Abstract
The present vaccine against influenza virus has the inevitable risk of antigenic discordance between the vaccine and the circulating strains, which diminishes vaccine efficacy. This necessitates new approaches that provide broader protection against influenza. Here we designed a vaccine using the hypervariable receptor-binding domain (RBD) of viral hemagglutinin displayed on a nanoparticle (np) able to elicit antibody responses that neutralize H1N1 influenza viruses spanning over 90 years. Co-display of RBDs from multiple strains across time, so that the adjacent RBDs are heterotypic, provides an avidity advantage to cross-reactive B cells. Immunization with the mosaic RBD–np elicited broader antibody responses than those induced by an admixture of nanoparticles encompassing the same set of RBDs as separate homotypic arrays. Furthermore, we identified a broadly neutralizing monoclonal antibody in a mouse immunized with mosaic RBD–np. The mosaic antigen array signifies a unique approach that subverts monotypic immunodominance and allows otherwise subdominant cross-reactive B cell responses to emerge.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All data needed to evaluate the conclusions in the paper are present in the paper and/or the Supplementary information. Coordinates for 441D6 Fab structure and cryo-electron microscopy density map have been deposited in the Protein Data Bank (PDB) under accession code 5TR8 and electron microscopy Data Bank (EMDB) under accession code EMD-7021. Hemagglutinin RBD–np and antibody sequences are deposited in the GenBank under accession codes MK273069–MK273076 and MK283705–MK283710. Requests for materials should be addressed to M.K. and B.S.G.
Change history
12 April 2019
In the version of this article initially published, the labels (50 Å) above the scale bars in Fig. 1b were incorrect. The correct size is 50 nm. The error has been corrected in the HTML and PDF versions of the article.
References
Morens, D. M. & Taubenberger, J. K. Historical thoughts on influenza viral ecosystems, or behold a pale horse, dead dogs, failing fowl, and sick swine. Influenza Other Respir. Viruses 4, 327–337 (2010).
Bedford, T. et al. Integrating influenza antigenic dynamics with molecular evolution. eLife 3, e01914 (2014).
Koel, B. F. et al. Substitutions near the receptor binding site determine major antigenic change during influenza virus evolution. Science 342, 976–979 (2013).
Smith, D. J. et al. Mapping the antigenic and genetic evolution of influenza virus. Science 305, 371–376 (2004).
Webster, R. G., Laver, W. G., Air, G. M. & Schild, G. C. Molecular mechanisms of variation in influenza viruses. Nature 296, 115–121 (1982).
Air, G. M. Influenza virus antigenicity and broadly neutralizing epitopes. Curr. Opin. Virol. 11, 113–121 (2015).
Wrammert, J. et al. Broadly cross-reactive antibodies dominate the human B cell response against 2009 pandemic H1N1 influenza virus infection. J. Exp. Med. 208, 181–193 (2011).
Joyce, M. G. et al. Vaccine-induced antibodies that neutralize group 1 and group 2 influenza a viruses. Cell 166, 609–623 (2016).
Whittle, J. R. et al. Flow cytometry reveals that H5N1 vaccination elicits cross-reactive stem-directed antibodies from multiple Ig heavy-chain lineages. J. Virol. 88, 4047–4057 (2014).
Li, G. M. et al. Pandemic H1N1 influenza vaccine induces a recall response in humans that favors broadly cross-reactive memory B cells. Proc. Natl Acad. Sci. USA 109, 9047–9052 (2012).
Schmidt, A. G. et al. Viral receptor-binding site antibodies with diverse germline origins. Cell 161, 1026–1034 (2015).
Andrews, S. F. et al. Preferential induction of cross-group influenza A hemagglutinin stem-specific memory B cells after H7N9 immunization in humans. Sci. Immunol. 2, eaan2676 (2017).
Ekiert, D. C. et al. Cross-neutralization of influenza A viruses mediated by a single antibody loop. Nature 489, 526–532 (2012).
Lee, P. S. et al. Heterosubtypic antibody recognition of the influenza virus hemagglutinin receptor binding site enhanced by avidity. Proc. Natl Acad. Sci. USA 109, 17040–17045 (2012).
Whittle, J. R. et al. Broadly neutralizing human antibody that recognizes the receptor-binding pocket of influenza virus hemagglutinin. Proc. Natl Acad. Sci. USA 108, 14216–14221 (2011).
Lee, P. S.et al. Receptor mimicry by antibody F045-092 facilitates universal binding to the H3 subtype of influenza virus. Nat. Commun. 5, 3614 (2014).
Dreyfus, C. et al. Highly conserved protective epitopes on influenza B viruses. Science 337, 1343–1348 (2012).
Ekiert, D. C. et al. Antibody recognition of a highly conserved influenza virus epitope. Science 324, 246–251 (2009).
Okuno, Y., Isegawa, Y., Sasao, F. & Ueda, S. A common neutralizing epitope conserved between the hemagglutinins of influenza A virus H1 and H2 strains. J. Virol. 67, 2552–2558 (1993).
Corti, D. et al. A neutralizing antibody selected from plasma cells that binds to group 1 and group 2 influenza A hemagglutinins. Science 333, 850–856 (2011).
Iba, Y. et al. Conserved neutralizing epitope at globular head of hemagglutinin in H3N2 influenza viruses. J. Virol. 88, 7130–7144 (2014).
Krause, J. C. et al. A broadly neutralizing human monoclonal antibody that recognizes a conserved, novel epitope on the globular head of the influenza H1N1 virus hemagglutinin. J. Virol. 85, 10905–10908 (2011).
Xiong, X. et al. Structures of complexes formed by H5 influenza hemagglutinin with a potent broadly neutralizing human monoclonal antibody. Proc. Natl Acad. Sci. USA 112, 9430–9435 (2015).
Impagliazzo, A. et al. A stable trimeric influenza hemagglutinin stem as a broadly protective immunogen. Science 349, 1301–1306 (2015).
Yassine, H. M. et al. Hemagglutinin-stem nanoparticles generate heterosubtypic influenza protection. Nat. Med. 21, 1065–1070 (2015).
Mallajosyula, V. V. et al. Influenza hemagglutinin stem-fragment immunogen elicits broadly neutralizing antibodies and confers heterologous protection. Proc. Natl Acad. Sci. USA 111, E2514–E2523 (2014).
Krammer, F., Pica, N., Hai, R., Margine, I. & Palese, P. Chimeric hemagglutinin influenza virus vaccine constructs elicit broadly protective stalk-specific antibodies. J. Virol. 87, 6542–6550 (2013).
Victora, G. D. & Nussenzweig, M. C. Germinal centers. Annu. Rev. Immunol. 30, 429–457 (2012).
Doria-Rose, N. A. & Joyce, M. G. Strategies to guide the antibody affinity maturation process. Curr. Opin. Virol. 11, 137–147 (2015).
Lanzavecchia, A., Fruhwirth, A., Perez, L. & Corti, D. Antibody-guided vaccine design: identification of protective epitopes. Curr. Opin. Immunol. 41, 62–67 (2016).
Kanekiyo, M. et al. Rational design of an epstein-barr virus vaccine targeting the receptor-binding site. Cell 162, 1090–1100 (2015).
Kanekiyo, M. et al. Self-assembling influenza nanoparticle vaccines elicit broadly neutralizing H1N1 antibodies. Nature 499, 102–106 (2013).
Zimmer, S. M. & Burke, D. S. Historical perspective--Emergence of influenza A (H1N1) viruses. N. Engl. J. Med. 361, 279–285 (2009).
Xu, R. et al. Structural basis of preexisting immunity to the 2009 H1N1 pandemic influenza virus. Science 328, 357–360 (2010).
Dintzis, H. M., Dintzis, R. Z. & Vogelstein, B. Molecular determinants of immunogenicity: the immunon model of immune response. Proc. Natl Acad. Sci. USA 73, 3671–3675 (1976).
Bachmann, M. F. et al. The influence of antigen organization on B cell responsiveness. Science 262, 1448–1451 (1993).
Tiller, T., Busse, C. E. & Wardemann, H. Cloning and expression of murine Ig genes from single B cells. J. Immunol. Methods 350, 183–193 (2009).
Gerhard, W., Yewdell, J., Frankel, M. E. & Webster, R. Antigenic structure of influenza virus haemagglutinin defined by hybridoma antibodies. Nature 290, 713–717 (1981).
Caton, A. J., Brownlee, G. G., Yewdell, J. W. & Gerhard, W. The antigenic structure of the influenza virus A/PR/8/34 hemagglutinin (H1 subtype). Cell 31, 417–427 (1982).
Hong, M. et al. Antibody recognition of the pandemic H1N1 Influenza virus hemagglutinin receptor binding site. J. Virol. 87, 12471–12480 (2013).
Yewdell, J. W., Gerhard, W. & Bachi, T. Monoclonal anti-hemagglutinin antibodies detect irreversible antigenic alterations that coincide with the acid activation of influenza virus A/PR/834-mediated hemolysis. J. Virol. 48, 239–248 (1983).
Raymond, D. D. et al. Conserved epitope on influenza-virus hemagglutinin head defined by a vaccine-induced antibody. Proc. Natl Acad. Sci. USA 115, 168–173 (2018).
Neu, K. E., Henry Dunand, C. J. & Wilson, P. C. Heads, stalks and everything else: how can antibodies eradicate influenza as a human disease? Curr. Opin. Immunol. 42, 48–55 (2016).
Lambert, L. C. & Fauci, A. S. Influenza vaccines for the future. N. Engl. J. Med. 363, 2036–2044 (2010).
Hai, R. et al. Influenza viruses expressing chimeric hemagglutinins: globular head and stalk domains derived from different subtypes. J. Virol. 86, 5774–5781 (2012).
Carter, D. M. et al. Elicitation of protective antibodies against a broad panel of H1N1 viruses in ferrets pre-immune to historical H1N1 influenza viruses.J. Virol. 91, e01283-17 (2017).
Carter, D. M. et al. Design and characterization of a computationally optimized broadly reactive hemagglutinin vaccine for H1N1 influenza viruses. J. Virol. 90, 4720–4734 (2016).
Belongia, E. A. et al. Repeated annual influenza vaccination and vaccine effectiveness: review of evidence. Expert Rev. Vaccines 16, 1–14 (2017).
Chai, N. et al. A broadly protective therapeutic antibody against influenza B virus with two mechanisms of action. Nat. Commun. 8, 014234 (2017).
Shen, C. et al. A multimechanistic antibody targeting the receptor binding site potently cross-protects against influenza B viruses. Sci. Transl. Med. 9, eaam5752 (2017).
Ngwuta, J. O. et al. Prefusion F-specific antibodies determine the magnitude of RSV neutralizing activity in human sera. Sci. Transl. Med. 7, 309ra162 (2015).
Ledgerwood, J. E. et al. Prime-boost interval matters: a randomized phase 1 study to identify the minimum interval necessary to observe the H5 DNA influenza vaccine priming effect. J. Infect. Dis. 208, 418–422 (2013).
Wei, C. J. et al. Induction of broadly neutralizing H1N1 influenza antibodies by vaccination. Science 329, 1060–1064 (2010).
Wheatley, A. K. et al. H5N1 vaccine-elicited memory B cells are genetically constrained by the IGHV locus in the recognition of a neutralizing epitope in the hemagglutinin stem. J. Immunol. 195, 602–610 (2015).
Brochet, X., Lefranc, M. P. & Giudicelli, V. IMGT/V-QUEST: the highly customized and integrated system for IG and TR standardized V-J and V-D-J sequence analysis. Nucleic Acids Res. 36, W503–W508 (2008).
Kong, W. P. et al. Protective immunity to lethal challenge of the 1918 pandemic influenza virus by vaccination. Proc. Natl Acad. Sci. USA 103, 15987–15991 (2006).
Yang, Z. Y. et al. Immunization by avian H5 influenza hemagglutinin mutants with altered receptor binding specificity. Science 317, 825–828 (2007).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Yu, X. et al. Neutralizing antibodies derived from the B cells of 1918 influenza pandemic survivors. Nature 455, 532–536 (2008).
Misasi, J. et al. Structural and molecular basis for Ebola virus neutralization by protective human antibodies. Science 351, 1343–1346 (2016).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Pettersen, E. F. et al. UCSF Chimera--a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Rohou, A. & Grigorieff, N. CTFFIND4: fFast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Scheres, S. H. Semi-automated selection of cryo-EM particles in RELION-1.3. J. Struct. Biol. 189, 114–122 (2015).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Acknowledgements
We thank J. Weir (Food and Drug Administration) for providing influenza viruses; R. Webby (St Jude Children’s Research Hospital) for influenza reverse genetics plasmids; M. Dillon, K. Wuddie, G. Sarbador, C. Chiedi, A. Taylor, H. Bao and D. Scorpio (VRC) for help with animal studies; A. Kumar (VRC) for help with protein production; V. Nair and E. Fischer (NIAID) for cryo-electron microscopy data collection; M. Chen (VRC), D. Angeletti and J. Yewdell (NIAID) for help with initial epitope mapping. This work used the computational resources of the NIH HPC Biowulf cluster (http://hpc.nih.gov) and also the Office of Cyber Infrastructure and Computational Biology (OCICB) High Performance Computing (HPC) cluster at the NIAID, NIH. Use of insertion device 22 (SER-CAT) at the Advanced Photon Source was supported by the US Department of Energy, Office of Science, Basic Energy Sciences (under contract W-31-109-Eng-38). Support for this work was provided by the NIAID Intramural Research Program to the VRC and Division of Intramural Research. This work was also supported in part with federal funds from the Frederick National Laboratory for Cancer Research, NIH (under contract HHSN261200800001E).
Author information
Authors and Affiliations
Contributions
M.K. and B.S.G. conceptualized and devised studies. M.K. designed immunogens. M.K., R.A.G., H.M.Y. and S.B.B. performed animal studies. M.G.J. crystallized and solved Fab structure. J.R.G. and A.K.H. determined cryo-electron microscopy structure. S.F.A., A.K.W., B.E.F., D.R.A. and M.S.P. performed FACS and single-cell PCR. M.K., R.A.G., S.F.A. and H.M.Y. produced and characterized proteins. A.C. made viruses by reverse genetics. R.A.G., H.M.Y., K.L., E.S.Y. and W.-P.K. performed virus neutralization assays. I.S.G. performed mathematical simulations. Y.T. and U.B. performed electron microscopy experiments. H.A. performed challenge studies. K.L.Z. and J.E.L. conducted clinical trials and provided human samples. M.K., M.G.J., R.A.G., J.R.G., S.F.A., H.M.Y., A.K.W., A.K.H., R.A.K., P.D.K., A.B.M., J.R.M. and B.S.G. analyzed data. M.K. and B.S.G. wrote paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The US National Institutes of Health has filed a patent application on the basis of the studies presented in this paper; M.K., H.M.Y. and B.S.G. are listed as inventors on the patent application.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Fig. 1 Biochemical characterization of hemagglutinin RBD–np.
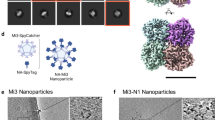
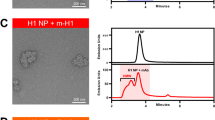
Chromatograms of homogeneously assembled and heterogeneously co-assembled hemagglutinin RBD–np on anion-exchange column (a) and size exclusion column (b). NC99 and CA09 hemagglutinin RBD–np were produced in Expi293 cells by transient transfection and the culture supernatants were purified. Anion-exchange chromatography was carried out using a HiTrap Q HP column eluted with linear gradient of NaCl (a). Fractions containing RBD–np corresponding to the shaded area were collected and subjected to size exclusion chromatography (SEC). SEC was carried out using a Superose 6 matrix to confirm particle formation (b). Experiments were independently performed three times with similar results. Purified RBD–np expressing various hemagglutinin RBD from indicated H1N1 strains homogeneously (c) or heterogeneously co-assembled (mosaic) (d) were analyzed on SDS–PAGE and blue staining. For comparison, mechanical mixtures of 2, 4, 6 or 8 different RBD–np were prepared and analyzed on the same gel (admix) (d). Experiments were independently performed two times with similar results. Size exclusion chromatography profiles of hemagglutinin RBD–np shown in panel c and d (e and f, respectively). Experiments were performed two times with similar results.
Supplementary Fig. 2 Antibody responses elicited by hemagglutinin RBD–np and commercial vaccines to influenza virus.
Serum HAI and neutralization antibody titers to CA09 (a) and BR07 (b). BALB/c mice were immunized with NC99 RBD–np (valence 1) or various preparations of various RBD–np either as admixtures (admix) or as co-assembled particles (Mosaic). Represented data were generated using sera collected 2 weeks after the second immunization with adjuvant (2 µg of RBD–np per immunization). Shown results were obtained from a representative experiment (n = 5). Each dot indicates individual mouse with mean of each group indicated as lines. Animal experiments were independently performed two times with similar results. Statistical analyses are done with one-way ANOVA with Tukey’s multiple comparisons post-hoc test. Non-statistical significance between groups is not shown for clarity. (c) Microneutralization (MN) titers against WS33, KW86, and CC03 viruses of pooled immune sera. Mice (n = 5) were immunized with 2 µg of mosaic RBD–np (8-valent), or 1.5 µg of commercial TIV (from indicated season) formulated with SAS and delivered at weeks 0 and 4. Immune sera were collected at week 6, RDE-treated, heat-inactivated, and pooled for microneutralization assays. Animal experiment was performed once.
Supplementary Fig. 3 Hemagglutinin-binding and H1N1 neutralization of monoclonal antibodies.
(a) Hemagglutinin-binding of monoclonal antibodies measured by biolayer interferometry. Recombinant hemagglutinin trimer proteins were immobilized on HIS1K biosensors through hexa-histidine tag and used to assess binding of monoclonal antibodies. All assays were performed in the Octet HTX instrument (fortéBio) at 30ºC with agitation at 1,000 rpm. PBS with 1% BSA was used as assay buffer for all steps. Plotted data indicate the response value during the mAb association step at 300 s. Control monoclonal antibodies 3u-u, 2D1 and CH65 were shown as gray bars. Experiments were performed two times with similar results. (b) Neutralization capacity of monoclonal antibodies. Pseudotype neutralization assays were used to assess neutralization potency of isolated monoclonal antibodies. Homologous NC99 and heterologous BJ95 strains were selected for screening. Experiments were performed two times with similar results.
Supplementary Fig. 4 Hemagglutinin-binding kinetics of 441D6 Fab by biolayer interferometry.
Binding kinetics measurements were carried out using biolayer interferometry with Octet HTX instrument (fortéBio). Recombinant hemagglutinin trimers were immobilized either on HIS1K or SA sensors through hexa-histidine tag or conjugated biotin, respectively. Fab 441D6 was prepared by introducing HRV3C cleavage site in the heavy chain hinge and digesting with HRV3C enzyme followed by Protein A affinity purification to remove Fc and uncleaved antibodies as described elsewhere7. Dilution series of Fab 441D6 was made in assay buffer (PBS with 1% BSA). All assays were performed in the Octet HTX instrument (fortéBio) at 30ºC with agitation at 1,000 rpm. Represented data contain measured sensograms (red) and global fitting with 1:1 binding model (black) generated by Octet Analysis software (version 9.0, fortéBio). Experiments were independently performed two times with similar results.
Supplementary Fig. 5 Structure determination of HA–441D6 complex by cryo-electron microscopy.
(a) Micrographs indicated the HA–441D6 complex partitioned equally between the carbon surface and carbon holes. (b) Picked particles contributing to the determined structure are indicated by green circles, visualized on the same micrograph shown in a. (c). Model-free 2D class averages reveal 3-fold symmetry of the complex. Experiment was performed once. (d) Class occupancy from 2D classification is plotted corresponding to classes in c. (e) Fourier-shell correlation analysis of ‘gold-standard’ refinement using a cutoff of 0.143 estimates the model resolution as 8.0 Å.
Supplementary Fig. 6 electron microscopy analysis of mosaic RBD–np particle and its complex with 441D6 Fab.
(a) Raw electron microscopy images and representative 2D classes of mosaic RBD–np. Individual particles were visualized by PTA negative staining, revealing the ferritin core particle as either a filled or hollow white circle, depending on stochastic penetration of the ferritin core by the negative stain. RBD domains were seen as points immediately surrounding the ferritin core. The inset shows 4 representative class averages from 60,842 picked particles, after initial picking of 65,517 particles in 174 micrographs. (b) Mosaic RBD–np in complex with 441D6 Fab was similarly analyzed by PTA negative staining electron microscopy. The extra density surrounding the ferritin core is consistent with the presence of a coat of bound 441D6 Fab. The inset shows representative class averages of the complex, but unlike the RBD–np alone, the particles that stained hollow or filled required separate processing as they did not naturally partition into distinct classes during classification. From a total pool of 9,187 particles in 134 micrographs, final class averages of filled particles were made from 6,926 particles, and hollow particles were made from 957 particles. Experiment was performed once. (c) Rotationally averaged images of RBD–np particles (bottom) and 441D6 Fab complexes (top) were synthesized from class averages, and then juxtaposed to illustrate the bulk effect of 441D6 binding. The inner sphere represents the ferritin core, while the middle density of lesser intensity is either RBD or RBD+Fab. The outermost layer resulted from lighter staining of the electron microscopy grid carbon surface, since the negative stain accumulated in higher amounts proximal to particles. (d) Relative mean intensity of the rotational averages was plotted as a function of particle radius. High intensity corresponds to white in images in c. The outer radius of the RBD–np is 13 nm, while mosaic RBD–np–441D6 Fab complexes extend to 15.5 nm from the particle center. We interpret the first minima near 7 nm for the 441D6 complex as accumulation of negative stain between the ferritin core and Fabs that are tethered via the RBD domains at varying orientations.
Supplementary Fig. 7 Therapeutic effect of 441D6 in mouse infection model.
Therapeutic administration of control VRC01 (anti-HIV) and 441D6 monoclonal antibodies in mice infected with A/California/07/2009 (H1N1) (a) or A/Puerto Rico/8/1934 (H1N1) (b). BALB/c mice were intranasally infected with a 10 × LD50 of either H1N1 viruses 24 hours before receiving monoclonal antibodies via intraperitoneal injection. Clinical symptoms and body weight were monitored throughout the study. Lines in plots showing body weight change indicate mean of group with standard deviations including only live animals at indicated time points. Experiment was performed once (n = 10).
Supplementary Fig. 8 Generation of 441D6KO hemagglutinin and its recognition by immune serum.
(a) Generation and characterization of hemagglutinin with 441D6KO mutation. Two point mutations, E116K and K172E were introduced in the NC99 hemagglutinin (highlighted in black on the hemagglutinin molecular surface). Antigenicity of 441D6KO hemagglutinin was assessed by ELISA using 441D6, CH65, and anti-His (Covance) antibodies. Binding curves of antibodies to WT and 441D6KO hemagglutinin were depicted. Assays were independently performed two times with similar results. (b) Serum binding antibody titers to WT and 441D6KO hemagglutinin. The endpoint titers in immune sera raised by NC99 HA-, admix (8-valent)-, mosaic (8-valent)-, and sequential RBD–np immunization were measured by ELISA (n = 10, biologically independent samples). Assays were independently performed two times with similar results. (c) Serum binding antibody titers to WT and 441D6KO hemagglutinin. The endpoint titers were determined as above (n = 26, biologically independent samples). Assays were independently performed two times with similar results. Statistical analyses were done with nonparametric t-test using Wilcoxon matched-paired signed rank test (two-tailed).
Supplementary information
Supplementary Figs. 1-8
Supplementary Figs. 1-8 Supplementary Table 1
Rights and permissions
About this article
Cite this article
Kanekiyo, M., Joyce, M.G., Gillespie, R.A. et al. Mosaic nanoparticle display of diverse influenza virus hemagglutinins elicits broad B cell responses. Nat Immunol 20, 362–372 (2019). https://doi.org/10.1038/s41590-018-0305-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0305-x
This article is cited by
-
Enhancing antibody responses by multivalent antigen display on thymus-independent DNA origami scaffolds
Nature Communications (2024)
-
Vaccine design via antigen reorientation
Nature Chemical Biology (2024)
-
Nanoparticle display of neuraminidase elicits enhanced antibody responses and protection against influenza A virus challenge
npj Vaccines (2024)
-
An attachment glycoprotein nanoparticle elicits broadly neutralizing antibodies and protects against lethal Nipah virus infection
npj Vaccines (2024)
-
Bringing immunofocusing into focus
npj Vaccines (2024)