Abstract
The activation of natural killer (NK) cells depends on a change in the balance of signals from inhibitory and activating receptors. The activation threshold values of NK cells are thought to be set by engagement of inhibitory receptors during development. Here, we found that the activating receptor NKG2D specifically set the activation threshold for the activating receptor NCR1 through a process that required the adaptor DAP12. As a result, NKGD2-deficient (Klrk1−/−) mice controlled tumors and cytomegalovirus infection better than wild-type controls through the NCR1-induced production of the cytokine IFN-γ. Expression of NKG2D before the immature NK cell stage increased expression of the adaptor CD3ζ. Reduced expression of CD3ζ in Klrk1−/− mice was associated with enhanced signal transduction through NCR1, and CD3ζ deficiency resulted in hyper-responsiveness to stimulation via NCR1. Thus, an activating receptor developmentally set the activity of another activating receptor on NK cells and determined NK cell reactivity to cellular threats.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. The NCBI SRA accession codes are: SRX4548789, SRX4548788, SRX4548787, SRX4548786, SRX4548785, SRX4548784, SRX4548783, SRX4548782, SRX4548781.
References
Spits, H., Bernink, J. H. & Lanier, L. NK cells and type 1 innate lymphoid cells: partners in host defense. Nat. Immunol. 17, 758–764 (2016).
Elliott, J. M. & Yokoyama, W. M. Unifying concepts of MHC-dependent natural killer cell education. Trends Immunol. 32, 364–372 (2011).
Long, E. O., Kim, H. S., Liu, D., Peterson, M. E. & Rajagopalan, S. Controlling natural killer cell responses: integration of signals for activation and inhibition. Annu. Rev. Immunol. 31, 227–258 (2013).
Koch, J., Steinle, A., Watzl, C. & Mandelboim, O. Activating natural cytotoxicity receptors of natural killer cells in cancer and infection. Trends Immunol. 34, 182–191 (2013).
Zafirova, B., Wensveen, F. M., Gulin, M. & Polić, B. Regulation of immune cell function and differentiation by the NKG2D receptor. Cell. Mol. Life Sci. 68, 3519–3529 (2011).
Jelenčić, V., Lenartić, M., Wensveen, F. M. & Polić, B. NKG2D: A versatile player in the immune system. Immunol. Lett. 189, 48–53 (2017).
Goh, W. & Huntington, N. D. Regulation of murine natural killer cell development. Front. Immunol. 8, 130 (2017).
Huntington, N. D., Vosshenrich, C. A. & Di Santo, J. P. Developmental pathways that generate natural-killer-cell diversity in mice and humans. Nat. Rev. Immunol. 7, 703–714 (2007).
Zafirova, B. et al. Altered NK cell development and enhanced NK cell-mediated resistance to mouse cytomegalovirus in NKG2D-deficient mice. Immunity 31, 270–282 (2009).
Guerra, N. et al. NKG2D-deficient mice are defective in tumor surveillance in models of spontaneous malignancy. Immunity 28, 571–580 (2008).
Sheppard, S. et al. Characterization of a novel NKG2D and NKp46 double-mutant mouse reveals subtle variations in the NK cell repertoire. Blood 121, 5025–5033 (2013).
Gilfillan, S., Ho, E. L., Cella, M., Yokoyama, W. M. & Colonna, M. NKG2D recruits two distinct adapters to trigger NK cell activation and costimulation. Nat. Immunol. 3, 1150–1155 (2002).
Lanier, L. L. DAP10- and DAP12-associated receptors in innate immunity. Immunol. Rev. 227, 150–160 (2009).
Hamerman, J. A. & Lanier, L. L. Inhibition of immune responses by ITAM-bearing receptors. Sci. STKE 2006, re1 (2006).
Arase, H. et al. Negative regulation of expression and function of Fc gamma RIII by CD3 zeta in murine NK cells. J. Immunol. 166, 21–25 (2001).
Utsuyama, M. & Hirokawa, K. Radiation-induced-thymic lymphoma occurs in young, but not in old mice. Exp. Mol. Pathol. 74, 319–325 (2003).
Overwijk, W. W. & Restifo, N. P. B16 as a mouse model for human melanoma. Curr. Protoc. Immunol. 39, 20.1.1–20.1.29 (2001).
Lenartić, M. et al. NKG2D promotes B1a cell development and protection against bacterial infection. J. Immunol. 198, 1531–1542 (2017).
Schuster, I. S., Coudert, J. D., Andoniou, C. E. & Degli-Esposti, M. A. “Natural regulators”: NK cells as modulators of T cell immunity. Front. Immunol. 7, 235 (2016).
Soderquest, K. et al. Cutting edge: CD8+ T cell priming in the absence of NK cells leads to enhanced memory responses. J. Immunol. 186, 3304–3308 (2011).
Takeda, K. et al. IFN-γ production by lung NK cells is critical for the natural resistance to pulmonary metastasis of B16 melanoma in mice. J. Leukoc. Biol. 90, 777–785 (2011).
Dong, Z. et al. The adaptor SAP controls NK cell activation by regulating the enzymes Vav-1 and SHIP-1 and by enhancing conjugates with target cells. Immunity 36, 974–985 (2012).
Glasner, A. et al. Recognition and prevention of tumor metastasis by the NK receptor NKp46/NCR1. J. Immunol. 188, 2509–2515 (2012).
Lakshmikanth, T. et al. NCRs and DNAM-1 mediate NK cell recognition and lysis of human and mouse melanoma cell lines in vitro and in vivo. J. Clin. Invest. 119, 1251–1263 (2009).
Wensveen, F. M. et al. NK cells link obesity-induced adipose stress to inflammation and insulin resistance. Nat. Immunol. 16, 376–385 (2015).
Davis, A. H., Guseva, N. V., Ball, B. L. & Heusel, J. W. Characterization of murine cytomegalovirus m157 from infected cells and identification of critical residues mediating recognition by the NK cell receptor Ly49H. J. Immunol. 181, 265–275 (2008).
Coudert, J. D., Scarpellino, L., Gros, F., Vivier, E. & Held, W. Sustained NKG2D engagement induces cross-tolerance of multiple distinct NK cell activation pathways. Blood 111, 3571–3578 (2008).
Deng, W. et al. Antitumor immunity. A shed NKG2D ligand that promotes natural killer cell activation and tumor rejection. Science 348, 136–139 (2015).
Eckelhart, E. et al. A novel Ncr1-Cre mouse reveals the essential role of STAT5 for NK-cell survival and development. Blood 117, 1565–1573 (2011).
Schwenk, F., Baron, U. & Rajewsky, K. A cre-transgenic mouse strain for the ubiquitous deletion of loxP-flanked gene segments including deletion in germ cells. Nucleic Acids Res. 23, 5080–5081 (1995).
Challen, G. A., Boles, N., Lin, K. K. & Goodell, M. A. Mouse hematopoietic stem cell identification and analysis. Cytometry. A. 75, 14–24 (2009).
Karo, J. M., Schatz, D. G. & Sun, J. C. The RAG recombinase dictates functional heterogeneity and cellular fitness in natural killer cells. Cell 159, 94–107 (2014).
Viant, C. et al. SHP-1-mediated inhibitory signals promote responsiveness and anti-tumour functions of natural killer cells. Nat. Commun. 5, 5108 (2014).
Wu, N. et al. A hematopoietic cell-driven mechanism involving SLAMF6 receptor, SAP adaptors and SHP-1 phosphatase regulates NK cell education. Nat. Immunol. 17, 387–396 (2016).
Chen, L. et al. Discovery of a novel shp2 protein tyrosine phosphatase inhibitor. Mol. Pharmacol. 70, 562–570 (2006).
Mandelboim, O. & Porgador, A. NKp46. Int. J. Biochem. Cell. Biol. 33, 1147–1150 (2001).
Myers, M. D., Dragone, L. L. & Weiss, A. Src-like adaptor protein down-regulates T cell receptor (TCR)-CD3 expression by targeting TCRzeta for degradation. J. Cell. Biol. 170, 285–294 (2005).
Goodfellow, H. S. et al. The catalytic activity of the kinase ZAP-70 mediates basal signaling and negative feedback of the T cell receptor pathway. Sci. Signal. 8, ra49 (2015).
Hamerman, J. A., Tchao, N. K., Lowell, C. A. & Lanier, L. L. Enhanced Toll-like receptor responses in the absence of signaling adaptor DAP12. Nat. Immunol. 6, 579–586 (2005).
Sjölin, H. et al. DAP12 signaling regulates plasmacytoid dendritic cell homeostasis and down-modulates their function during viral infection. J. Immunol. 177, 2908–2916 (2006).
Mulrooney, T. J., Posch, P. E. & Hurley, C. K. DAP12 impacts trafficking and surface stability of killer immunoglobulin-like receptors on natural killer cells. J. Leukoc. Biol. 94, 301–313 (2013).
Della Chiesa, M. et al. Evidence that the KIR2DS5 gene codes for a surface receptor triggering natural killer cell function. Eur. J. Immunol. 38, 2284–2289 (2008).
Fauriat, C., Ivarsson, M. A., Ljunggren, H. G., Malmberg, K. J. & Michaëlsson, J. Education of human natural killer cells by activating killer cell immunoglobulin-like receptors. Blood 115, 1166–1174 (2010).
Carr, W. H. et al. Cutting Edge: KIR3DS1, a gene implicated in resistance to progression to AIDS, encodes a DAP12-associated receptor expressed on NK cells that triggers NK cell activation. J. Immunol. 178, 647–651 (2007).
Deng, G. M., Beltran, J., Chen, C., Terhorst, C. & Tsokos, G. C. T cell CD3ζ deficiency enables multiorgan tissue inflammation. J. Immunol. 191, 3563–3567 (2013).
Yang, M. et al. K33-linked polyubiquitination of Zap70 by Nrdp1 controls CD8(+) T cell activation. Nat. Immunol. 16, 1253–1262 (2015).
Takaki, R., Watson, S. R. & Lanier, L. L. DAP12: an adapter protein with dual functionality. Immunol. Rev. 214, 118–129 (2006).
Kiel, M. J. et al. SLAM family receptors distinguish hematopoietic stem and progenitor cells and reveal endothelial niches for stem cells. Cell 121, 1109–1121 (2005).
Wang, Y. et al. Long noncoding RNA derived from CD244 signaling epigenetically controls CD8+ T-cell immune responses in tuberculosis infection. Proc. Natl Acad. Sci. USA 112, E3883–E3892 (2015).
Acknowledgements
We thank S. Slavić Stupac, M. Samsa, E. Marinović, A. Miše and K. Mikić for technical assistance. We thank S. Jonjić for discussions and overall support to this work. We thank O. Mandelboim (Hebrew University Hadassah Medical School), M. Colonna (Washington University School of Medicine in St. Louis, St. Louis, MO, USA), M. Busslinger (Research Institute of Molecular Pathology, Vienna, Austria), A. Waisman (Institute of Molecular Biology, Mainz, Germany), D.R. Littman (NYU School of Medicine, New York, NY, USA), J. McGlade (University of Toronto, Toronto, Canada) and K. Rajewsky (MDC Center for Molecular Medicine, Berlin, Germany) for providing mouse lines. This work was supported by an EFIS-IL Short-term Fellowship grant to V.J., European Research Council under the European Union’s Seventh Framework Programme (FP/2007–2013)/ERC Grant Agreement no. 311335, Swedish Research Council, Norwegian Research Council, Swedish Foundation for Strategic Research, Wallenberg Foundation, Swedish Cancer Foundation, Swedish Childhood Cancer Foundation, as well as the Stockholm County Council and Karolinska Institutet Center for Innovative Medicine to Y.T.B., Netherlands Organization for Scientific Research (91614029), the European Commission (PCIG14-GA-2013-630827), a University of Rijeka Support grant (865.10.2101) and a Croatian Science Foundation Grant (IP-2016-06-8027) to F.M.W., a European Social Fund grant (HR.3.2.01-0263), University of Rijeka Support grant (803.10.1103) and Croatian Science Foundation grant (IP-2016-06-9306) to B.P. and the grant KK.01.1.1.01.0006, awarded to the Scientific Centre of Excellence for Virus Immunology and Vaccines and co-financed by the European Regional Development Fund.
Author information
Authors and Affiliations
Contributions
V.J. carried out most of the experiments and analyzed data. M.Š., I.K., M.L., S.M., B.L. and F.M.W. performed and analyzed experiments. B.P. directed the research. B.P., V.J. and F.M.W. designed experiments and wrote the paper. V.S. and M.P. designed and performed qPCR on NK precursors. T.D.H. and Y.T.B. designed and performed RNA-seq experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Better control of B16 melanoma by Klrk1−/− mice is NK cell and IFNγ mediated.
(a) FACS analysis of indicated organs from C57BL/6J and C57BL/6J RITL mice. (b) NKG2D ligand expression on indicated cells and 3T3-H60 cells as positive control labeled with NKG2D-hIg fusion protein (full line) or irrelevant fusion protein (dashed line). (c,g) Survival curves for C57BL/6J and Klrk1−/− mice after B16 cells i.v. injection without (c) or with CD4+ T cell depletion (g). (d) Representative x-Ray pictures of tumors from C57BL/6J or Klrk1−/− after B16 s.c. injection. (e) NKG2D expression on NKT cells from Klrk1−/− (n = 4), Klrk1fl/fl (n = 7), Klrk1fl/flCD4Cre (n = 6) mice (ANOVA, with Bonferroni’s post-test correction, *** p < 0.001). (f) Survival curves of indicated littermate mice after B16 cells i.v. injection. (h) Killing capacity of NK cells from C57BL/6J and Klrk1−/− mice towards B16-CFSE labeled after 14h at indicated effector to target ratios (n = 3). (i) Ifng mRNA expression normalized to 18S in B16 tumors isolated on 10th day from C57BL/6J and Klrk1−/− mice (data pooled from two experiments). (j) IFN-γ production by NK cells from indicated mice (n = 4) after DNAM-1 stimulation. (k) B16 cells stained with irrelevant fusion protein (dashed line), NCR1-hIg (full line) or NCR1-hIG plus anti NCR1 antibody to prevent binding of fusion protein (red line). c,f and g were analyzed by the Kaplan–Meier model followed by Log-rank (Mantel-Cox) test (n = 10 mice per group). h and j were analyzed using two-tailed unpaired t-test (ns, not significant). e, h-j show mean ± s.e.m. All graphs are representative from two independent experiments.
Supplementary Figure 2 Klrk1−/− mice show better control of both WT and Δm157 mCMV strains.
C57BL/6J and Klrk1−/− mice (n = 6 mice per goup) were infected with mCMV MW97.01 and Δm157 mCMV, which was generated on the backbone of the MW97.01 BAC. On day four after infection, viral titers in spleen were determined by plaque assay. * p > 0.05, ** p > 0.005 (two-sided Mann-Whitney test). Shown are mean ± s.e.m. Graphs are representative from two independent experiments.
Supplementary Figure 3 Deletion of NKG2D expression on NK cells.
(a) NKG2D expression was analyzed on NK cells from Klrk1−/− (n = 7), Klrk1fl/fl (n = 5), Klrk1fl/fl NCR1cre (n = 5) mice. Graph show geometric mean values (ANOVA, with Bonferroni’s post-test correction, shown mean ± s.e.m, *** p < 0.001). (b) Klrk1Δ/Δ mice were generated by crossing Klrk1fl/fl mice with Deletercre. NK cells from Klrk1Δ/Δ (n = 4) and C57BL/6J (n = 5) mice were stimulated through NCR1 and IFN-γ production was analyzed (two-tailed unpaired t-test, shown mean ± s.e.m; *P < 0.05). (c) Rag1CreEYFPStop-FloxiDTR mice were injected with PBS (-DT) or with diphtheria toxin (+DT). After 24h, immune cells in blood were analyzed. Gated is for viable cells. Shown are representative graphs from 3 (a,b) and 2 (c) independent experiments.
Supplementary Figure 4 Characterization of NK cells.
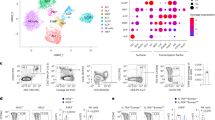
(a) Gating strategy for development of NK cells in BM. First lineage+ cells were excluded using anti CD4, CD8, Gr-1 and Ter119 antibodies. In lineage− population different stages of NK cell development were determined. Stage 1(CD122+NK1.1−CD11b−c-kit−), stage 2 (CD122+NK1.1+NCR1−CD11b−c-kit−), stage 3 (CD122+NK1.1+NCR1+CD11b−c-kit−), stage 4 (CD122+NK1.1+NCR1+CD11b−c-kit+), stage 5 (CD122+NK1.1+NCR1+CD11b+c-kit−/+), stage 6 (CD122+NK1.1+NCR1+CD11b+c-kit−). EYFP+ and EYFP− NK cells from Rag1cre Rosa26-floxed STOP-YFP (EYFPStop-Flox) Klrk1+/+ (n = 4 mice in EYFP+ group and 3 in EYFP− group) (b) or Rag1creEYFPStop-Flox Klrk1−/− (n = 4 mice per group) (c) were analyzed. Graph shows percentages of Klrg1 and CD11b positive NK cells. (d) Graph shows percentage of Ly49I+ NK cells in C57BL/6J (n = 3) and Klrk1−/− (n = 4) littermates. b-d two-tailed unpaired t-test was used; ns, not significant, *P < 0.05, **p < 0.01, *** p < 0.001. b-d shown are mean ± s.e.m.
Supplementary Figure 5 Analysis of NK cells.
(a) NK cells from Tyrobp+/+ and Tyrobp−/− littermates were stimulated through the indicated receptors and IFN-γ production was analyzed by intracellular flow cytometry (n = 5 mice per group). b-c NK cells from Stage 1 and 2 were sorted by FACS Aria using gating strategy described in Supplementary fig. 4a. Using qPCR we analyzed expression of NKG2D-S isoform (b) and DAP12 molecule (c). Graphs show relative expression to Rplp0 (n = 5 mice per group). (d) NK cells from C57BL/6J or Cd247−/− mice were stained for CD3ζ expression and analyzed by flow cytometry. Figure shows representative histograms. (e) NK cells from Tyrobp+/+ and Tyrobp−/− littermates were analyzed for expression of CD3ζ, FcRγ, Zap-70 and Syk expression. Figure shows graphs of geometric mean values for indicated molecules. As controls, isotype stained samples were used (n = 5 mice per group). (f) NK cells from Klrk1−/−, C57BL/6J, Hcst−/− and Tyrobp−/− mice were stimulated through CD16 receptor and IFN-γ production was analyzed (n = 5 mice per group). Representative data (shown are mean ± s.e.m.) from two (a-c, e) or three (d, f) independent experiments. To analyze a-c and e two-tailed unpaired t-test was used (ns, not significant, *P < 0.05, **p < 0.01, *** p < 0.001), for f ANOVA, with Bonferroni’s post-test correction was used (**p < 0.01, *** p < 0.001).
Supplementary Figure 6 Hyper-reactivity of Klrk1−/− NK cells is the consequence of reduced expression of CD3ζ.
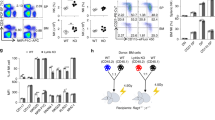
(a) Cluster analysis of genes differentially expressed in Klrk1−/− NK cells. mRNA of C57BL/6J and Klrk1−/− NK cells was analyzed by RNAseq. To identify potential interaction partners with CD3ζ and Zap-70, cluster analysis of these molecules and differentially expressed genes was performed. Unconnected nodes were excluded. (b) NK cells from Balb/c or Sla−/− mice were stained for SLAP-1 expression and analyzed by flow cytometry. The figure shows representative histograms. (c) Splenic NK cells from Balb/c (n = 10) and Sla−/− (n = 8) mice were analyzed for expression of NCR1. Figure shows graphs of geometric mean values. d-e NK cells from Tyrobp+/+ and Tyrobp−/− littermates were stimulated through the NCR1 receptor and phosphorylation of Syk (d) and PLC-γ (e) was analyzed (n = 5 mice per group). (f) NCR1 expression was analyzed on NK cells from C57BL/6J (n = 5) and cd247−/− (n = 4) mice. The graph shows geometric mean values. To analyze c-f two-tailed unpaired t-test was used (ns, not significant, *P < 0.05). Shown are mean ± s.e.m.
Supplementary information
Rights and permissions
About this article
Cite this article
Jelenčić, V., Šestan, M., Kavazović, I. et al. NK cell receptor NKG2D sets activation threshold for the NCR1 receptor early in NK cell development. Nat Immunol 19, 1083–1092 (2018). https://doi.org/10.1038/s41590-018-0209-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0209-9
This article is cited by
-
KLRK1 as a prognostic biomarker for lung adenocarcinoma cancer
Scientific Reports (2022)
-
CAR-engineered NK cells; a promising therapeutic option for treatment of hematological malignancies
Stem Cell Research & Therapy (2021)
-
It takes a village
Nature Immunology (2018)