Abstract
Colonization of the upper respiratory tract by pneumococcus is important both as a determinant of disease and for transmission into the population. The immunological mechanisms that contain pneumococcus during colonization are well studied in mice but remain unclear in humans. Loss of this control of pneumococcus following infection with influenza virus is associated with secondary bacterial pneumonia. We used a human challenge model with type 6B pneumococcus to show that acquisition of pneumococcus induced early degranulation of resident neutrophils and recruitment of monocytes to the nose. Monocyte function was associated with the clearance of pneumococcus. Prior nasal infection with live attenuated influenza virus induced inflammation, impaired innate immune function and altered genome-wide nasal gene responses to the carriage of pneumococcus. Levels of the cytokine CXCL10, promoted by viral infection, at the time pneumococcus was encountered were positively associated with bacterial load.
Similar content being viewed by others
Main
Pneumonia is a major global health problem; it kills more children under 5 years of age than any other disease1. The burden of disease is aggravated by old age, chronic lung disease, immunosuppression and viral co-infection. Secondary pneumonia following pandemic and seasonal influenza virus infection is a significant cause of mortality worldwide2.
Nasopharyngeal colonization by Streptococcus pneumoniae (pneumococcus) is common, with 40–95% of infants and 10–25% of adults colonized at any given time3. Such pneumococcal carriage is important as the prerequisite of infection4, the primary reservoir for transmission5 and the predominant source of immunizing exposure and immunological boosting in both children and adults6,7.
Immune dysregulation caused by infection with respiratory viruses such as influenza leads to increased carriage load8. Increased carriage load has been associated with pneumonia incidence and severity, as well as with within-household S. pneumoniae transmission5,9,10,11. The mechanisms and markers associated with this pathogen synergy have been difficult to study in human subjects due to the rapidly progressing nature of the disease.
One safe way to simulate influenza infection in the nose is using live attenuated influenza vaccine (LAIV), consisting of cold-adapted influenza viruses. LAIV has been shown to affect subsequent susceptibility to S. pneumoniae and to lead to increased carriage load in mouse models of infection and in vaccinated children12,13. Furthermore, LAIV administration prior to S. pneumoniae challenge led to a 50% increase in S. pneumoniae acquisition, as detected by molecular methods, as well as a tenfold increase in nasopharyngeal bacterial load14.
In mouse models of pneumococcal carriage, recruitment of neutrophils and monocytes to the nasopharynx dependent on the TH17 subset of helper T cells mediates immunological control and clearance15,16,17. Influenza virus infection promotes type I interferons, which interfere with recruitment of these phagocytes, although interferon-γ (IFN-γ) is postulated to impair phagocytosis by macrophages through downregulation of the scavenger receptor MARCO (macrophage receptor with collagenous structure)18,19,20. However, the precise immune mechanisms and gene regulators involved in the control and clearance of pneumococcal carriage in humans have not been revealed21. Moreover, how these mechanisms are altered during human influenza virus infection remains largely unknown.
Systems-biology approaches have allowed the identification of immune mechanisms associated with protection from infectious diseases and with robust immune responses during vaccination22,23,24,25,26,27,28. Here, we applied systems biology to nasal samples collected in the setting of human challenge with LAIV and S. pneumoniae, to emulate nasal effects of influenza infection on S. pneumoniae carriage. We identified the key cellular mechanisms that control newly acquired pneumococcal carriage in humans, and how they are disrupted following nasal influenza infection.
Results
LAIV-induced inflammation leads to increased pneumococcal carriage load and acquisition
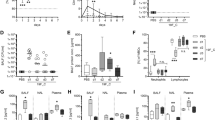
In a double-blinded controlled randomized clinical trial, we administered LAIV (n = 55) 3 days prior to S. pneumoniae inoculation (day 0). To verify the requisite topical application for an effect on pneumococcal carriage, we administered tetravalent inactivated influenza vaccine as a control (n = 62). LAIV infection led to transiently increased pneumococcal acquisition at day 2 (60.0% and 40.3%, as detected by molecular methods, in LAIV groups and control groups, respectively)14. LAIV also increased S. pneumoniae carriage load in the first 14 days following pneumococcal inoculation14 (Supplementary Fig. 1). We collected a series of nasal microbiopsies and nasal lining fluid throughout the study to assess ongoing cellular and cytokine responses. Participants were grouped into those who did not become colonized following S. pneumoniae challenge (carriage−) and those who did (carriage+), as determined by classical microbiology (Fig. 1a). To investigate whether LAIV-induced immune responses were associated with a predisposition to pneumococcal carriage, we measured the concentrations of 30 cytokines and of proteins, including interleukin 1 receptor antagonist (IL-1RA), in nasal lining fluid (Fig. 1b). After multiple-testing correction, we found that at day 0, directly prior to S. pneumoniae inoculation, LAIV significantly increased concentrations of 20 cytokines, including CXCL10 (IP-10), tumor-necrosis factor (TNF), interleukin 10 (IL-10), IFN-γ and IL-15 (Fig. 1b and Supplementary Table 1). In contrast, the control group did not show any significant increase in cytokine response at day 0. Following S. pneumoniae inoculation, S. pneumoniae carriage in the absence of LAIV was associated with increased concentration of epidermal growth factor at day 2 and decreased concentration of IL-1RA at day 9 after S. pneumoniae inoculation (compared with that at baseline), neither of which remained significant after multiple-testing correction. No other cytokines, including IL-17A or CCL2, were significantly altered by carriage alone (Fig. 1b).
a, Experimental design of the study. Analyzed time points are indicated by black circles. b, Heat map showing for each cytokine the median log2[fold change] compared with baseline for the time points 0, 2,7 and 9 days; n = 19 per group. c, The delta in median log2[fold change] following LAIV vaccination just prior to inoculation with S. pneumoniae for subjects becoming carriage+ or carriage− (excluding subjects becoming positive by PCR only, who resemble subjects that become carriage+ by culture as well). The color of each bar represents the median induction in the entire LAIV group. **P = 0.0097 by two-tailed Wilcoxon test for LAIV carriage− subjects comparing IL-10 day 0 with baseline; P = 0.073 for the LAIV carriage+ group. ***P = 0.0008 by two-tailed Wilcoxon test for LAIV carriage+ subjects comparing CXCL10 day 0 with baseline; P = 0.051 for the LAIV carriage− group. d, Pneumococcal load (median and interquartile range of CFU ml−1 in nasal wash shown) for all carriage+ subjects with high (top quartile, n = 9) or low (all subjects below top quartile, n = 28) CXCL10 concentrations at day 0. P = 0.019 by two-tailed Mann-Whitney test of area under the curve of log-transformed load over time. e, Scatter plot showing correlation of CXCL10 concentration at baseline with S. pneumoniae load for a second validation cohort (n = 52) with an asymptomatic upper respiratory tract virus infection (n = 15) or not. Spearman correlation test results and linear regression line with 95% confidence interval (gray shading) are shown.
Even before bacterial inoculation, nasal inflammatory responses to LAIV differed between those who went on to become carriers and those who were protected from carriage (Fig. 1c). In particular, IL-10 was significantly increased in LAIV-vaccinated subjects who did not acquire S. pneumoniae following inoculation (5.8-fold increase), but not in those who became carriers following inoculation (2.0-fold increase). In contrast, CXCL10 was significantly increased in subjects who went on to become carriers (2.4-fold increase), but not in those who remained carriage-negative (1.5-fold increase). Moreover, subjects with increased concentrations of CXCL10 before inoculation displayed higher pneumococcal load following S. pneumoniae inoculation (Fig. 1d). This suggests that differences in the response to influenza virus are associated with secondary susceptibility to S. pneumoniae. To test whether this was specific for LAIV infection, we measured CXCL10 in nasal washes from an independent cohort in which a subset of subjects had an asymptomatic viral upper respiratory tract infection the week before S. pneumoniae inoculation that did not progress to symptomatic infection. These comprised infections with rhinovirus (n = 12), coronavirus (n = 5), respiratory syncytial virus (n = 2) and parainfluenzavirus (n = 1)29. The predominant virus, rhinovirus, was recently shown to associate with increased pneumococcal acquisition and transmission30. In these virus-infected subjects, CXCL10 concentration was increased (Supplementary Fig. 2), and baseline CXCL10 concentration correlated with increased pneumococcal load also in this second cohort (Fig. 1e). Unfortunately, sample sizes were too small to further investigate in depth the effects of infection with the different viruses, which are likely to have divergent consequences. Although the correlation was modest in this validation cohort, suggesting that other host and environmental factors are involved, we identified a biomarker, CXCL10, that was predictive of S. pneumoniae load.
Early neutrophil degranulation in response to carriage is impaired by LAIV infection
In mouse models, neutrophil recruitment after onset of carriage contributes to control of the bacteria15. We observed pre-existing high numbers of neutrophils in the human nasal lining, and pneumococcal carriage did not lead to significant further recruitment of neutrophils (Supplementary Fig. 3a,b). To investigate whether luminal neutrophils were involved in the early control of carriage, we measured concentrations of myeloperoxidase, a marker for neutrophil degranulation31, in nasal washes. Concentrations were increased (2.2-fold) at 2 days after challenge in control carriage+ but not carriage− individuals (Fig. 2a). This neutrophil activation was impaired in the LAIV group, who displayed high carriage load during early carriage and had increased acquisition compared with that of controls. Together, these findings suggest that neutrophil degranulation is important for the initial control of carriage. To investigate whether neutrophils were also impaired systemically following LAIV, as reported during wild-type influenza infection32, we isolated blood neutrophils before, and at 3 days after, LAIV administration from a subset of subjects. We confirmed that opsonophagocytic killing (OPK) of S. pneumoniae by blood neutrophils was decreased following LAIV (Fig. 2b). This effect could be mimicked by the addition of TNF, but not CXCL10, to neutrophils from healthy donors in vitro, decreasing killing capacity in a dose-dependent manner (Fig. 2c,d). NanoString expression analysis of 594 genes revealed ten differentially expressed genes in blood neutrophils 3 days post-LAIV (Supplementary Table 2). Among those were the genes encoding MAP4K2 (mitogen-activated protein kinase kinase kinase kinase 2; 3.2-fold increase), which acts on the TNF signal-transduction pathway33, and the co-inhibitory receptor TIGIT (T cell immunoreceptor with immunoglobulin and ITIM domains; 3.6-fold increase; Fig. 2e). TIGIT expression levels were also negatively correlated with neutrophil killing capacity (r = −0.73; Fig. 2f). TIGIT is an immunological checkpoint protein that has been described to promote regulatory T cell function34, but its expression on neutrophils has not been previously appreciated, to our knowledge. Incubation of whole blood with recombinant TNF increased TIGIT levels on neutrophil surfaces within 30 min in a dose-dependent manner (Fig. 2g).
a, Concentrations of myeloperoxidase in nasal wash of volunteers before or 2 days after S. pneumoniae inoculation. Median and interquartile range are shown (for n = 9 LAIV carriage− and LAIV carriage+ and for n = 10 control carriage− and control carriage+ subjects). *P = 0.014 by two-tailed Wilcoxon paired test. b, S. pneumoniae OPK capacity of blood neutrophils before and 3 days following LAIV (n = 6) or control (tetravalent inactivated influenza vaccine or no, n = 7) vaccination. Individual subjects are shown and connected by lines. *P = 0.031 by two-tailed Wilcoxon paired test. NS, not significant. c,d, Effect of exogenous TNF (c; n = 10) and CXCL10 (d; n = 8) on OPK activity of blood neutrophils of healthy volunteers. **P = 1.15 × 10−5 by Friedman test. Neutrophils from six subjects were used in three independent experiments. Individual samples are depicted and connected by dashed lines. e, Normalized MAP4K2 and TIGIT counts on sorted neutrophils before LAIV or in control arm (n = 6, red) and following LAIV (n = 4, blue). Individual samples are shown, and paired samples are connected by black lines. **P = 0.008 and ***P = 3.2 × 10−5 two-tailed unadjusted P values using a negative binomial generalized linear model (DESeq2). f, Correlation between OPK activity and TIGIT counts (n = 10). Spearman rho (r) and P value are shown. Regression line and 95% confidence intervals (shaded area) are shown. g, Levels of TIGIT on blood neutrophil surface measured by flow cytometry after a 30-min incubation without or with 1 ng ml−1 TNF or 100 ng ml−1 TNF (n = 4). *P = 0.042 by Friedman test. Individual subjects are depicted by dots and connected by lines.
Taken together, these findings indicate that inflammation following LAIV impairs the response of nasal and systemic neutrophils to pneumococcus, which could be mimicked by addition of exogenous TNF to neutrophils and is associated with an upregulation of TIGIT.
Pneumococcal carriage–induced monocyte recruitment to the nose is impaired by LAIV infection
Immunophenotyping revealed a significant recruitment of monocytes to the nose following establishment of carriage (Fig. 3a and Supplementary Fig. 4). Monocyte numbers increased as early as 2 days following S. pneumoniae inoculation, peaked at 9 days (median 4.8× increase) and remained elevated 29 days after S. pneumoniae inoculation. In contrast, there was no recruitment of CD3+ T cells to the nose (Supplementary Fig. 4b). LAIV infection prior to pneumococcal carriage impaired the recruitment of monocytes to the nose (Fig. 3a). Moreover, peak pneumococcal load was associated with increased monocyte recruitment in the control group, but not in the LAIV group (Fig. 3b,c). Indeed, for subjects in the control group with very low carriage densities, which were detectable only by molecular methods, no monocyte recruitment was observed (Supplementary Fig. 4c). This suggests that a minimum S. pneumoniae load is required for sensing and monocyte recruitment and that LAIV infection interferes with this process. Although CCL2 (MCP-1) was not substantially induced following S. pneumoniae carriage, its concentration was correlated with numbers of monocytes at all time points (Supplementary Fig. 5a). Furthermore, stratification of individuals showed that those with increased CCL2 concentration at day 2 after S. pneumoniae inoculation exhibited increased monocyte recruitment (Supplementary Fig. 5b). Concentrations of IL-6, IFN-γ and TNF were also correlated with numbers of monocytes at each time point, but stratification of individuals did not reveal differential recruitment of monocytes (Supplementary Fig. 5a,b). In a second, independent cohort that did not receive any vaccine, monocytes were increased at day 9 following S. pneumoniae inoculation, which was correlated with an increased CCL2 concentration in nasal fluid, validating these results (Supplementary Table 3 and Supplementary Fig. 5c).
a, Median and interquartile range of nasal monocyte numbers normalized to epithelial cell numbers are shown for control carriage+ (n = 24), control carriage− (n = 37), LAIV carriage+ (n = 25) and LAIV carriage− (n = 30) groups. The dashed green line shows the baseline level in the control carriage+ group. *P = 0.038 at day 2 and P = 0.030 at day 29; **P = 0.002 by two-tailed Wilcoxon paired non-parametric test. b,c, Levels of maximum pneumococcal (S. pneumoniae) load in nasal wash are shown for the control group (b; n = 22) and LAIV group (c; n = 23) and are correlated with the maximum monocyte recruitment (fold change to baseline). Individual subjects are shown, and Spearman correlation analysis is shown.
Thus, acquisition of pneumococcal carriage led to a recruitment of monocytes to the nasopharynx, a process that was associated with pneumococcal load and CCL2 induction and that was inhibited by LAIV infection.
Nasal responses associated with pneumococcal clearance are impaired by LAIV
To assess anti-pneumococcal responses induced by carriage, we collected nasal cells 29 days after S. pneumoniae inoculation, stimulated them in vitro with heat-killed S. pneumoniae and measured concentrations of 30 cytokines in the supernatant. Increased production (fold change > 2 and q < 0.05 (P value corrected for multiple testing) to unstimulated control) of TNF, macrophage inflammatory protein 1α (MIP-1α), IL-10, IL-6 and granulocyte–macrophage colony-stimulating factor upon restimulation was observed in the control carriage+ group (Fig. 4a and Supplementary Fig. 6a). In the LAIV carriage+ group, however, this boosting of anti-pneumococcal cytokine responses by rechallenge was absent (Fig. 4a and Supplementary Fig. 6a). The production of the above five cytokines correlated with decreased pneumococcal load at day 29 after S. pneumoniae inoculation, suggesting these responses are involved in S. pneumoniae clearance (Fig. 4b). To test whether monocytes/macrophages were the source of these cytokines, we compared the cytokine signature from whole nasal cells with that from alveolar macrophages exposed to S. pneumoniae in vitro (Fig. 4c). Relative cytokine production was highly correlated between the two cell populations, suggesting that nasal monocytes/macrophages could be the source of these cytokines. This is supported by the observation that in carriers with low carriage load (detectable only by molecular methods), absence of monocyte recruitment was associated with absent S. pneumoniae-specific responses (Supplementary Fig. 6b).
a, Whole nasal cells were collected from 48 subjects 28 days post-inoculation and stimulated for 18 h with heat-killed S. pneumoniae. Supernatant was collected, and concentrations of 30 cytokines were measured by multiplex enzyme-linked immunosorbent assay. The median and interquartile range for cytokines induced at least twofold in at least one condition are displayed. b, Correlations between cytokine production following S. pneumoniae stimulation and pneumococcal load are shown (n = 22). Spearman non-parametric correlation test results and regression lines with shaded 95% confidence intervals are shown per cytokine. c, The cytokine profile from alveolar macrophages (median for six volunteers shown) exposed to S. pneumoniae for 18 h was compared with that of stimulated whole nasal cells (median of control carriage+ group shown). Spearman non-parametric correlation test results and regression lines with shaded 95% confidence intervals are shown.
In conclusion, carriage led to increased responses of nasal cells to pneumococcal stimulation, which was potentially due to the infiltration of monocytes. This was impaired by prior LAIV infection and was correlated with clearance of pneumococcal carriage (Supplementary Fig. 7).
LAIV alters nasal gene-expression responses to carriage
To identify gene signatures associated with the observed responses to pneumococcal carriage and infection with LAIV, we performed RNA sequencing on whole nasal cells at days −5, 2 and 9 after S. pneumoniae inoculation (Fig. 5 and Supplementary Table 4). Carriage without LAIV induced 834 and 176 differentially expressed genes (DEGs) at days 2 and 9, respectively (Fig. 5a). These genes were enriched for pathways associated with gap junction trafficking and regulation (including GJA1, TJP1 and multiple GJB genes) and degradation of the extracellular matrix (including COL17A1, COL12A1, LAMA3 and KLK7). In the carriage− group, a smaller number of DEGs was observed (161 and 248 at days 2 and 9, respectively).
a, The number of differentially expressed genes (DEGs) between each time point and the baseline for each group are shown. Upregulated and downregulated genes are depicted in red and blue, respectively. Connections between bars show the number of common genes between LAIV and control conditions, where colors reflect distinct pathways. b, Circular representation of DEGs and gene set enrichment analysis for LAIV carriage+ and control carriage+ groups at days 2 and 9 after S. pneumoniae inoculation. The individual log2[fold change] values (baseline-normalized) were used as ranks in a single sample gene set enrichment analysis to identify consistently enriched pathways among subjects. Genes and pathways are connected by lines.
In the LAIV carriage+ group, 936 and 711 DEGs were observed at day 2 and day 9, respectively. Surprisingly, despite the high concentrations of inflammatory cytokines observed in the LAIV carriage− group, only a relatively small number of DEGs were observed at days 2 and 9 (126 and 153, respectively). DEGs of carriage+ subjects receiving LAIV and DEGs of carriage+ subjects without LAIV showed very little overlap, with only 38 DEGs at day 2 and 2 DEGs at day 9 in common. Very little overlap was observed at the pathway level between these groups, indicating LAIV alters the natural responses to pneumococcus (Fig. 5b and Supplementary Table 5). This could reflect transcriptome kinetics, due to altered differentiation and cellular activation, or it could reflect changes in cell migration to the nasal mucosa.
The LAIV carriage+ group showed enrichment for genes encoding molecules involved in the Toll-like receptor 3 (TLR3) signaling cascade, in retinoic acid–inducible gene I (RIG-I)/melanoma differentiation-associated protein 5 (MDA5)-mediated induction of IFN-α/β pathways and in IFN-γ signaling, which is in agreement with the induction of antiviral responses following LAIV vaccination35. Moreover, TLR4 signaling was also enriched in this group. The pneumococcal protein pneumolysin is sensed by TLR436, and it is possible that the increased pneumococcal load following LAIV vaccination led to increased pneumolysin sensing. O-linked glycosylation of mucins, which are used by S. pneumoniae as a carbohydrate source for growth37, was also enriched in the LAIV carriage+ group (including the genes ST3GAL4, GALNT7, GCNT3 and B4GALT5). ST3GAL4 encodes a sialyl transferase, and cleavage of sialic acids by the influenza neuraminidase has previously been shown to promote pneumococcal growth38. This finding supports a LAIV-mediated effect on pneumococcal growth through alterations of host factors. Common genes and pathways between the LAIV-vaccinated carriers and control carriers include ‘innate immune system’ and ‘signaling by interleukins’ (IL1B, CLEC4E, CD55 and IL1RN). In conclusion, the genome-wide transcriptomic response to pneumococcal carriage was substantially altered on both the gene level and the pathway level by LAIV.
Gene modules associated with recruitment of monocytes
To identify sets of coexpressed genes post-LAIV and carriage, we used CEMiTool on the baseline-normalized data of LAIV and control groups, separately39. This modular expression analysis revealed genes that may act together or are similarly regulated during the immune responses to carriage and infection.
Genes in the control cohort were grouped into four coexpression modules, of which three were significantly enriched for known reactome pathways (Supplementary Data 1). Module M1 was enriched in the carriage+ group at day 9 after S. pneumoniae inoculation (Fig. 6a). Numbers of monocytes were correlated with the average fold change count in this module, suggesting that these genes reflect the infiltration of monocytes (Fig. 6b). To further investigate these monocytes, we performed gene set enrichment analysis on the module M1 genes using a list of genes from distinct monocyte subsets (Fig. 6c)40. These genes were enriched for classical CD14+CD16– monocytes and not for other monocyte subsets. Moreover, this module was enriched for genes related to ‘chemokine receptors bind chemokines’ and ‘IFN-α/β signaling’ (Fig. 6d). Type I interferons have been shown to be required for the clearance of pneumococcal carriage in mouse models41, and these findings suggest that their activity in monocytes might be critical for this. CEMiTool also integrates coexpression analysis with protein–protein interaction data. Expression of the gene encoding the chemokine CXCL6 and that of the gene encoding its receptor CXCR2 were identified as hubs in module M1 (Fig. 6e and Supplementary Data 1). CXCR2 engagement has been shown to induce attachment of monocytes to the endothelial layer, initiating chemotaxis, which suggests this interaction could contribute to monocyte recruitment42. Module M3 was enriched for genes related to ‘extracellular matrix organization’ and ‘collagen formation’ (Fig. 7).
Raw counts were normalized using log[CPM], and log2[fold change] values were calculated for each time point against the baseline after which coexpression modules were extracted. a, Gene set enrichment analyses showing the module activity at each time point for carriage+ and carriage− groups. Symbol size and color reflect normalized enrichment score (NES). b, Correlation with average fold change counts of all M1 genes at day 9 with paired numbers of monocytes from the volunteer’s other nostril. Individual subjects are shown, along with regression line with 95% confidence interval and Spearman correlation analysis (n = 13). c, The genes of M1 present with genes highly expressed in CD14+CD16− (578 genes), CD14+CD16+ (108 genes) and CD14−CD16+ (162 genes) 40, showing the overlapping number of genes between M1 and monocyte subsets in parentheses. The overlap for significance was analyzed using the χ2 test. d, Over-representation analysis of module M1 using gene sets from the Reactome Pathway database. e, Interaction plot for M1, with gene nodes highlighted.
Raw counts were normalized using log[CPM], and log2[fold change] values were calculated for each time point against the baseline after which coexpression modules were extracted. a, Over-representation analysis of module M3 of the control group using gene sets from the Reactome Pathway database. b, Interaction plot for M3, with gene nodes highlighted.
For LAIV, we identified six distinct coexpression modules (Supplementary Data 2), which were strongly enriched for genes related to ‘diseases associated with O-glycosylation of proteins’ (module M1), ‘immunoregulatory interactions between a lymphoid and a non-lymphoid cell’ (module M3), ‘chemokine receptors bind chemokines’ (module M4) and ‘interferon signaling’ (module M5; Fig. 8). Indeed, the hubs of module M5 are well known type I interferon–related genes, such as ISG15, OAS1, OASL, IFIT1–3 and IFITM1. Taken together, our findings reveal that a strong local antiviral response is elicited in response to LAIV infection.
Raw counts were normalized using log[CPM], and log2[fold change] values were calculated for each time point against the baseline after which coexpression modules were extracted. a, Gene set enrichment analyses showing the module activity at each time point for carriage+ and carriage− LAIV groups. b, Over-representation analysis of module M5 of the LAIV group using gene sets from the Reactome Pathway database. c, Interaction plot for M5, with gene nodes highlighted.
Discussion
This study addresses fundamental questions about the immune responses that control and clear S. pneumoniae carriage and how influenza infection can alter this control. By using a double experimental human challenge model with LAIV and S. pneumoniae, we revealed that S. pneumoniae carriage led to quick degranulation of pre-existing nasal neutrophils in the human nose and recruitment of monocytes, promoting bacterial clearance. LAIV infection impaired these immune responses following carriage. LAIV is an attenuated influenza strain, and wild-type influenza viruses might have even more pronounced effects on the host response to pneumococcus. Carriage in the absence of LAIV was associated with only limited inflammation, corroborating the view of S. pneumoniae as a commensal bacterium that can asymptomatically colonize healthy adults43. In contrast, robust proinflammatory cytokine responses were measured following LAIV at both the protein- and gene-expression level. These results provide an explanation for the published finding that LAIV increased acquisition of S. pneumoniae and carriage load14.
In addition, our findings that LAIV led to impaired blood neutrophil killing capacity and that the addition of TNF, the nasal levels of which were increased following LAIV, to neutrophils in vitro impaired their activity highlight the crucial role of neutrophils in susceptibility to secondary bacterial infection44. The association of TIGIT with this impaired neutrophil function following influenza infection warrants further investigation, as TIGIT-blocking therapeutics are currently being developed for treatment of cancer and human immunodeficiency virus45.
We identified CXCL10 as a marker for increased susceptibility to S. pneumoniae, and we propose that this should be further investigated as a potential therapeutic target for secondary bacterial infections associated with viral infections. Our data showed that individuals with higher concentrations of CXCL10 prior to S. pneumoniae inoculation had higher bacterial loads. In a previous study, children with pneumonia with viral and bacterial (predominantly pneumococcal) co-infection had increased amounts of CXCL10 compared with those of children with only viral or bacterial pneumonia, a finding that is associated with disease severity46. Data in mice suggest that CXCL10 plays a direct role during pneumonia. Mice with genetic ablation of CXCR3, the receptor for CXCL10, CXCL9 and CXCL11, showed increased survival, decreased lung inflammation and less invasion following infection, depending on the pneumococcal inoculation strain used47. Moreover, addition of exogenous CXCL10 prior to infection of mice with influenza virus or respiratory syncytial virus increased pneumonia severity48.
Our results support previous findings from mouse models showing that CCL2 signaling and monocyte recruitment are key mediators of pneumococcal carriage clearance16. However, contrary to key mechanisms described in mouse models, we did not observe any production of IL-17A or neutrophil recruitment to the nose following carriage or associated with carriage clearance15,16,17, underlining the importance of confirming mouse findings with human data.
One limitation of this study is that only one pneumococcal serotype 6B isolate was used; future studies using other isolates with a more or less invasive phenotype will be able to address how generalizable these findings are across pneumococcal isolates. Nonetheless, the observation that carriage load and duration decline in parallel for all serotypes following repeated exposure suggests that immunological control of newly acquired S. pneumoniae is mediated by similar mechanisms independent of the colonizing serotype49.
In conclusion, this study highlights the importance of innate immunity—which was impaired by pre-existing viral infections—in the control of carriage load and clearance of S. pneumoniae. Secondary bacterial infection following viral respiratory tract infection creates a large disease burden worldwide, and disrupting viral–bacterial synergy through host-directed therapy could prove an attractive addition to current therapeutic and vaccination options50.
Methods
Study design and sample collection
Healthy adult volunteers were 1:1 randomized to receive either intranasal LAIV (2015/2016 Fluenz Tetra or FluMist Tetra, AstraZeneca) or intramuscular quadrivalent inactivated influenza vaccination (Fluarix Tetra, GlaxoSmithKline), as described previously14. The control group also received a nasal saline spray, while the LAIV group also received an intramuscular saline injection. Three days post-vaccination, all subjects were inoculated with 80,000 colony-forming units (CFU) per nostril of S. pneumoniae 6B type as described previously6,51. Nasal microbiopsies (ASL Rhino-Pro, Arlington Scientific) and nasal lining fluid samples (Nasosorption, Hunt Developments) were collected and stored at −80 °C as previously described52.
Clinical trial details
The double-blinded, randomized clinical LAIV-EHPC (experimental human pneumococcal carriage) trial was registered on EudraCT (number 2014-004634-26) on 28 April 2015 and ISRCTN (number 16993271) on 2 September 2015 and was co-sponsored by the Royal Liverpool University Hospital and the Liverpool School of Tropical Medicine. Key eligibility criteria included capacity to give informed consent, no immunocompromised state or contact with susceptible individuals, no pneumococcal or influenza vaccine or infection in the last 2 years and not having taken part in EHPC studies in the past 3 years. The primary endpoint was the occurrence of pneumococcal colonization as determined by the presence of pneumococcus in nasal wash samples at any time point post-inoculation up to and including day 29, detected using classical microbiology or lytA quantitative PCR as previously described6,51,53. In this study, 130 volunteers were inoculated with pneumococcus, giving an 80% power to identify a 50% increase in carriage acquisition. Of 130 vaccinated volunteers, 5 were natural pneumococcal carriers (2 in the LAIV arm and 3 in the control arm) and were excluded from further analysis. Another 8 subjects in the LAIV arm were excluded following a systematic LAIV dispensing error by a single practitioner, as recommended by the trial steering group. This resulted in final totals of 55 subjects analyzed in the LAIV arm and 62 subjects in the control arm. Key secondary endpoints included the load of pneumococcal colonization in nasal wash samples at each time point following pneumococcal inoculation (days 2, 7, 9, 14, 22 and 29), detected using classical microbiology; the area under the curve of pneumococcal colonization load following pneumococcal inoculation (days 2, 7, 9, 14, 22 and 29), detected using classical microbiology or by molecular methods (lytA); and the immunological mechanisms associated with altered susceptibility to pneumococcus following LAIV. The outcomes reported in this manuscript were a priori included in the study protocol.
Ethics statement
All volunteers gave written informed consent, and research was conducted in compliance with all relevant ethical regulations. Ethical approval was given by the East Liverpool National Health Service Research and Ethics Committee/Liverpool School of Tropical Medicine Research and Ethics Committee, reference numbers 15/NW/0146 and 14/NW/1460 and Human Tissue Authority licensing number 12548.
Flow cytometry analysis
Immunophenotyping of nasal cells obtained by curettes was performed as previously described52. In brief, cells were dislodged from curettes and stained with LIVE/DEAD Fixable Violet Dead Cell Stain (Thermo Fisher Scientific) and an antibody cocktail containing Epcam-PE (9C4; BioLegend), HLADR-PECy7 (L243; BioLegend), CD16-APC (3G8; BioLegend), CD66b-FITC (G10F5, BioLegend), CD3-APCH7 (SK7; BD Biosciences), CD14-PercpCy5.5 (MφP9, BD Biosciences) and CD45-PACOrange (HI30, Thermo Fisher Scientific). Whole blood was stained for 15 min at room temperature with TIGIT-PECy7 (A15153G, BioLegend) and CD16-APC, followed by 2 × 10 min incubation steps with FACSLysis buffer (BD Biosciences) to remove erythrocytes. Samples were acquired on an LSR II flow cytometer (BD Biosciences) and analyzed using Flowjo X (Treestar). Fluorescent minus one controls for each of the included antibodies were used to validate results. For the LAIV and control cohorts, but not the additional validation cohort (Supplementary Fig. 5c), 84 of 553 samples (15.2%) had less than 500 immune cells or 250 epithelial cells and were excluded from further analysis.
Neutrophil opsonophagocytic killing
Neutrophil killing capacity was evaluated as previously described with minor modifications54. Briefly, neutrophils were isolated through density-gradient centrifugation, followed by 45 min incubation with serotype 6B pneumococci (inoculation strain, multiplicity of infection 100:1), baby rabbit complement (Mast Group) and human intravenous immunoglobulin (Gamunex; Grifols). In some experiments, recombinant TNF or CXCL10 (Bio-Techne) was added.
Luminex analysis of nasal lining fluid or stimulated nasal cells
Nasal cells collected in RPMI medium containing 1% penicillin–streptomycin–neomycin (Thermo Fisher Scientific) and 10% heat-inactivated FBS (Thermo Fisher Scientific) were incubated with 50 μg ml−1 DNase I (Sigma Aldrich) at room temperature for 20 min and filtered over a 70 μm filter (Thermo Fisher Scientific). Cells were spun down at 440g for 5 min, resuspended, counted and incubated at 250,000 cells ml−1 in 96-well or 384-well plates (Thermo Fisher Scientific). Heat-killed S. pneumoniae inoculation strain was added at a concentration of 5 µg ml−1 of total protein (corresponding to 4.3 × 107 CFU ml−1), and cells were incubated for 18 h. Bacterial protein concentration was measured by Bradford assay, using BSA as standard, and titration experiments were performed to determine dose. Supernatant was collected and stored at −80 °C until analysis. For nasosorption filters, cytokines were eluted from stored filters using 100 μl of assay buffer (Thermo Fisher Scientific) by centrifugation, then the eluate was cleared by further centrifugation at 16,000g. Prior to analysis, samples were centrifuged for 10 min at 16,000g to clear samples. These were acquired on an LX200 using a 30-plex magnetic human Luminex cytokine kit (Thermo Fisher Scientific) and analyzed with xPonent3.1 software following the manufacturer’s instructions. Samples were analyzed in duplicate, and nasosorption samples with a coefficient of variation (CV) of >25% were excluded.
RNA extraction and sequencing
Nasal cells were collected in RNALater (Thermo Fisher Scientific) at −80 °C until extraction. Extraction was performed using the RNEasy Micro Kit (Qiagen) with on-column DNA digestion. Extracted RNA was quantified using a Qubit (Thermo Fisher Scientific). Sample integrity assessment (Bioanalyzer; Agilent), library preparation and RNA sequencing (Illumina HiSeq4000, 20 million reads, 100 paired-end reads) were performed at the Beijing Genome Institute.
NanoString
Purified blood neutrophils were stored in RLT buffer (Qiagen) with 1% 2-mercaptoethanol (Sigma Aldrich) at −80 °C until RNA extraction as above. The single-cell immunology v2 kit (NanoString) was used with 20 pre-amp cycles for all samples. Hybridized samples were prepared on a Prep Station and scanned on a nCounter MAX (NanoString). Raw counts were analyzed using DESeq2 with internal normalization, which gave lower variance than normalizing to included housekeeping genes. DEGs were identified using a model matrix correcting for repeated individual measurements.
RNA sequencing analysis
Quality control of raw sequencing data was done using FastQC. Mapping to a human reference genome assembly (GRCh38) was done using STAR 2.5.0a55. Read counts from the resulting binary alignment map files were obtained with featureCounts using a general transfer format gene annotation from the Ensembl database56,57. The R/Bioconductor package DESeq2 was used to identify differentially expressed genes among the samples, after removing absent features (zero counts in more than 75% of samples)58. Genes with a false-discovery rate value of <0.1 and an absolute fold change of >1.5 were identified as differentially expressed.
Coexpression analysis
For coexpression analysis, counts were normalized using log[counts per million (CPM)], and the log2[fold change] was calculated for each time point in a subject-wise manner. The coexpression analysis was performed separately for each group (control and LAIV) using the CEMiTool package developed by our group and available at Bioconductor (https://bioconductor.org/packages/release/bioc/html/CEMiTool.html)39. This package unifies the discovery and the analysis of coexpression gene modules, evaluating whether modules contain genes that are over-represented by specific pathways or that are altered in a specific sample group. A P-value = 0.05 was applied for filtering genes with low expression levels.
Statistical analysis
All experiments were performed randomized and blinded. Two-tailed statistical tests were used throughout the study. When log-normalized data were not normally distributed, non-parametric tests were performed and multiple-correction testing (Benjamini-Hochberg) was applied for gene expression and Luminex analysis.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Raw RNA sequencing data have been deposited in the Gene Expression Omnibus repository, accession number GSE117580. All other underlying data are provided in the manuscript.
References
Liu, L. et al. Global, regional, and national causes of under-5 mortality in 2000–15: an updated systematic analysis with implications for the Sustainable Development Goals. Lancet 388, 3027–3035 (2016).
Morens, D. M., Taubenberger, J. K. & Fauci, A. S. Predominant role of bacterial pneumonia as a cause of death in pandemic influenza: implications for pandemic influenza preparedness. J. Infect. Dis. 198, 962–970 (2008).
Goldblatt, D. et al. Antibody responses to nasopharyngeal carriage of Streptococcus pneumoniae in adults: a longitudinal household study. J. Infect. Dis. 192, 387–393 (2005).
Simell, B. et al. The fundamental link between pneumococcal carriage and disease. Expert Rev. Vaccines 11, 841–855 (2012).
Melegaro, A., Gay, N. J. & Medley, G. F. Estimating the transmission parameters of pneumococcal carriage in households. Epidemiol. Infect. 132, 433–441 (2004).
Ferreira, D. M. et al. Controlled human infection and rechallenge with Streptococcus pneumoniae reveals the protective efficacy of carriage in healthy adults. Am. J. Respir. Crit. Care Med. 187, 855–864 (2013).
McCool, T. L., Cate, T. R., Moy, G. & Weiser, J. N. The immune response to pneumococcal proteins during experimental human carriage. J. Exp. Med. 195, 359–365 (2002).
Mina, M. J. & Klugman, K. P. The role of influenza in the severity and transmission of respiratory bacterial disease. Lancet Respir. Med. 2, 750–763 (2014).
Alpkvist, H. et al. Clinical and microbiological factors associated with high nasopharyngeal pneumococcal density in patients with pneumococcal pneumonia. PLoS ONE 10, e0140112 (2015).
Wolter, N. et al. High nasopharyngeal pneumococcal density, increased by viral coinfection, is associated with invasive pneumococcal pneumonia. J. Infect. Dis. 210, 1649–1657 (2014).
Albrich, W. C. et al. Pneumococcal colonisation density: a new marker for disease severity in HIV-infected adults with pneumonia. BMJ Open 4, e005953 (2014).
Thors, V. et al. The effects of live attenuated influenza vaccine on nasopharyngeal bacteria in healthy 2 to 4 year olds. A randomized controlled trial. Am. J. Respir. Crit. Care Med. 193, 1401–1409 (2016).
Mina, M. J., McCullers, J. A. & Klugman, K. P. Live attenuated influenza vaccine enhances colonization of Streptococcus pneumoniae and Staphylococcus aureus in mice. mBio 5, e01040-13 (2014).
Rylance, J. et al. Effect of live attenuated influenza vaccine on pneumococcal carriage. bioRxiv https://doi.org/10.1101/343319 (2018).
Lu, Y. J. et al. Interleukin-17A mediates acquired immunity to pneumococcal colonization. PLoS Pathog. 4, e1000159 (2008).
Zhang, Z., Clarke, T. B. & Weiser, J. N. Cellular effectors mediating Th17-dependent clearance of pneumococcal colonization in mice. J. Clin. Investig. 119, 1899–1909 (2009).
Lu, Y. J. et al. GMP-grade pneumococcal whole-cell vaccine injected subcutaneously protects mice from nasopharyngeal colonization and fatal aspiration-sepsis. Vaccine 28, 7468–7475 (2010).
Nakamura, S., Davis, K. M. & Weiser, J. N. Synergistic stimulation of type I interferons during influenza virus coinfection promotes Streptococcus pneumoniae colonization in mice. J. Clin. Investig. 121, 3657–3665 (2011).
Sun, K. & Metzger, D. W. Inhibition of pulmonary antibacterial defense by interferon-γ during recovery from influenza infection. Nat. Med 14, 558–564 (2008).
Li, W., Moltedo, B. & Moran, T. M. Type I interferon induction during influenza virus infection increases susceptibility to secondary Streptococcus pneumoniae infection by negative regulation of γδ T cells. J. Virol. 86, 12304–12312 (2012).
Jochems, S. P., Weiser, J. N., Malley, R. & Ferreira, D. M. The immunological mechanisms that control pneumococcal carriage. PLoS Pathog. 13, e1006665 (2017).
Querec, T. D. et al. Systems biology approach predicts immunogenicity of the yellow fever vaccine in humans. Nat. Immunol. 10, 116–125 (2009).
Oh, J. Z. et al. TLR5-mediated sensing of gut microbiota is necessary for antibody responses to seasonal influenza vaccination. Immunity 41, 478–492 (2014).
Li, S. et al. Metabolic phenotypes of response to vaccination in humans. Cell 169, 862–877 (2017).
Kazmin, D. et al. Systems analysis of protective immune responses to RTS,S malaria vaccination in humans. Proc. Natl Acad. Sci. USA 114, 2425–2430 (2017).
Nakaya, H. I. et al. Systems biology of vaccination for seasonal influenza in humans. Nat. Immunol. 12, 786–795 (2011).
Nakaya, H. I. et al. Systems analysis of immunity to influenza vaccination across multiple years and in diverse populations reveals shared molecular signatures. Immunity 43, 1186–1198 (2015).
Li, S. et al. Molecular signatures of antibody responses derived from a systems biology study of five human vaccines. Nat. Immunol. 15, 195–204 (2014).
Glennie, S. et al. Modulation of nasopharyngeal innate defenses by viral coinfection predisposes individuals to experimental pneumococcal carriage. Mucosal Immunol 9, 56–67 (2015).
Karppinen, S. et al. Acquisition and transmission of Streptococcus pneumoniae are facilitated during rhinovirus infection in families with children. Am. J. Respir. Crit. Care Med. 196, 1172–1180 (2017).
Segal, A. W. How neutrophils kill microbes. Annu. Rev. Immunol. 23, 197–223 (2005).
Craft, A. W., Reid, M. M. & Low, W. T. Effect of virus infections on polymorph function in children. Br. Med. J. 1, 1570 (1976).
Yuasa, T., Ohno, S., Kehrl, J. H. & Kyriakis, J. M. Tumor necrosis factor signaling to stress-activated protein kinase (SAPK)/Jun NH2-terminal kinase (JNK) and p38. Germinal center kinase couples TRAF2 to mitogen-activated protein kinase/ERK kinase kinase 1 and SAPK while receptor interacting protein associates with a mitogen-activated protein kinase kinase kinase upstream of MKK6 and p38. J. Biol. Chem. 273, 22681–22692 (1998).
Anderson, A. C., Joller, N. & Kuchroo, V. K. Lag-3, Tim-3, and TIGIT: co-inhibitory receptors with specialized functions in immune regulation. Immunity 44, 989–1004 (2016).
Wu, W. et al. RIG-I and TLR3 are both required for maximum interferon induction by influenza virus in human lung alveolar epithelial cells. Virology 482, 181–188 (2015).
Malley, R. et al. Recognition of pneumolysin by Toll-like receptor 4 confers resistance to pneumococcal infection. Proc. Natl Acad. Sci. USA 100, 1966–1971 (2003).
Paixao, L. et al. Host glycan sugar-specific pathways in Streptococcus pneumonia: galactose as a key sugar in colonisation and infection. PLoS ONE 10, e0121042 (2015). correction 10, e0127483 (2015).
Siegel, S. J., Roche, A. M. & Weiser, J. N. Influenza promotes pneumococcal growth during coinfection by providing host sialylated substrates as a nutrient source. Cell Host Microbe 16, 55–67 (2014).
Russo, P. S. T. et al. CEMiTool: a Bioconductor package for performing comprehensive modular co-expression analyses. BMC Bioinformatics 19, 56 (2018).
Kwissa, M. et al. Dengue virus infection induces expansion of a CD14+CD16+ monocyte population that stimulates plasmablast differentiation. Cell Host Microbe 16, 115–127 (2014).
Parker, D. et al. Streptococcus pneumoniae DNA initiates type I interferon signaling in the respiratory tract. mBio 2, e00016-00011 (2011).
Gerszten, R. E. et al. MCP-1 and IL-8 trigger firm adhesion of monocytes to vascular endothelium under flow conditions. Nature 398, 718–723 (1999).
Weiser, J. N. The pneumococcus: why a commensal misbehaves. J. Mol. Med. 88, 97–102 (2010).
Rynda-Apple, A., Robinson, K. M. & Alcorn, J. F. Influenza and bacterial superinfection: illuminating the immunologic mechanisms of disease. Infect. Immun. 83, 3764–3770 (2015).
Cox, M. A., Nechanitzky, R. & Mak, T. W. Check point inhibitors as therapies for infectious diseases. Curr. Opin. Immunol. 48, 61–67 (2017).
Hoffmann, J. et al. Viral and bacterial co-infection in severe pneumonia triggers innate immune responses and specifically enhances IP-10: a translational study. Sci. Rep. 6, 38532 (2016).
Seyoum, B., Yano, M. & Pirofski, L. A. The innate immune response to Streptococcus pneumoniae in the lung depends on serotype and host response. Vaccine 29, 8002–8011 (2011).
Luo, H., Wang, D., Che, H. L., Zhao, Y. & Jin, H. Pathological observations of lung inflammation after administration of IP-10 in influenza virus- and respiratory syncytial virus-infected mice. Exp. Ther. Med. 3, 76–79 (2012).
Hogberg, L. et al. Age- and serogroup-related differences in observed durations of nasopharyngeal carriage of penicillin-resistant pneumococci. J. Clin. Microbiol. 45, 948–952 (2007).
Madhi, S. A., Klugman, K. P. & Vaccine Trialist Group. A role for Streptococcus pneumoniae in virus-associated pneumonia. Nat. Med. 10, 811–813 (2004).
Gritzfeld, J. F. et al. Experimental human pneumococcal carriage. J. Vis. Exp. 72, e50115 (2013).
Jochems, S. P. et al. Novel analysis of immune cells from nasal microbiopsy demonstrates reliable, reproducible data for immune populations, and superior cytokine detection compared to nasal wash. PLoS ONE 12, e0169805 (2017).
Gritzfeld, J. F. et al. Density and duration of experimental human pneumococcal carriage. Clin. Microbiol. Infect. 20, O1145–O1151 (2014).
Morton, B. et al. Augmented passive immunotherapy with P4 peptide improves phagocyte activity in severe sepsis. Shock 46, 635–641 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Yates, A. et al. Ensembl 2016. Nucleic Acids Res. 44, D710–D716 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
S.B.G. and D.M.F. are supported by the Medical Research Council (grant MR/M011569/1), the Bill and Melinda Gates Foundation (grant OPP1117728) and the National Institute for Health Research Local Comprehensive Research Network. Flow cytometric acquisition was performed on a BD LSR II funded by a Wellcome Trust Multi-User Equipment Grant (104936/Z/14/Z). S.P.J. received support from the Royal Society of Tropical Medicine and Hygiene for NanoString analysis. H.I.N. is supported by the São Paulo Research Foundation (FAPESP; grants 2013/08216-2 and 2012/19278-6). We thank all volunteers for participating in this study, and R. Robinson, C. Lowe, L. Lazarova, K. Piddock and I. Wheeler for clinical support. We also acknowledge M. Mina for his input in study design.
Author information
Authors and Affiliations
Contributions
S.P.J. contributed to conceiving, designing, performing and analyzing experiments and writing the paper. F.M. and H.I.N. contributed to analyzing experiments and writing the paper. B.F.C., M.H., E.M., E.S., J.F.G., C.S., J.Reiné, S.P., E.N., E.L.G., W.A.A.d.S.P. and D.B. contributed to conducting and analyzing experiments. A.H.-W., H.H., C.H., H.A., S.Z., V.C., J.Rylance and S.B.G. contributed to sample collection and/or designing the study. D.M.F. contributed to conceiving, designing and analyzing experiments, designing the study and writing the paper. All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Pneumococcal load in the LAIV and control cohorts.
Mean and standard error of log-transformed carriage load (colony-forming units (CFU)/mL nasal wash) for carriage+ volunteers (defined by detection of S. pneumoniae at any timepoint) are shown for the control carriage+ group (green, n = 24 subjects) and LAIV carriage+ group (orange, n = 25). Load for samples with undetectable pneumococcal load was set at 0.1 CFU/mL nasal wash.
Supplementary Figure 2 CXCL10 concentrations are increased by virus infection.
(a), Concentrations of CXCL10 at day 0 in the LAIV cohort, measured by Luminex in nasosorption (n = 77). (b), Concentrations of CXCL10 at day 5 to S. pneumoniae inoculation in a cohort with known viral URT state measured by ELISA (n = 82). Oropharyngeal swabs collected 5 d before S. pneumoniae inoculation were assessed for a viral multiplex PCR panel for detection of influenza A and B (n = 0), respiratory syncytial virus (n = 2), human metapneumovirus (n = 0), human rhinovirus (n = 12), parainfluenza viruses 1–4 (n = 1), and coronaviruses OC43, NL63, 229E, and HKU1 (n = 5). Individual volunteers are shown and box plots depict median and interquartile ranges, with whiskers extending to 1.5 × interquartile range or maximum value, per group. *P = 0.017, **P = 6.2 × 10–5 by two-tailed Mann–Whitney test.
Supplementary Figure 3 Nasal granulocyte numbers over time and baseline immune cell composition.
(a), Numbers of granulocytes, predominantly neutrophils based on CD16 expression, at the nasal mucosa. Median and interquartile range are shown for cell numbers normalized to numbers of epithelial cells for control carriage+ (n = 24), control carriage– (n = 37), LAIV carriage+ (n = 25) and LAIV carriage– (n = 30) groups. The x axis shows days relative to inoculation. (b), Absolute numbers of cells observed per nasal sample (median and IQR are shown for 117 subjects at baseline).
Supplementary Figure 4 Flow cytometry gating strategy and longitudinal levels of nasal T cells and monocytes.
(a), Gating strategy of nasal cells for one representative volunteer. (b), Numbers of T cells at the nasal mucosa. Median and interquartile range are shown for cell numbers normalized to numbers of epithelial cells for control carriage+ (n = 24), control carriage– (n = 37), LAIV carriage+ (n = 25) and LAIV carriage– (n = 30) groups. The x axis shows days relative to inoculation. (c), Median and interquartile range of monocyte numbers in the control group over time with carriage detected by classical microbiology (carriage+, n = 24), molecular methods only (lytA+, n = 16) or not at all (carriage–, n = 22). *P = 0.038 at day 2 and P = 0.030 at day 29, **P = 0.002 by two-tailed Wilcoxon paired non-parametric test.
Supplementary Figure 5 Monocyte recruitment associates with CCL2 concentration and is reproducible in an independent volunteer cohort.
(a), Spearman’s correlation between the 30 measured cytokines and nasal monocyte numbers for each timepoint (n = 73). The length of the bar depicts the rho value, cyan bars represent cytokines that correlate significantly at all days, red bars associate for a specific day, and transparent red bars are not significantly associated for that day. (b), Monocyte numbers over time in those volunteers who most upregulate CCL2, IFN-γ, IL-6 or TNF at day 2 (top quartile FC induction, n = 17 versus below top quartile, n = 56) or not. *P = 0.028 by two-tailed Mann–Whitney test of monocyte number area under curve of inducers and non-inducers. (c), Numbers of monocytes and CCL2 in a second, independent cohort of seven carriers without any vaccination. Symbols represent individual subjects, with color corresponding to day relative to S. pneumoniae inoculation. The effects of CCL2 on monocyte numbers by generalized linear regression analysis are shown, correcting for repeated individual measurements.
Supplementary Figure 6 Nasal cell responses to in vitro stimulation with pneumococcus.
Whole nasal cells were collected 29 d after S. pneumoniae inoculation and stimulated for 18 h with heat-killed pneumococcus (n = 48). Supernatant was collected and the concentrations of 30 cytokines were measured by Luminex. Four cytokines were not measured above the limit of detection (IL-2, IL-4, MIG and eotaxin), and these were excluded from analysis. (a), Responses for all measured cytokines for the four groups are shown. (b), Production in carriage– volunteers subdivided into those who had very low carriage loads, detectable only by molecular methods (lytA+, n = 9), or not at all (carriage–, n = 5). The dotted line indicates a twofold increase over the unstimulated control. The median and interquartile range of FC to unstimulated are shown An asterisk indicates significantly induced cytokines (shown in red) (FC > 2, q < 0.05, two-tailed paired Wilcoxon test followed by Benjamini–Hochberg correction).
Supplementary Figure 7 Immune mechanisms associating with control of S. pneumoniae carriage and their disruption by LAIV.
Carriage in the absence of influenza leads to quick degranulation of nasal neutrophils followed by recruitment of monocytes to the nose, associating with the start of clearance. Influenza infection leads to inflammation, impairing this innate control of carriage.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 2 and 3
Supplementary Data 1
CEMiTool report of control group
Supplementary Data 2
CEMiTool report of LAIV group
Supplementary Table 1
Cytokine induction following LAIV or control vaccination
Supplementary Table 4
List of differentially expressed genes
Supplementary Table 5
List of pathways
Rights and permissions
About this article
Cite this article
Jochems, S.P., Marcon, F., Carniel, B.F. et al. Inflammation induced by influenza virus impairs human innate immune control of pneumococcus. Nat Immunol 19, 1299–1308 (2018). https://doi.org/10.1038/s41590-018-0231-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0231-y
This article is cited by
-
Exploring the immune-inflammatory mechanism of Maxing Shigan Decoction in treating influenza virus A-induced pneumonia based on an integrated strategy of single-cell transcriptomics and systems biology
European Journal of Medical Research (2024)
-
NEDD4 Regulated Pyroptosis Occurred from Co-infection between Influenza A Virus and Streptococcus pneumoniae
Journal of Microbiology (2023)
-
A novel Streptococcus pneumoniae human challenge model demonstrates Treg lymphocyte recruitment to the infection site
Scientific Reports (2022)
-
Asplenia and spleen hypofunction
Nature Reviews Disease Primers (2022)
-
Innate immune responses at the asymptomatic stage of influenza A viral infections of Streptococcus pneumoniae colonized and non-colonized mice
Scientific Reports (2021)