Abstract
Crosstalk between mesenchymal stromal cells (MSCs) and hematopoietic stem cells (HSCs) is essential for hematopoietic homeostasis and lineage output. Here, we investigate how transcriptional changes in bone marrow (BM) MSCs result in long-lasting effects on HSCs. Single-cell analysis of Cxcl12-abundant reticular (CAR) cells and PDGFRα+Sca1+ (PαS) cells revealed an extensive cellular heterogeneity but uniform expression of the transcription factor gene Ebf1. Conditional deletion of Ebf1 in these MSCs altered their cellular composition, chromatin structure and gene expression profiles, including the reduced expression of adhesion-related genes. Functionally, the stromal-specific Ebf1 inactivation results in impaired adhesion of HSCs, leading to reduced quiescence and diminished myeloid output. Most notably, HSCs residing in the Ebf1-deficient niche underwent changes in their cellular composition and chromatin structure that persist in serial transplantations. Thus, genetic alterations in the BM niche lead to long-term functional changes of HSCs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding authors upon request. Source data for Extended Data Fig. 1d are provided with the paper. All high-throughput data in this study were deposited at the Gene Expression Omnibus (GEO) under series GSE127970 (CAR, PαS RNA-seq and ATAC-seq), series GSE128089 (HSC RNA-seq and ATAC-seq), and series GSE128743 (single-cell RNA-seq). A web application for scRNA-seq data visualization is available at https://hematopoietic-niche-regulation-by-ebf1.ie-freiburg.mpg.de.
References
Joseph, C. et al. Deciphering hematopoietic stem cells in their niches: a critical appraisal of genetic models, lineage tracing, and imaging strategies. Cell Stem Cell 13, 520–533 (2013).
Mendelson, A. & Frenette, P. S. Hematopoietic stem cell niche maintenance during homeostasis and regeneration. Nat. Med. 20, 833–846 (2014).
Morrison, S. J. & Scadden, D. T. The bone marrow niche for haematopoietic stem cells. Nature 505, 327–334 (2014).
Acar, M. et al. Deep imaging of bone marrow shows non-dividing stem cells are mainly perisinusoidal. Nature 526, 126–130 (2015).
Cordeiro Gomes, A. et al. Hematopoietic stem cell niches produce lineage-instructive signals to control multipotent progenitor differentiation. Immunity 45, 1219–1231 (2016).
Ding, L. & Morrison, S. J. Haematopoietic stem cells and early lymphoid progenitors occupy distinct bone marrow niches. Nature 495, 231–235 (2013).
Itkin, T. et al. Distinct bone marrow blood vessels differentially regulate haematopoiesis. Nature 532, 323–328 (2016).
Mendez-Ferrer, S. et al. Mesenchymal and haematopoietic stem cells form a unique bone marrow niche. Nature 466, 829–834 (2010).
Kfoury, Y. & Scadden, D. T. Mesenchymal cell contributions to the stem cell niche. Cell Stem Cell 16, 239–253 (2015).
Pinho, S. & Frenette, P. S. Haematopoietic stem cell activity and interactions with the niche. Nat. Rev. Mol. Cell Biol. 20, 303–320 (2019).
Omatsu, Y. et al. The essential functions of adipo-osteogenic progenitors as the hematopoietic stem and progenitor cell niche. Immunity 33, 387–399 (2010).
Zhou, B. O., Yue, R., Murphy, M. M., Peyer, J. G. & Morrison, S. J. Leptin-receptor-expressing mesenchymal stromal cells represent the main source of bone formed by adult bone marrow. Cell Stem Cell 15, 154–168 (2014).
Greenbaum, A. et al. CXCL12 in early mesenchymal progenitors is required for haematopoietic stem-cell maintenance. Nature 495, 227–230 (2013).
Nagasawa, T. The chemokine CXCL12 and regulation of HSC and B lymphocyte development in the bone marrow niche. Adv. Exp. Med. Biol. 602, 69–75 (2007).
Morikawa, S. et al. Prospective identification, isolation, and systemic transplantation of multipotent mesenchymal stem cells in murine bone marrow. J. Exp. Med. 206, 2483–2496 (2009).
Asada, N. et al. Differential cytokine contributions of perivascular haematopoietic stem cell niches. Nat. Cell Biol. 19, 214–223 (2017).
Kunisaki, Y. et al. Arteriolar niches maintain haematopoietic stem cell quiescence. Nature 502, 637–643 (2013).
Chen, J. Y. et al. Hoxb5 marks long-term haematopoietic stem cells and reveals a homogenous perivascular niche. Nature 530, 223–227 (2016).
Chan, C. K. et al. Endochondral ossification is required for haematopoietic stem-cell niche formation. Nature 457, 490–494 (2009).
Mizoguchi, T. et al. Osterix marks distinct waves of primitive and definitive stromal progenitors during bone marrow development. Dev. Cell 29, 340–349 (2014).
Omatsu, Y., Seike, M., Sugiyama, T., Kume, T. & Nagasawa, T. Foxc1 is a critical regulator of haematopoietic stem/progenitor cell niche formation. Nature 508, 536–540 (2014).
Kieslinger, M., Hiechinger, S., Dobreva, G., Consalez, G. G. & Grosschedl, R. Early B cell factor 2 regulates hematopoietic stem cell homeostasis in a cell-nonautonomous manner. Cell Stem Cell 7, 496–507 (2010).
Qian, H. et al. Molecular characterization of prospectively isolated multipotent mesenchymal progenitors provides new insight into the cellular identity of mesenchymal stem cells in mouse bone marrow. Mol. Cell. Biol. 33, 661–677 (2013).
Seike, M., Omatsu, Y., Watanabe, H., Kondoh, G. & Nagasawa, T. Stem cell niche-specific Ebf3 maintains the bone marrow cavity. Genes Dev. 32, 359–372 (2018).
Lin, H. & Grosschedl, R. Failure of B-cell differentiation in mice lacking the transcription factor EBF. Nature 376, 263–267 (1995).
Jimenez, M. A., Akerblad, P., Sigvardsson, M. & Rosen, E. D. Critical role for Ebf1 and Ebf2 in the adipogenic transcriptional cascade. Mol. Cell. Biol. 27, 743–757 (2007).
Hesslein, D. G. et al. Ebf1-dependent control of the osteoblast and adipocyte lineages. Bone 44, 537–546 (2009).
Grün, D. et al. De novo prediction of stem cell identity using single-cell transcriptome data. Cell Stem Cell 19, 266–277 (2016).
Herman, J. S., Sagar & Grun, D. FateID infers cell fate bias in multipotent progenitors from single-cell RNA-seq data. Nat. Methods 15, 379–386 (2018).
Logan, M. et al. Expression of Cre recombinase in the developing mouse limb bud driven by a Prxl enhancer. Genesis 33, 77–80 (2002).
Baryawno, N. et al. A cellular taxonomy of the bone marrow stroma in homeostasis and leukemia. Cell 177, 1915–1932.e16 (2019).
Tikhonova, A. N. et al. Author Correction: The bone marrow microenvironment at single-cell resolution. Nature 572, E6 (2019).
Fretz, J. A. et al. Altered metabolism and lipodystrophy in the early B-cell factor 1-deficient mouse. Endocrinology 151, 1611–1621 (2010).
Pinho, S. et al. Lineage-biased hematopoietic stem cells are regulated by distinct niches. Dev. Cell 44, 634–641.e4 (2018).
Ruppert, R. et al. Kindlin-3-mediated integrin adhesion is dispensable for quiescent but essential for activated hematopoietic stem cells. J. Exp. Med. 212, 1415–1432 (2015).
Ellis, S. J. & Tanentzapf, G. Integrin-mediated adhesion and stem-cell-niche interactions. Cell Tissue Res. 339, 121–130 (2010).
Boller, S. et al. Pioneering activity of the C-terminal domain of EBF1 shapes the chromatin landscape for B cell programming. Immunity 44, 527–541 (2016).
Li, R. et al. Dynamic EBF1 occupancy directs sequential epigenetic and transcriptional events in B-cell programming. Genes Dev. 32, 96–111 (2018).
Nerlov, C. & Graf, T. PU.1 induces myeloid lineage commitment in multipotent hematopoietic progenitors. Genes Dev. 12, 2403–2412 (1998).
Iwasaki, H. et al. Distinctive and indispensable roles of PU.1 in maintenance of hematopoietic stem cells and their differentiation. Blood 106, 1590–1600 (2005).
Friedman, A. D. C/EBPα induces PU.1 and interacts with AP-1 and NF-κB to regulate myeloid development. Blood Cells Mol. Dis. 39, 340–343 (2007).
Obier, N. et al. Cooperative binding of AP-1 and TEAD4 modulates the balance between vascular smooth muscle and hemogenic cell fate. Development 143, 4324–4340 (2016).
Kurotaki, D. et al. Transcription factor IRF8 governs enhancer landscape dynamics in mononuclear phagocyte progenitors. Cell Rep. 22, 2628–2641 (2018).
Vierbuchen, T. et al. AP-1 transcription factors and the BAF complex mediate signal-dependent enhancer selection. Mol. Cell 68, 1067–1082.e12 (2017).
Jones, M. et al. Ash1l controls quiescence and self-renewal potential in hematopoietic stem cells. J. Clin. Invest. 125, 2007–2020 (2015).
Min, I. M. et al. The transcription factor EGR1 controls both the proliferation and localization of hematopoietic stem cells. Cell Stem Cell 2, 380–391 (2008).
Laurenti, E. et al. CDK6 levels regulate quiescence exit in human hematopoietic stem cells. Cell Stem Cell 16, 302–313 (2015).
Comazzetto, S. et al. Restricted hematopoietic progenitors and erythropoiesis require SCF from leptin receptor+ niche cells in the bone marrow. Cell Stem Cell 24, 477–486.e6 (2019).
Medyouf, H. The microenvironment in human myeloid malignancies: emerging concepts and therapeutic implications. Blood 129, 1617–1626 (2017).
Raaijmakers, M. H. Niche contributions to oncogenesis: emerging concepts and implications for the hematopoietic system. Haematologica 96, 1041–1048 (2011).
Gyory, I. et al. Transcription factor Ebf1 regulates differentiation stage-specific signaling, proliferation, and survival of B cells. Genes Dev. 26, 668–682 (2012).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Hashimshony, T. et al. CEL-Seq2: sensitive highly-multiplexed single-cell RNA-Seq. Genome Biol. 17, 77 (2016).
Sagar, Herman, J. S., Pospisilik, J. A. & Grun, D. High-throughput single-cell RNA sequencing and data analysis. Methods Mol. Biol. 1766, 257–283 (2018).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Grun, D., Kester, L. & van Oudenaarden, A. Validation of noise models for single-cell transcriptomics. Nat. Methods 11, 637–640 (2014).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Piper, J. et al. Wellington: a novel method for the accurate identification of digital genomic footprints from DNase-seq data. Nucleic Acids Res. 41, e201 (2013).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Montague, T. G., Cruz, J. M., Gagnon, J. A., Church, G. M. & Valen, E. CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res. 42, W401–W407 (2014).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Acknowledgements
We are grateful to E. Trompouki for critical reading of the manuscript and M. Rott for assistance with the manuscript preparation. We thank S. Muellerke, I. Falk, K. Schlegel, D. Komandin and F. Ludin for technical assistance and members of the R. Grosschedl laboratory for discussions. We thank the Deep Sequencing, Imaging, FACS and Animal facilities of the Max Planck Insitute of Immunobiology and Epigenetics. This work was supported by the funds from the Max Planck Society and German Research Foundation. D. Grün was supported by the Max Planck Society, the German Research Foundation (DFG) (GR4980/1-1, GR4980/3-1, and GRK2344 MeInBio) and by the DFG under Germany’s Excellence Strategy (CIBSS-EXC-2189 Project ID 390939984).
Author information
Authors and Affiliations
Contributions
M.D. designed and performed experiments. S.R. and P.C. performed bioinformatics analysis. J.S.H. performed and analyzed single-cell RNA-seq experiments and created a shiny web application. D.G. supervised J.S.H. E.L. performed tail vein injections. R.G. conceived and supervised the study. R.G. and M.D. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information L. A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Ebf1 deficiency alters MSC composition and gene expression.
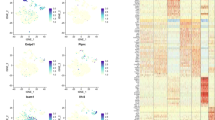
a, Representative gating strategy used to identify and isolate CAR and PaS cells. b, Heat map depicting log2-transformed normalized expression of signature genes (rows) across the cells in each cluster (columns). Key genes are indicated on the right. Color code of clusters corresponds to Fig.1a. All clusters were compared in a pairwise fashion against each other and differentially expressed genes (adjusted P-value<0.05) appearing in at least 3 comparisons. Data represent 912 single-sorted wild-type cells from 3 independent experiments (n=4 mice). Negative binomial distributions reflecting the gene expression variability within each subgroup were inferred, based on a background model for the expected transcript count variability, a P-value for the observed difference in transcript counts between the two subgroups is computed as described for DESeq2. The P-values were corrected for multiple testing via the Benjamini–Hochberg method. c, t-SNE maps showing normalized expression of Ptprc, Rag1 and J chain in sequences CAR (CD45–CD31–Lin–PDGFRα+Sca1–) and PαS (CD45–CD31–Lin–PDGFRα+Sca1+) cells. d, Representative immunoblot analysis of EBF1 protein level in total cells extracts from 1x104 sorted cells (n=2 independent experiments). Actin was used as a loading control. EBF1 band is marked with an arrow. e, t-SNE map showing clusters of cells with similar transcriptomes within CAR and PαS populations, derived by RaceID3 algorithm. Data represent 1777 single-sorted wild-type and knockout cells from 3 independent experiments (n=4 mice per genotype). f, t-SNE map showing distribution of CAR and PαS cells from Ebf1+/+Prx1Cre mice (black) and from Ebf1fl/flPrx1Cre mice (blue) within established clusters. g, MA plot showing differential gene expression between Ebf1+/+Prx1Cre and Ebf1fl/flPrx1Cre cells from scRNA-seq analysis. Upregulated genes are shown in red and downregulated genes are shown in blue. Source data.
Extended Data Fig. 2 Ebf1 deficiency alters MSC in vitro differentiation.
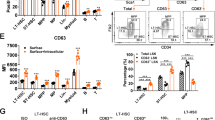
Integrated analysis of the scRNA-seq dataset from CAR and PαS cells from Ebf1+/+Prx1Cre and Ebf1fl/flPrx1Cre mice with data published by Tikhonova et al. and Baryawno et al. a, UMAP plot depicting cells collored by used technology CEL-Seq2 (blue) utilized by Derecka et al. study or 10x (red) utlized by Tikhonova et al. study. b, UMAP plot depicting cells collored by used technology CEL-Seq2 (blue) utilized by Derecka et al. study or 10x (red) utlized by Baryawno et al. study. c, Percentage of Annexin V positive CAR and PαS cells in the BM. Data represent the mean ± s.e.m, for 3 independent experiments, n=13 Ebf1+/+Prx1Cre mice and n=12 Ebf1fl/flPrx1Cre mice. Percentage of CAR and PαS cells within each cell cycle phase. Data represent the mean ± s.e.m, for 3 independent experiments, n=11 (CAR) and n=14 (PαS) Ebf1+/+Prx1Cre mice and n=11 (CAR) and n=12 (PαS) Ebf1fl/flPrx1Cre mice (*P=0.02 for G0, *P=0.04 for G1, *P=0.01 for s/G2/M). d, Histomorphometric analysis of tibiae from n=12 Ebf1+/+Prx1Cre and n=12 Ebf1fl/flPrx1Cre mice showing bone volume as a percentage of tissue volume (BV/TV) in the trabecular area of the bone (*P=0.01) and osteoblast surface as a percentage of bone surface (Ob.S/BS) in the trabecular (*P=0.03) and endocortical area of the bone. e, Quantification of CFU-F colonies from sorted CAR and PaS cells. Data are presented as mean ± s.e.m for 3 independent experiments, n=9 for Ebf1+/+Prx1Cre and n=10 for Ebf fl/flPrx1Cre mice (*p=0.03, ***p=0.0003). f, Representative images showing multilineage differetiation of CFU-F colonies from PαS cells from Ebf1+/+Prx1Cre and Ebf1fl/flPrx1Cre mice (n=3 independent experiments). Adipocytes were stained with Oild Red O, osteoblasts with Alizarin Red S and chondrocytes with Toluidine Blue. g, Relative mRNA expression levels of mesenchymal lineage-specific markers from PαS-derived CFU-F colonies sub-cloned into secondary cultures and subjected to adipogenic, osteogenic and chondrogenic in vitro differentiation. Data are represented as mean fold change ± s.e.m over Ebf1+/+Prx1Cre cells set as 1, for 3 independent experiments, n=26 clones from Ebf1+/+Prx1Cre mice and n=35 clones from Ebf1fl/flPrx1Cre mice (Adipogenic differentiation); n=32 clones for each genotype (Osteogenic fifferentiation); n=38 clones from Ebf1+/+Prx1Cre mice and n=48 clones from Ebf1fl/flPrx1Cre mice (Chondrogenic differentiation). Statistical analysis in c, d, e, g was performed using two-tailed unpaired Student’s t-test.
Extended Data Fig. 3 Prx1Cre mediated deletion of Ebf1.
a, Total BM cellularity. Data represent the mean ± s.e.m, n=22 Ebf1+/+Prx1Cre mice and n=20 Ebf1fl/flPrx1Cre mice. b, Quantification of CFU colonies from the spleen and peripheral blood. Data represent the mean ± s.e.m, for 3 independent experiments, n=12 Ebf1+/+Prx1Cre mice and n=10 Ebf1fl/flPrx1Cre mice. SP-spleen, PB-peripheral blood. c, Frequency of neutrophils (CD11b+CD11c–Ly6G+), monocytes (CD11b+CD11c–Ly6Chi) and macrophages (MHC-II+CD11b+F4/80+) in the BM. Data represent the mean ± s.e.m, n=10 Ebf1+/+Prx1Cre mice and n=12 Ebf1fl/flPrx1Cre mice (*P=0.03, **P=0.001). d, Absolute numbers of B cells (CD19+B220+) with n=14 mice per group, T cells (CD3+) with n=16 Ebf1+/+Prx1Cre mice and n=15 Ebf1fl/flPrx1Cre mice and erythroid cells (Ter119+) with n=15 Ebf1+/+Prx1Cre mice and n=14 Ebf1fl/flPrx1Cre mice, in the BM. e, Quantification of early B cell subsets: common lymphoid progenitors-CLP (Lin–Sca1loc-kitloIL-7Rα+CD135+) with n=10 mice per group, pro-B cells (B220+CD43+), pre-B cells (B220loCD43–) and rec-B cells (B220hiCD43–) with n=14 mice per group, in the BM. Statistical analysis in a-e was performed using two-tailed unpaired Student’s t-test.
Extended Data Fig. 4 LepRCre and NG2Cre-ERTM dependent deletion of Ebf1.
a, Total BM cellularity. Data represent the mean ± s.e.m, n=22 Ebf1+/+LepRCre mice and n=20 Ebf1fl/flLepRCre mice. b, Absolute numbers of MPP3 (CD135–CD48+CD150–LSK) and MPP4 (CD135+ CD150–LSK) in the BM. Data represent the mean ± s.e.m, n=17 Ebf1+/+LepRCre mice and n=20 Ebf1fl/flLepRCre mice (*P=0.02). c, Numbers of CLPs (Lin–Sca1loc-kitloIL-7Rα+CD135+) in the BM. Data represent the mean ± s.e.m, n=12 Ebf1+/+LepRCre mice and n=13 Ebf1fl/flLepRCre mice (*P=0.03). d, Frequency of neutrophils (CD11b+CD11c–Ly6G+) (*P=0.03), monocytes (CD11b+CD11c–Ly6Chi) and macrophages (MHC-II+CD11b+F4/80+) (*P=0.01) in the BM. Data represent the mean ± s.e.m, n=8 mice per group. e, Absolute numbers of T cells (CD3+) and erythroid cells (Ter119+) in the BM. Data represent the mean ± s.e.m, n=22 (T cells) and n=19 (erythroid) Ebf1+/+LepRCre mice and n=19 Ebf1fl/flLepRCre mice. f, Numbers of B lineage cells (CD19+B220+) with n=13 mice per group, including pro-B cells (B220+CD43+), pre-B cells (B220loCD43–) and rec-B cells (B220hiCD43–) with n=12 mice per grop, in the BM of Ebf1fl/flLepRCre and Ebf1+/+LepRCre mice. Data represent the mean ± s.e.m (**p=0.03). g, Total BM cellularity. Data represent the mean ± s.e.m, n=16 Ebf1+/+ NG2Cre-ERTM mice and n=17 Ebf1fl/flNG2Cre-ERTM mice. h, Absolute numbers of MPP3 (CD135–CD48+CD150–LSK) and MPP4 (CD135+ CD150–LSK) in the BM. Data represent the mean ± s.e.m, n=16 mice per group. I, Numbers of B cells (CD19+B220+), T cells (CD3+) and erythroid cells (Ter119+) in the BM of Ebf1fl/flNG2Cre-ERTM and Ebf1+/+NG2Cre-ERTM mice. Data represent the mean ± s.e.m, n=12 mice per group. Statistical analysis in a-i was performed using two-tailed unpaired Student’s t-test.
Extended Data Fig. 5 Chromatin accessibility in CAR and PαS cells from Ebf1+/+ and Ebf1–/– mice.
a, Frequency and absolute number of HSCs in the spleens of Ebf1+/+Prx1Cre and Ebf1fl/flPrx1Cre mice after G-CSF treatment. G-CSF was administered subcutaneously once a day for six consecutive days. Mice were analyzed on day 7 from the start of G-CSF treatment. Data represent the mean ± s.e.m, for 3 independent experiments, n=7 mice per group (*p=0.03, two-tailed unpaired Student’s t-test). b, Venn diagram showing the numbers of identified ATAC peaks in CAR and PαS cells from Ebf1+/+ and Ebf1–/– mice. c, Percentage of ATAC peaks in different genomic regions in CAR and PαS cells. d, Motifs identified in clusters presented in Fig.4g. e, Venn diagrams comparing differentially expressed genes with differential ATAC peaks containing EBF motif in CAR and PαS cells from Ebf1+/+ and Ebf1–/– mice. Upregulated genes are in red and downregulated genes are in green. All genes marked by black circle in WT only cluster (blue box) show loss of ATAC signal within EBF motif in Ebf1–/– (KO) cells. Genes marked by black circle in common cluster show loss of EBF1-specific footprint in Ebf1-/- cells. f, Digital genomic footprinting analysis showing average normalized Tn5 insertion profiles around EBF footprinted motifs in merged ATAC peaks. Insertions on the forward and reverse strands are indicated in red and blue, respectively.
Extended Data Fig. 6. Transplantation of HSCs from Ebf1-deficient niche to wild-type recipients.
a, Percent of chimerism in peripheral blood (PB) during the course of experiment. Percentage of CD45.2+ donor-derived cells in PB within myeloid cells (Mac1+) b, B cells (CD19+) c, and T cells (CD3+) d over time. Data represent the mean ± s.e.m with n=18 recipients of HSCs from Ebf1+/+Prx1Cre mice and n=18 recipients of HSCs from Ebf1fl/flPrx1Cre mice in the primary bone marrow transplantation and n=20 recipients of HSCs from Ebf1+/+Prx1Cre mice and n=29 recipients of HSCs from Ebf1fl/flPrx1Cre mice in the secondary bone marrow transplantation.
Extended Data Fig. 7 Niche-dependent changes in HSCs chromatin accessibility and transcriptome.
a, ATAC signal ±3 Kb around the center of the peak. b, Motifs identified in clusters with lost and gained accessibility in HSCs from Ebf1+/+Prx1Cre and Ebf1fl/flPrx1Cre mice. c, Heatmaps showing footprinted motif co-occurrence enrichment clustering. d, GSEA plot showing enrichments of myeloid cells differentiation signature in HSCs derived from wild-type niche. The Normalized Enrichment Score (NES) was computed using a Kolmogorov–Smirnov statistic. Significance was assessed using a two-tailed t-test comparing to a null distribution computed via 1000 gene-set permutations. Correction for multiple testing was performed using FDR adjustment, n=183. e, Representative tracks showing differential merged ATAC signal in HSCs from Ebf1+/+Prx1Cre and Ebf1fl/flPrx1Cre mice correlated with genes differentially expressed between cluster 2 and 3 (Fig 6e). Region on chromosome 8 with Pcm1 and Asah1 serves as a control without differential ATAC signal. The scale in the y-axis represents RPKM.
Extended Data Fig. 8 Niche-dependent changes in HSCs composition.
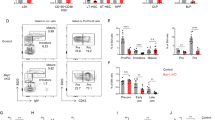
a, t-SNE maps highlighting normalized expression of Mki67 in sequenced HSCs. b, Representative tracks showing differential merged ATAC signal in HSCs from Ebf1+/+Prx1Cre and Ebf1fl/flPrx1Cre mice correlated with genes differentially expressed between cluster 2 and 3 (Fig. 7e). Region on chromosome 1 with Fbxo36 and Cab39 serves as a control without differential ATAC signal. The scale in the y-axis represents RPKM. c, t-SNE map showing clusters of cells with similar transcriptomes within LSK (Lin–Sca-1+c-kit+) populations, derived by RaceID3 algorithm. Data represent 2844 single-sorted wild-type and knockout cells from 2 independent experiments (n=5 mice for Ebf1+/+Prx1Cre group and n=6 mice for Ebf1fl/flPrx1Cre group). d, t-SNE map displaying clusters with statistically significant enrichment of wild-type cells (cluster 6) or Ebf1-deficient cells (cluster 3 and 10) within LSK population. Data represent 2844 single-sorted wild-type and knockout cells from 2 independent experiments (n=5 mice for Ebf1+/+Prx1Cre group and n=6 mice for Ebf1fl/flPrx1Cre group). Fisher’s exact test was used for statistics (exact P values and cell numbers for each cluster are provided in Supplementary Table 5). e, t-SNE maps showing normalized expression of Mpo in sequenced LSKs. f, t-SNE maps showing normalized expression of Ctsg in sequenced LSKs. g, MA plot showing differential gene expression between cluster 10 and cluster 11 from scRNA-seq analysis of LSKs. Genes upregulated in cluster 11 are shown in red and downregulated genes are shown in blue.
Supplementary information
Supplementary Information
Supplementary Methods.
Supplementary Table
Supplementary Tables 1–6.
Source data
Source Data Extended Data Fig. 1
Unprocessed Western Blots
Rights and permissions
About this article
Cite this article
Derecka, M., Herman, J.S., Cauchy, P. et al. EBF1-deficient bone marrow stroma elicits persistent changes in HSC potential. Nat Immunol 21, 261–273 (2020). https://doi.org/10.1038/s41590-020-0595-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0595-7
This article is cited by
-
Single-cell RNA sequencing of human non-hematopoietic bone marrow cells reveals a unique set of inter-species conserved biomarkers for native mesenchymal stromal cells
Stem Cell Research & Therapy (2023)
-
Age-Progressive and Gender-Dependent Bone Phenotype in Mice Lacking Both Ebf1 and Ebf2 in Prrx1-Expressing Mesenchymal Cells
Calcified Tissue International (2022)