Abstract
CD8+ T cell exhaustion dampens antitumor immunity. Although several transcription factors have been identified that regulate T cell exhaustion, the molecular mechanisms by which CD8+ T cells are triggered to enter an exhausted state remain unclear. Here, we show that interleukin-2 (IL-2) acts as an environmental cue to induce CD8+ T cell exhaustion within tumor microenvironments. We find that a continuously high level of IL-2 leads to the persistent activation of STAT5 in CD8+ T cells, which in turn induces strong expression of tryptophan hydroxylase 1, thus catalyzing the conversion to tryptophan to 5-hydroxytryptophan (5-HTP). 5-HTP subsequently activates AhR nuclear translocation, causing a coordinated upregulation of inhibitory receptors and downregulation of cytokine and effector-molecule production, thereby rendering T cells dysfunctional in the tumor microenvironment. This molecular pathway is not only present in mouse tumor models but is also observed in people with cancer, identifying IL-2 as a novel inducer of T cell exhaustion.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The ChIP–seq data reported in this paper have been deposited in the Genome Sequence Archive in National Genomics Data Center, Beijing Institute of Genomics (China National Center for Bioinformation), Chinese Academy of Sciences, under accession number CRA003170. The TCGA data reported in this paper derive from UCSC xena (https://xenabrowser.net/datapages/). Source data are provided with this paper.
References
Borghaei, H. et al. Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N. Engl. J. Med. 373, 1627–1639 (2015).
Neelapu, S. S. et al. Axicabtagene ciloleucel CAR T-cell therapy in refractory large B-cell lymphoma. N. Engl. J. Med. 377, 2531–2544 (2017).
Zaretsky, J. M. et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. N. Engl. J. Med. 375, 819–829 (2016).
Majzner, R. G. & Mackall, C. L. Tumor antigen escape from CAR T-cell therapy. Cancer Discov. 8, 1219–1226 (2018).
Fourcade, J. et al. Upregulation of Tim-3 and PD-1 expression is associated with tumor antigen-specific CD8+ T cell dysfunction in melanoma patients. J. Exp. Med. 207, 2175–2186 (2010).
Baitsch, L. et al. Exhaustion of tumor-specific CD8+ T cells in metastases from melanoma patients. J. Clin. Invest. 121, 2350–2360 (2011).
Ma, X. et al. Cholesterol induces CD8+ T cell exhaustion in the tumor microenvironment. Cell Metab. 30, 143–156.e5 (2019).
Scott, A. C. et al. TOX is a critical regulator of tumour-specific T cell differentiation. Nature 571, 270–274 (2019).
Alfei, F. et al. TOX reinforces the phenotype and longevity of exhausted T cells in chronic viral infection. Nature 571, 265–269 (2019).
Zhang, L. et al. Lineage tracking reveals dynamic relationships of T cells in colorectal cancer. Nature 564, 268–272 (2018).
Zheng, C. et al. Landscape of infiltrating T cells in liver cancer revealed by single-cell sequencing. Cell 169, 1342–1356.e16 (2017).
Tang, Z., Kang, B., Li, C., Chen, T. & Zhang, Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 47, W556–W560 (2019).
Ehrlich, A. K. et al. AhR activation increases IL-2 production by alloreactive CD4+ T cells initiating the differentiation of mucosal-homing Tim3+ Lag3+ Tr1 cells. Eur. J. Immunol. 47, 1989–2001 (2017).
Mujib, S. et al. Antigen-independent induction of Tim-3 expression on human T cells by the common γ-chain cytokines IL-2, IL-7, IL-15, and IL-21 is associated with proliferation and is dependent on the phosphoinositide 3-kinase pathway. J. Immunol. 188, 3745–3756 (2012).
Zhang, Z. N. et al. Elevation of Tim-3 and PD-1 expression on T cells appears early in HIV infection, and differential Tim-3 and PD-1 expression patterns can be induced by common γ-chain cytokines. Biomed. Res. Int. 2015, 916936 (2015).
Beltra, J. C. et al. IL2Rβ-dependent signals drive terminal exhaustion and suppress memory development during chronic viral infection. Proc. Natl Acad. Sci. USA 113, E5444–E5453 (2016).
Ross, S. H. & Cantrell, D. A. Signaling and function of interleukin-2 in T lymphocytes. Annu. Rev. Immunol. 36, 411–433 (2018).
Malek, T. R. The biology of interleukin-2. Annu. Rev. Immunol. 26, 453–479 (2008).
Opitz, C. A. et al. An endogenous tumour-promoting ligand of the human aryl hydrocarbon receptor. Nature 478, 197–203 (2011).
Liu, Y. et al. Tumor-repopulating cells induce PD-1 expression in CD8+ T cells by transferring kynurenine and AhR activation. Cancer Cell 33, 480–494.e7 (2018).
Gutierrez-Vazquez, C. & Quintana, F. J. Regulation of the immune response by the aryl hydrocarbon receptor. Immunity 48, 19–33 (2018).
Bittinger, M. A., Nguyen, L. P. & Bradfield, C. A. Aspartate aminotransferase generates proagonists of the aryl hydrocarbon receptor. Mol. Pharmacol. 64, 550–556 (2003).
Rothhammer, V. & Quintana, F. J. The aryl hydrocarbon receptor: an environmental sensor integrating immune responses in health and disease. Nat. Rev. Immunol. 19, 184–197 (2019).
Xing, X. et al. SUMOylation of AhR modulates its activity and stability through inhibiting its ubiquitination. J. Cell. Physiol. 227, 3812–3819 (2012).
Wojciechowski, W. et al. Cytokine-producing effector B cells regulate type 2 immunity to H. polygyrus. Immunity 30, 421–433 (2009).
Granucci, F. et al. Inducible IL-2 production by dendritic cells revealed by global gene expression analysis. Nat. Immunol. 2, 882–888 (2001).
Liu, C. et al. Neuropilin-1 is a T cell memory checkpoint limiting long-term antitumor immunity. Nat. Immunol. 21, 1010–1021 (2020).
Martinez, G. J. et al. The transcription factor NFAT promotes exhaustion of activated CD8+ T cells. Immunity 42, 265–278 (2015).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8+ T cell exhaustion. Nature 571, 211–218 (2019).
Spranger, S. et al. Up-regulation of PD-L1, IDO, and Tregs in the melanoma tumor microenvironment is driven by CD8+ T cells. Sci. Transl. Med. 5, 200ra116 (2013).
Kalia, V. & Sarkar, S. Regulation of effector and memory CD8 T cell differentiation by IL-2-A balancing act. Front. Immunol. 9, 2987 (2018).
Smith, K. A. Interleukin-2: inception, impact, and implications. Science 240, 1169–1176 (1988).
Cheng, L. E., Ohlén, C., Nelson, B. H. & Greenberg, P. D. Enhanced signaling through the IL-2 receptor in CD8+ T cells regulated by antigen recognition results in preferential proliferation and expansion of responding CD8+ T cells rather than promotion of cell death. Proc. Natl Acad. Sci. USA 99, 3001–3006 (2002).
Shevach, E. M. Application of IL-2 therapy to target T regulatory cell function. Trends Immunol. 33, 626–632 (2012).
Van Parijs, L. & Abbas, A. K. Homeostasis and self-tolerance in the immune system: turning lymphocytes off. Science 280, 243–248 (1998).
Liao, W., Lin, J. X. & Leonard, W. J. Interleukin-2 at the crossroads of effector responses, tolerance, and immunotherapy. Immunity 38, 13–25 (2013).
Rosenberg, S. A. Raising the bar: the curative potential of human cancer immunotherapy. Sci. Transl. Med. 4, 127ps128 (2012).
Subramanian Vignesh, K., Landero Figueroa, J. A., Porollo, A., Caruso, J. A. & Deepe, G. S. Jr. Granulocyte macrophage-colony stimulating factor induced Zn sequestration enhances macrophage superoxide and limits intracellular pathogen survival. Immunity 39, 697–710 (2013).
Lin, J. X. et al. The role of shared receptor motifs and common Stat proteins in the generation of cytokine pleiotropy and redundancy by IL-2, IL-4, IL-7, IL-13, and IL-15. Immunity 2, 331–339 (1995).
Man, K. et al. Transcription factor IRF4 promotes CD8+ T cell exhaustion and limits the development of memory-like T cells during chronic infection. Immunity 47, 1129–1141.e5 (2017).
Ames, R. Y., Ting, L.-M., Gendlina, I., Kim, K. & Macian, F. The transcription factor NFAT1 participates in the induction of CD4+ T cell functional exhaustion during Plasmodium yoelii infection. Infect. Immun. 85, e00364–00317 (2017).
Doering, T. A. et al. Network analysis reveals centrally connected genes and pathways involved in CD8+ T cell exhaustion versus memory. Immunity 37, 1130–1144 (2012).
Shin, H. et al. A role for the transcriptional repressor Blimp-1 in CD8+ T cell exhaustion during chronic viral infection. Immunity 31, 309–320 (2009).
Welsh, R. M. Blimp hovers over T cell immunity. Immunity 31, 178–180 (2009).
Zaid, A. et al. Persistence of skin-resident memory T cells within an epidermal niche. Proc. Natl Acad. Sci. USA 111, 5307–5312 (2014).
Murray, I. A., Patterson, A. D. & Perdew, G. H. Aryl hydrocarbon receptor ligands in cancer: friend and foe. Nat. Rev. Cancer 14, 801–814 (2014).
Boyman, O. & Sprent, J. The role of interleukin-2 during homeostasis and activation of the immune system. Nat. Rev. Immunol. 12, 180–190 (2012).
Herr, F. et al. IL-2 phosphorylates STAT5 to drive IFN-γ production and activation of human dendritic cells. J. Immunol. 192, 5660–5670 (2014).
Wilhelm, C. et al. An IL-9 fate reporter demonstrates the induction of an innate IL-9 response in lung inflammation. Nat. Immunol. 12, 1071–1077 (2011).
Tanaka, Y., Nagai, Y., Okumura, M., Greene, M. I. & Kambayashi, T. PRMT5 is required for T cell survival and proliferation by maintaining cytokine signaling. Front Immunol. 11, 621 (2020).
Acknowledgements
This work was supported by National Natural Science Foundation of China (81788101, 81773062, 91942314) and Chinese Academy of Medical Sciences (CAMS) Initiative for Innovative Medicine (CAMS-I2M) 2016-I2M-1-007.
Author information
Authors and Affiliations
Contributions
B.H. conceived of the project. Y.L., N.Z., L.Z., J.W., Y.Z., T.Z., Y.F., Y.G., X.L., J.L., Z.W., J.X., Huafeng Zhang, J.M., K.T., Y.F., F.C. and Haizeng Zhang performed the experiments. Y.L., N.Z., L.Z., J.W., Y.X., J.D., C.Z., B.D., Y.Z., Q.G. and P.Y. developed methodology. Y.L., B.H., F.X.-F.Q., N.Z. and L.Z. performed data analysis. B.H. and Y.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Immunology thanks Yang-Xin Fu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 IL-2 signaling induces CD8+ T cells exhaustion.
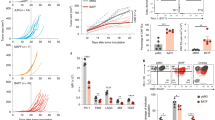
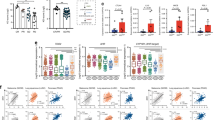
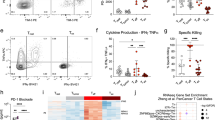
a, TCGA database RNA-Seq analysis of correlation between the expression of IL-2 and exhausted T cell signature genes in breast (n = 195) and lung (n = 61) paracancerous tissues, uterine corpus endometrial carcinoma (n = 176), and colon paracancerous tissues (n = 43); or IL-2R and exhausted T cell signature genes in colon cancer (n = 590), breast cancer (n = 1080), lung squamous cell carcinoma (n = 482) or lung adenocarcinoma (n = 508) patients. GEO database RNA-Seq analysis of correlation between the expression of IL-2 and exhausted T cell signature genes in liver normal tissue from chronic hepatitis B patients (n = 124, GSE84044). b, Activated OVA-CTLs were treated with IL-2, IL-2 + anti-CD25, IL-2 + anti-CD122 or IL-2 + anti-CD25 + anti-CD122 neutralizing antibodies for 72 hr. Expression profiles of IRs and cytokines were determined by flow cytometry. c, Activated CD45.1+ gp-100-CTLs were adoptively transferred into C57BL/6 J mice bearing with 5 × 5 mm B16 melanoma. At the same time, these mice were intraperitoneally injected with PBS, IL-2, anti-IL-2 neutralizing antibody or anti-CD25 + anti-CD122 neutralizing antibody every two days for 3 times. At day 7, CD45.1+ transferred cells were isolated from the tumors for flow cytometric analysis of the expression of IRs (PD-1, LAG3, TIM3) and cytokines. d, The same as (c), except that CD45.1+ OVA-CTLs were transferred into mice. e-g, Activated CTLs were treated with IL-2 for the indicated time periods. Expression profiles of IRs (e), viable cell number (f) and the level of cytokines from the supernatants (g) were determined by flow cytometry or ELISA. h-j, Splenocytes from OVA-OT1 mice were stimulated by OVA peptides for 48 hr, and then treated with IL-2 for indicated times periods. The expression of IRs (h), the level of cytokines (i) and viable cells number (j) were determined by flow cytometry. k,l, CTLs were treated with different doses of IL-2 for 48 hr (k) or IL-2 for indicated time periods (l). The expression of IRs was determined by flow cytometry. In c,d, n = 5 mice; b,e-l, n = 3 independent experiments. Two- tailed Student’s t test (f) or by one-way ANOVA Bonferroni’s test (b-e, g-l), or Pearson’s correlation test (a). The data represent mean ± SD.
Extended Data Fig. 2 The Tph1-5HTP pathway regulates CD8+ T cell exhaustion induced by IL-2.
a, Activated CTLs were treated with STAT5 inhibitor (Stat5in), fludarabine, Stattic, LY294002, SCH772984, SP00125 or SB203580 for 48 hr. The expression of IRs was determined by flow cytometry. b,h The knockdown efficiency of STAT5 (b) or TPH1 (h) was confirmed by western blot. c,d, The resting or activated CTLs were treated with IL-2 for 48 hr (c). Or, activated CTLs were treated with IL-2 or IL-2 + Stat5in for 48 hr (d). Tph1 mRNA was determined by real-time PCR. e, The expression of Tph1 from resting or activated Stat5 shRNAs- CD8+ T cells treated with IL-2 or IL-2 + Stat5in for 48 hr. f, The same as (c), except that activated scramble or Stat5 shRNAs-CTLs were used. g, Activated CTLs were treated with IL-2, Stat5in or IL-2 + Stat5in for 48 hr. The level of 5HTP was measured by HPLC-MS. i-k, Activated CTLs were treated with different doses of 5HTP as indicated for 48 hr (i,j) or 5HTP for the indicated time periods (k). Flow cytometric analysis of the expression of IRs, TNF or IFN-γ. l,m, The same as (i-k), except that resting CD8+ T cells were used. In a-m, n = 3 independent experiments. One-way ANOVA Bonferroni’s test (a, c, d, f, g, i-m). The data represent mean ± SD.
Extended Data Fig. 3 The Kyn-AhR pathway regulates IL-2 induced CD8+ T cells exhaustion.
a,b, Activated CTLs were treated with 5HTP or different doses of carbidopa (a), serotonin or melatonin (b) for 48 hr. The expression of IRs was determined by flow cytometry. c, Resting or activated CTLs were treated with 5HTP for 48 hr. The mRNA expression of Cyp1a1 and Cyp1b1 was determined by real-time PCR. d,e, Activated murine CTLs were treated with StemRegenin 1 (SR1), CH223191 (CH), 5HTP, 5HTP/SR1 or 5HTP/CH for 48 hr. Flow cytometric analysis of the expression of IRs (d), or TNF and IFN-γ (e). f, The same as (d,e), except that CTLs were treated with IL-2, IL-2 + SR1 or IL-2 + CH. g, NIH3T3 cells expressing Pdcd-1, Lag3, Havcr2 or Entpd1 promoter-luciferase reporter PGL3 were co-transfected with AhR-plasmid. Cells were treated with 5HTP for 48 hr and analyzed by luciferase assay. h, The same as g, except that cells expressed Pdcd-1, Havcr2 or Entpd1 promoter mutant-luciferase reporter PGL3. i, The same as f, except that cells were treated with Kyn or tryptophan (Trp) for 48 hr. Flow cytometric analysis of the expression of IRs, TNF and IFN-γ. In a-i, n = 3 independent experiments. One-way ANOVA Bonferroni’s test (a, c, d, e, f, g, h, i). The data represent mean ± SD.
Extended Data Fig. 4 IL-2 treatment stabilizes AhR.
a, Activated human CD8+ T cells were treated with IL-2 or IL-2 antibody for 48 hr. The mRNA expression of AhR was measured by real-time PCR. b, Activated CTLs were treated with IL-2 in the presence of cycloheximide (CHX) (2 μg/mL) for the indicated time periods. The level of AhR was determined by western blot analysis. c,The same as (b), except that stably overexpressing HA-AhR human CD8+ T cells were used and the level of AhR was determined by western blot with anti-HA antibody. d, Activated CTLs were treated with IL- 2, anti-IL-2 antibody or IL-2 + anti-IL-2 antibody for 48 hr. AhR level was determined by western blot. e,I, The same as d, except that cell lysates were immunoprecipitated with anti-AhR antibody and immunoblotted with an anti-ubiquitination I, SUMO1 or SUMO2/3 (i) antibody. f,j, Activated human CD8+ T cells stably overexpressing HA-AhR were transfected with a Flag-UbG76V (f) or Flag-SUMO1 plasmid (j). Then, cells were treated with IL- 2, anti-IL-2 antibody or IL-2 + anti-IL-2 antibody. Cell lysates were immunoprecipitated with anti-HA antibody and immunoblotted with an anti-ubiquitination or Flag antibody. g,h, Anti-CD3 antibody-activated human (g) or mouse (h) CD8+ T cells were treated with IL-2 in the presence of MG132 (10 μM) for the indicated time periods. Cells were lysed to perform western blot analysis with an anti-AhR antibody. k, Activated HA-AhR-human CD8+ T cells were transfected with Flag-SUMO1 plasmid. Cells were stained with anti-Flag and anti-HA antibodies and observed under a confocal microscope. Bar, 10 μm. In a-k, n = 3 independent experiments. The data represent mean ± SD.
Extended Data Fig. 5 The Tph1-5HTP-AhR pathway regulates IL-2-induced CD8+ T cells exhaustion in vivo.
a, The mRNA expression of Tph1 in CD8+ T cells from tumor tissues of C57BL/6 J mice with 3 × 3 mm or 8 × 8 mm MC38 tumors. b, Correlation between Tph1 expression and IL-2 levels in MC38 tumors. c, Activated CD45.1+ OVA-CTLs were transferred into C57BL/6 J mice bearing with OVA-B16 melanoma, and then mice were intratumorally injected with 5HTP. CD45.1+ cells were isolated from the tumors for flow cytometric analysis of the expression of IRs, or TNF and IFN-γ. d, C57BL/6 J mice bearing MC38 tumor were intratumorally injected with Stat5in, fenchlonine. CD8+ T cells were isolated from the tumor for flow-cytometric analysis of the expression of IRs, or TNF and IFN-γ. e, The same as c, except that mice were treated with Stat5in or fenchlonine. f,g, Activated Stat5 shRNAs (f) or Tph1 shRNAs-CD45.1+ gp-100-CTLs (g) were transferred into CD45.2+ C57BL/6 J mice with B16 melanoma. At day 7, CD45.1+ T cells were isolated from tumor tissues and the expression of IRs, TNF and IFN-γ were determined by flow cytometry. h-j, The same as (f,g), except that cells were transferred into mice every five days for 3 times. Some mice were intraperitoneally injected with anti-CD25 + anti-CD122 antibody. Tumor growth and mouse survival were analyzed. k, The same as (d), except that mice were intratumorally injected with CH223191 or SR1. l, The same as c, except that mice were intratumorally injected with CH223191or SR1. m, C57BL/6 J mice with 3 × 3 mm MC38 tumor were injected with IL-2, SR1, Stat5in, IL-2 + SR1 or IL-2 + Stat5in every two days for 5 times. Tumor growth was analyzed. n, The same as (m), except that mice with 8 × 8 mm MC38 tumors were treated with IL-15. o,p, Activated CTLs were treated with IL-2 or IL-15 for 48 hr (o) or 72 hr (p). The expression of Tph1 (o) or IRs (p) was determined by real-time PCR or flow cytometry, respectively. In a, d-g, k, l, n = 5 mice; b, n = 15 mice; c, n = 6 mice; h-j, n = 10 mice; m, n, n = 8 mice; o, p, n = 3 independent experiments. Two-tailed Student’s t test (a, c, n), Pearson’s correlation test (b), one-way ANOVA Bonferroni’s test (d-g, h (left), i (left), j-m, o, p), or Log-rank survival analysis (h (right), i (right)). The data represent mean ± SD (a, c-g, k, l, o, p) or mean ± SEM (h-j, m, n).
Extended Data Fig. 6 IL-2 is released by CD45+CD3+ T cells to induce CD8+ T cells exhaustion in the tumor microenvironment.
a, IL-2 mRNA expression in murine CD8+ T cells (mCD8+ T cells) or the PB of healthy volunteer (hCD8+ T cells), MC38, 4T1, B16, HCT15, HCT116, MCF-7 or MP-1 human primary melanoma cells. b, IL-2 mRNA expression in CD45− and CD45+ cells from 5 × 5 mm MC38 tumor tissues. c, The CD11c+ IL-2+, B220+ IL-2+ or CD11b+ F4/80+ IL-2+ cells from 3 × 3 mm or 8 × 8 mm MC38 tumor tissues were determined by flow cytometry. d,e The CD3+ IL-2+ CD4+ or CD3+ IL-2+ CD8+ cells from 3 × 3 mm or 8 × 8 mm B16 melanoma tumor tissues (d) or H22 ascites (e) were determined by flow cytometry. f,g, C57BL/6 J mice with 3 × 3 mm or 8 × 8 mm MC38 tumors were treated with IL-2 or IL2 + anti-IL-2Ra antibody. The CD3+, CD3+CD8+ (f), or PD1+TCF1−TIM3+ (g) T cells in the tumors were analyzed by flow cytometry. h, Activated OVA-CTLs were treated with IL-2 for indicated times periods. The PD1+TCF1−TIM3+ T cells were analyzed by flow cytometry. i,j, The same as (f,g), except that mice were i.p. injected with anti-IL-2 (i), anti- anti-CD4 or CTLA-4 (j) neutralizing antibody. Tumor growth was analyzed. k-m, C57BL/6 J mice with 8 × 8 mm B16 melanoma were intraperitoneally injected with anti-CD4 neutralizing antibody. CD8+ T cells were isolated from the tumor for flow-cytometric analysis of the expression of IRs (k), or TNF and IFN-γ (l). Tumor growth and mouse survival was analyzed (m). n, The same as (k,l), except that BALB/c mice with H22 ascites were used. o, Activated murine CD4+ and CD8+ T cells were treated with IL-2 for 72 hr. The expression of IRs was determined by flow cytometry. In b-g, k, l, n, n = 5 mice; i, j, n = 8 mice; m, n = 7 mice; a, h, o, n = 3 independent experiments. Two-tailed Student’s t test (b, d, e, g, h, k, l, m, n), one-way ANOVA Bonferroni’s test (f, i, j, o), or Log-rank survival analysis (m (down)). The data represent mean ± SD (a-h, k, l, n, o) or mean ± SEM (i, j, m).
Extended Data Fig. 7 IL-2 induces CD8+ T cells exhaustion by activating TPH1-5HTP-AhR pathway in patients.
a, IL-2 mRNA expression in CD45+ and CD45− cells from colon or breast cancer tissues of patients (n = 5). b, The CD3+ CD8+ IL-2+ or CD3+ CD4+ IL-2+ cells from the fresh tissues of colon cancer patients (n = 5) was determined by flow cytometry. c, Resting CD8+ T cells from the PB of colon or breast cancer patients (n = 6) were treated with IL-2 for 72 hr. The expression of IRs and cytokines was determined by flow cytometry. d, The same as (c), except that resting or activated T cells from healthy donors were used. e, Resting or activated CD8+ T cells from PB of colon or lung cancer patients (n = 15 scopes from 3 patients) were treated with IL-2 for 48 hr. Cells were stained with anti-TPH1 antibody and observed by confocal microscopy. Bar, 10 μm. f, The same as (c), except that activated TPH1shRNAs- CD8+ T cells from healthy volunteers were used. g, Activated CD8+ T cells from the PB of healthy volunteers or colon cancer patients were treated with IL-2, IL-2 + Stat5in or IL-2 + fenchlonine for 72 hr. The expression of IRs and cytokines were determined by flow cytometry. h, The same as (c), except that STAT5 shRNAs-CD8+ T cells from healthy volunteers or colon cancer patients were used. i, Activated CD8+ T cells from the PB of colon or lung cancer patients were treated with IL-2 for 48 hr, or CD3+CD8+ T cells were sorted from peripheral blood (n = 50 scope from 10 patients) or tumor tissues (n = 30 scope from 6 patients) of colon or lung cancer patients. These cells were stained with anti-AhR antibody and observed by confocal microscopy. Bar, 10 μm. j, The same as (g), except CD8+ T cells from colon cancer patients were treated with IL-2, IL-2 + SR1 or IL-2 + CH223191. k, The same as (g), except that CD8+ T cells were from healthy donors or lung or breast cancer patients. In d, f-h, n = 3 independent experiments; j, k, n = 5. Two- tailed Student’s t test (a, b, c, d, i), one-way ANOVA Bonferroni’s test (e-h, j, k). The data represent mean ± SD.
Supplementary information
Supplementary Tables 1–7
Clinical information of cancer patients, reagent information, antibody information, primer sequences information and CHIP-seq information.
Source data
Source Data Fig. 1
Statistical source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Statistical source data
Source Data Fig. 3
Unprocessed fluorescent image
Source Data Fig. 4
Statistical source data
Source Data Fig. 4
Unprocessed western blot
Source Data Fig. 5
Statistical source data
Source Data Fig. 6
Statistical source data
Source Data Fig. 7
Statistical source data
Source Data Extended Data Fig. 1
Statistical source data
Source Data Extended Data Fig. 2
Statistical source data
Source Data Extended Data Fig. 2
Unprocessed western blot
Source Data Extended Data Fig. 3
Statistical source data
Source Data Extended Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 4
Unprocessed western blot and unprocessed fluorescent image
Source Data Extended Data Fig. 5
Statistical source data
Source Data Extended Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 7
Statistical source data
Source Data Extended Data Fig. 7
Unprocessed fluorescent image
Rights and permissions
About this article
Cite this article
Liu, Y., Zhou, N., Zhou, L. et al. IL-2 regulates tumor-reactive CD8+ T cell exhaustion by activating the aryl hydrocarbon receptor. Nat Immunol 22, 358–369 (2021). https://doi.org/10.1038/s41590-020-00850-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-00850-9
This article is cited by
-
Uremic toxins mediate kidney diseases: the role of aryl hydrocarbon receptor
Cellular & Molecular Biology Letters (2024)
-
Cellular metabolism regulates the differentiation and function of T-cell subsets
Cellular & Molecular Immunology (2024)
-
Tumour-infiltrating lymphocyte therapy for patients with advanced-stage melanoma
Nature Reviews Clinical Oncology (2024)
-
Stratifying ICIs-responsive tumor microenvironment in HCC: from parsing out immune-hypoxic crosstalk to clinically applicable MRI-radiomics models
British Journal of Cancer (2024)
-
Importance of CD8 Tex cell-associated gene signatures in the prognosis and immunology of osteosarcoma
Scientific Reports (2024)