain 132, 2487–2500 (2009).
King, I. L., Dickendesher, T. L. & Segal, B. M. Circulating Ly-6C+ myeloid precursors migrate to the CNS and play a pathogenic role during autoimmune demyelinating disease. Blood 113, 3190–3197 (2009).
Ajami, B., Bennett, J. L., Krieger, C., McNagny, K. M. & Rossi, F. M. V. Infiltrating monocytes trigger EAE progression, but do not contribute to the resident microglia pool. Nat. Neurosci. 14, 1142–1149 (2011).
Croxford, A. L. et al. The cytokine GM-CSF drives the inflammatory signature of CCR2+ monocytes and licenses autoimmunity. Immunity 43, 502–514 (2015).
Spath, S. et al. Dysregulation of the cytokine GM-CSF induces spontaneous phagocyte invasion and immunopathology in the central nervous system. Immunity 46, 245–260 (2017).
Jaitin, D. A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Locatelli, G. et al. Mononuclear phagocytes locally specify and adapt their phenotype in a multiple sclerosis model. Nat. Neuroscience. 21, 1196–1208 (2018).
Caravagna, C. et al. Diversity of innate immune cell subsets across spatial and temporal scales in an EAE mouse model. Sci. Rep. 8, 5146 (2018).
Lewis, N. D., Hill, J. D., Juchem, K. W., Stefanopoulos, D. E. & Modis, L. K. RNA sequencing of microglia and monocyte-derived macrophages from mice with experimental autoimmune encephalomyelitis illustrates a changing phenotype with disease course. J. Neuroimmunol. 277, 26–38 (2014).
Giladi, A. et al. Single-cell characterization of haematopoietic progenitors and their trajectories in homeostasis and perturbed haematopoiesis. Nat. Cell Biol. 20, 836–846 (2018).
Baran, Y. et al. MetaCell: analysis of single-cell RNA-seq data using K-nn graph partitions. Genome Biol. 20, 206–219 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Meredith, M. M. et al. Zinc finger transcription factor zDC is a negative regulator required to prevent activation of classical dendritic cells in the steady state. J. Exp. Med. 209, 1583–1593 (2012).
Satpathy, A. T. et al. Zbtb46 expression distinguishes classical dendritic cells and their committed progenitors from other immune lineages. J. Exp. Med. 209, 1135–1152 (2012).
Briseño, C. G. et al. Distinct transcriptional programs control cross-priming in classical and monocyte-derived dendritic cells. Cell Rep. 15, 2462–2474 (2016).
Wolf, Y. et al. Microglial MHC class II is dispensable for experimental autoimmune encephalomyelitis and cuprizone-induced demyelination. Eur. J. Immunol. 48, 1308–1318 (2018).
Brühl, H. et al. Targeting of Gr-1+,CCR2+ monocytes in collagen-induced arthritis. Arthritis Rheum. 56, 2975–2985 (2007).
Varol, C. et al. Monocytes give rise to mucosal, but not splenic, conventional dendritic cells. J. Exp. Med. 204, 171–180 (2007).
Matcovitch-Natan, O. et al. Microglia development follows a stepwise program to regulate brain homeostasis. Science 353, aad8670 (2016).
Shemer, A. et al. Engrafted parenchymal brain macrophages differ from microglia in transcriptome, chromatin landscape and response to challenge. Nat. Commun. 9, 5206 (2018).
Becher, B., Tugues, S. & Greter, M. GM-CSF: from growth factor to central mediator of tissue inflammation. Immunity 45, 963–973 (2016).
Esiri, M. M. & Reading, M. C. Macrophage populations associated with multiple sclerosis plaques. Neuropathol. Appl. Neurobiol. 13, 451–465 (1987).
Brück, W. et al. Monocyte/macrophage differentiation in early multiple sclerosis lesions. Ann. Neurol. 38, 788–796 (1995).
Melero-Jerez, C., Ortega, M. C., Moliné-Velázquez, V. & Clemente, D. Myeloid derived suppressor cells in inflammatory conditions of the central nervous system. Biochim. Biophys. Acta. 1862, 368–380 (2016).
McQualter, J. L. et al. Granulocyte macrophage colony-stimulating factor: a new putative therapeutic target in multiple sclerosis. J. Exp. Med. 194, 873–882 (2001).
Greter, M. et al. GM-CSF controls nonlymphoid tissue dendritic cell homeostasis but is dispensable for the differentiation of inflammatory dendritic cells. Immunity 36, 1031–1046 (2012).
Blecher-Gonen, R. et al. Single-cell analysis of diverse pathogen responses defines a molecular roadmap for generating antigen-specific immunity. Cell Syst. 8, 109–121.e6 (2019).
Hirako, I. C. et al. Splenic differentiation and emergence of CCR5+CXCL9+CXCL10+ monocyte-derived dendritic cells in the brain during cerebral malaria. Nat. Commun. 7, 13277 (2016).
Serbina, N. V., Salazar-Mather, T. P., Biron, C. A., Kuziel, W. A. & Pamer, E. G. TNF/iNOS-producing dendritic cells mediate innate immune defense against bacterial infection. Immunity 19, 59–70 (2003).
Paré, A. et al. IL-1β enables CNS access to CCR2hi monocytes and the generation of pathogenic cells through GM-CSF released by CNS endothelial cells. Proc. Natl Acad. Sci. USA 115, E1194–E1203 (2018).
Ronchi, F. et al. Experimental priming of encephalitogenic Th1/Th17 cells requires pertussis toxin-driven IL-1β production by myeloid cells. Nat. Commun. 7, 11541 (2016).
Fife, B. T. et al. CXCL10 (IFN-γ-inducible protein-10) control of encephalitogenic CD4+ T cell accumulation in the central nervous system during experimental autoimmune encephalomyelitis. J. Immunol. 166, 7617–7624 (2001).
Zilionis, R. et al. Single-cell transcriptomics of human and mouse lung cancers reveals conserved myeloid populations across individuals and species. Immunity 50, 1317–1334 (2019).
Cohen, M. et al. Lung single-cell signaling interaction map reveals basophil role in macrophage imprinting. Cell 175, 1031–1044.e18 (2018).
Hettinger, J. et al. Origin of monocytes and macrophages in a committed progenitor. Nat. Immunol. 14, 821–830 (2013).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods. 12, 357–360 (2015).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Supek, F., Bošnjak, M., Škunca, N. & Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6, e21800 (2011).
Acknowledgements
We thank V. Malchin for excellent technical assistance, J. Priller for support, the MDC animal facility (especially J. Bergemann) and the MDC and WIS FACS facilities (especially H.-P. Rahn). A.M. is a Heisenberg fellow supported by the DFG (MI1328). I.A. is supported by the Chan-Zuckerberg Initiative, the HHMI International Scholar award, the European Research Council Consolidator grant (724471-HemTree2.0), the Thompson Family Foundation, an MRA Established Investigator Award (509044), the Israel Science Foundation (703/15), the Ernest and Bonnie Beutler Research Program for Excellence in Genomic Medicine, the Helen and Martin Kimmel award for innovative investigation, an International Progressive MS Alliance/NMSS PA-1604-08459 and an Adelis Foundation grant. S.J. is supported by the International Progressive MS Alliance/NMSS PA-1604-08459.
Author information
Authors and Affiliations
Contributions
A.M. designed and, together with I.A., supervised the study. L.K.W., H.L., D.D., C.M., F.P. and A.S. performed the experiments. A.G. performed bioinformatical analysis. S.J., S.Y., M.M. and A.L. provided key reagents, mouse lines and intellectual input. A.M. and I.A. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Ioana Visan was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Quality control of the scRNA-seq data.
a, List of experiment and cell numbers used in this study. The number of cells represented here are numbers before exclusion of contaminating lymphocytes or neutrophils. b, Number of Illumina reads and c, total UMI per single cell. d, Fraction of analyzed cells after filtering. Cells are grouped and colored by experimental procedure.
Extended Data Fig. 2 Identification of mononuclear phagocyte subsets in the inflamed CNS.
a, Comparing Seurat and MetaCell clustering results. Rows represent the 55 identified metacells, grouped by their cell identity, and columns represent Seurat clusters. Color intensity in each entry depicts the number of cells assigned to a specific combination of MetaCells and Seurat clusters. b, Pairwise correlation analysis of the 10 distinct cell populations. Shown here are the numbers of differential expressed genes. c, Expression quantiles of key cell-type-specific marker genes on top of the 2D projection map. n = 2,897 single cells were analyzed. d, Top 10 differentially expressed genes in each cluster (log2 fold change). e, n = 2,897 single cells were classified into the 10 indicated metacell subsets (color bar) and the top 60 differential expressed genes were used for GO-enrichment of each cluster (these genes can be found in Supplementary Table 2). Circle color indicates p-value and size indicates number of genes. P-values indicate Benjamini Hochberg adjusted GSEA permutation tests.
Extended Data Fig. 3 Characterization of dendritic cells in the inflamed CNS.
a, EAE was induced in Zbtb46-GFP mice and cells were isolated from the inflamed spinal cord at the acute (day 15 PI; mean score: 2,7 ± 0,4 SEM; n = 5) and the chronic phase (day 30 PI; mean score 2,2 ± 0,4 SEM; n = 6). b, FACS analysis of LinnegLy6G−CD44highCX3CR1low-to-intCD11b+ cellular infiltrates into the spinal cord of acute (pool of n = 6 mice) and chronic diseased EAE Zbtb46Gfp/+ mice (pool of n = 7 mice). Shown is the gating for sorting GFP+ cells as indicated by the red square. c, Projection of 1,282 Zbtb46-GFP+ cells on the 2D projection as shown in Fig. 1. d, Upper panel: Expression profiles of 1,056 infiltrated Zbtb46-GFP+ cDC cells that clustered into 15 DC metacells according to their transcriptomic similarities. Colorbar represent grouping of cells into three major cDC clusters. Dark violet correspond to cDC1 subset. Lower panel: MFI of Zbtb46-GFP expression in the sorted cells is shown on the bottom of the heatmap. Red dots indicate cells isolated during the acute phase, while blue dots indicate cells from the chronic phase. e, Expression quantiles of key cell-type-specific marker genes on top of the projection map. Single cell data represented in a-e are representative of one experiment.
Extended Data Fig. 4 Temporal resolution of cDC infiltrates in the acute and chronic stages of EAE.
a, Projection of Zbtb46-GFP+ cells from Extended Data Fig. 3 separated according to the acute (left; pool of n = 5 mice) and chronic (right; pool of n = 6 mice) stage of EAE. n = 702 cells from acute and 580 from chronic disease stages were analyzed. b, Cell distribution of Zbtb46-GFP+ cells from both stages of disease. c, Differential gene expression between acute and chronic cDC. Values represent log-transformed normalized expression. Single cell data represented in a-c are representative of one experiment.
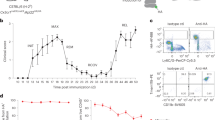
Extended Data Fig. 5 Effects of MC21 depletion.
a, Mice were immunized with MOG35-55 and animals received at the peak of disease six injections of either 50 μg of isotype control antibody (rat IgG2b) or 50 μg purified anti-CCR2 (MC21). Shown is the mean clinical course ±SEM. N = 6-7 mice per group and asterisk indicates statistical significance with * p < 0,05 and ** p < 0,005; unpaired two-tailed T-test. Data are representative of one experiment with six mice. b, FACS analysis (left) and quantification (right; mean ± SD) of Ly6C+ MHCII+ (IAb) monocytes in the blood of isotype or MC21 treated mice. N = 3 mice per group, asterisk indicates statistical significance with p < 0,05; unpaired two-tailed T-test. The experiment was repeated three times with similar results. c, Analysis and quantification of splenic immune cells in EAE mice that received two injections of 50 μg isotype control antibody or 50 μg purified anti-CCR2. Shown are % of the respective cell populations out of CD45+ cells (n = 4 animals per group; experiment was performed twice with similar results; mean ± SD; asterisk indicates statistical significance with p < 0,01; unpaired two-tailed T-test). Tregs were identified as CD4+FoxP3+. d, Analysis and quantification of blood immune cells in EAE mice that received two injections of 50μg isotype control antibody or 50μg purified anti-CCR2 (MC21). Shown are % of the respective cell populations out of CD45+ cells (n = 4 animals per group; experiment was performed twice with similar results; mean ± SD; asterisk indicates statistical significance with p < 0,05; unpaired two-tailed T-test). e, Repetition of Fig. 3 in an independent mouse facility and with purified MC21 antibody. Wt animals received either of 50μg isotype control antibody or 50μg purified anti-CCR2 at the peak of disease for two consecutive days. Shown are the EAE courses during the experiment (day 16 PI, mean score in each group: isotype: 2.7 ± 0.3 SEM; MC21 3.0 ± 0.3 SEM; asterisk indicates statistical significance with p < 0,01; unpaired two-tailed T-test;). f, Projection of CD44+Ly6G−CD11b+ non-neutrophilic, non-microglial cells from isotype- (left) and MC21-treated (right) animals on the metacell model from Fig. 1. g, Bar plots showing enrichment (log2 fold change) of myeloid groups in MC21-treated mice compared to isotype controls. Error bars represent 95% confidence intervals. 3 mice were pooled for MARS-seq analysis depicted in f,g and n = 232 cells from isotype- and 147 cells from MC21-treated mice were analyzed in f, g.
Extended Data Fig. 6 Expression of Cxcl9 and Cxcl10 in comparison to Arg1.
Shown is the log2 enrichment over median of Cxcl9 and Cxcl10 against Arg1 in the main 6 monocytic clusters.
Extended Data Fig. 7 Identification of MDP and Ly6C+ derived cells in the CNS after sequential transfer.
a, MDP were isolated from CD45.1/1 mice and BM Ly6C+ monocytes were extracted from Ubc-GFP mice as shown in Fig. 6a. b, Heatmap depicting gene expression across the transferred cells. c, Each cell was assigned to its GFP and Ly6C expression according to the indexed FACS measurement. Shown is the mean fluorescence intensity of each marker. d, 2×106 Ly6C+ monocytes were isolated from CD45.1/1 mice and transferred at the peak of disease into nine CD45.2/2 recipient mice (day 13 PI; mean clinical score ± SEM are shown). e, 4 days after transfer, only 40 transferred cells could be re-isolated from the pooled spinal cord of recipients that showed no Ly6C and no MHCII expression. f, scRNA-seq identified that the majority of grafted cells show a Ly6C− monocyte signature, while the remaining cells correspond to microglia-like cells and to Arg1+ subsets. Single cell data represented in d-f are representative of one experiment.
Supplementary information
Supplementary Tables
In Supplementary Table 1 (sheet 2), we represent the log2 enrichment over median (lfp, see Methods) of the identified 55 metacell clusters depicted in Fig. 1 combined with Extended Data Fig. 3. X1–X8, Arg1+ macrophages I; X9–X17, Nos2+ macrophages; X18–X21, Saa3+ monocytes; X22–X27, Cxcl10+ monocytes; X28-X34, Ly6C+ monocytes; X35–X36, microglia(-like) cells; X37–X52, cDC; X53, Ifit2+ monocytes; X54, Ly6C– monocytes; X55, Arg1+ macrophages II. UMI counts for each gene are listed at the end of the table. In Supplementary Table 2 (sheet 3), the 60 top differentially expressed genes are listed, which were used for gene-ontology enrichment analysis (see Extended Data Fig. 2).
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 7
Statistical Source Data
Rights and permissions
About this article
Cite this article
Giladi, A., Wagner, L.K., Li, H. et al. Cxcl10+ monocytes define a pathogenic subset in the central nervous system during autoimmune neuroinflammation. Nat Immunol 21, 525–534 (2020). https://doi.org/10.1038/s41590-020-0661-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0661-1
This article is cited by
-
Skull bone marrow channels as immune gateways to the central nervous system
Nature Neuroscience (2023)
-
Systemic innate myeloid responses to acute ischaemic and haemorrhagic stroke
Seminars in Immunopathology (2023)
-
Myelin insulation as a risk factor for axonal degeneration in autoimmune demyelinating disease
Nature Neuroscience (2023)
-
Identification of astrocyte regulators by nucleic acid cytometry
Nature (2023)
-
Central nervous system macrophages in progressive multiple sclerosis: relationship to neurodegeneration and therapeutics
Journal of Neuroinflammation (2022)