Abstract
The sensing of microbial genetic material by leukocytes often elicits beneficial pro-inflammatory cytokines, but dysregulated responses can cause severe pathogenesis. Genome-wide association studies have linked the gene encoding phospholipase D3 (PLD3) to Alzheimer’s disease and have linked PLD4 to rheumatoid arthritis and systemic sclerosis. PLD3 and PLD4 are endolysosomal proteins whose functions are obscure. Here, PLD4-deficient mice were found to have an inflammatory disease, marked by elevated levels of interferon-γ (IFN-γ) and splenomegaly. These phenotypes were traced to altered responsiveness of PLD4-deficient dendritic cells to ligands of the single-stranded DNA sensor TLR9. Macrophages from PLD3-deficient mice also had exaggerated TLR9 responses. Although PLD4 and PLD3 were presumed to be phospholipases, we found that they are 5′ exonucleases, probably identical to spleen phosphodiesterase, that break down TLR9 ligands. Mice deficient in both PLD3 and PLD4 developed lethal liver inflammation in early life, which indicates that both enzymes are needed to regulate inflammatory cytokine responses via the degradation of nucleic acids.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Schlee, M. & Hartmann, G. Discriminating self from non-self in nucleic acid sensing. Nat. Rev. Immunol. 16, 566–580 (2016).
Barbalat, R., Ewald, S. E., Mouchess, M. L. & Barton, G. M. Nucleic acid recognition by the innate immune system. Annu. Rev. Immunol. 29, 185–214 (2011).
Rigby, R. E., Leitch, A. & Jackson, A. P. Nucleic acid-mediated inflammatory diseases. BioEssays 30, 833–842 (2008).
Al-Mayouf, S. M. et al. Loss-of-function variant in DNASE1L3 causes a familial form of systemic lupus erythematosus. Nat. Genet. 43, 1186–1188 (2011).
Napirei, M. et al. Features of systemic lupus erythematosus in Dnase1-deficient mice. Nat. Genet. 25, 177–181 (2000).
Yasutomo, K. et al. Mutation of DNASE1 in people with systemic lupus erythematosus. Nat. Genet. 28, 313–314 (2001).
Kawane, K. et al. Requirement of DNase II for definitive erythropoiesis in the mouse fetal liver. Science 292, 1546–1549 (2001).
Rodero, M. P. et al. Type I interferon-mediated autoinflammation due to DNase II deficiency. Nat. Commun. 8, 2176 (2017).
Morita, M. et al. Gene-targeted mice lacking the Trex1 (DNase III) 3′–>5′ DNA exonuclease develop inflammatory myocarditis. Mol. Cell. Biol. 24, 6719–6727 (2004).
Lee-Kirsch, M. A. et al. Mutations in the gene encoding the 3′-5′ DNA exonuclease TREX1 are associated with systemic lupus erythematosus. Nat. Genet. 39, 1065–1067 (2007).
Rice, G. et al. Heterozygous mutations in TREX1 cause familial chilblain lupus and dominant Aicardi-Goutieres syndrome. Am. J. Hum. Genet. 80, 811–815 (2007).
Ipsaro, J. J., Haase, A. D., Knott, S. R., Joshua-Tor, L. & Hannon, G. J. The structural biochemistry of Zucchini implicates it as a nuclease in piRNA biogenesis. Nature 491, 279–283 (2012).
Waite, M. The PLD superfamily: insights into catalysis. Biochim. Biophys. Acta 1439, 187–197 (1999).
McDermott, M., Wakelam, M. J. & Morris, A. J. Phospholipase D. Biochem. Cell Biol. 82, 225–253 (2004).
Otani, Y. et al. PLD$ is involved in phagocytosis of microglia: expression and localization changes of PLD4 are correlated with activation state of microglia. PLoS One 6, e27544 (2011).
Fazzari, P. et al. PLD3 gene and processing of APP. Nature 541, E1–E2 (2017).
Gonzalez, A. C. et al. Unconventional trafficking of mammalian phospholipase D3 to lysosomes. Cell Rep. 22, 1040–1053 (2018).
Terao, C. et al. PLD4 as a novel susceptibility gene for systemic sclerosis in a Japanese population. Arthritis Rheum. 65, 472–480 (2013).
Okada, Y. et al. Meta-analysis identifies nine new loci associated with rheumatoid arthritis in the Japanese population. Nat. Genet. 44, 511–516 (2012).
Chen, W. C. et al. rs2841277 (PLD4) is associated with susceptibility and rs4672495 is associated with disease activity in rheumatoid arthritis. Oncotarget 8, 64180–64190 (2017).
Cruchaga, C. et al. Rare coding variants in the phospholipase D3 gene confer risk for Alzheimer’s disease. Nature 505, 550–554 (2014).
Zhang, D. F. et al. PLD3 in Alzheimer’s disease: a modest effect as revealed by updated association and expression analyses. Mol. Neurobiol. 53, 4034–4045 (2016).
Hooli, B. V. et al. PLD3 gene variants and Alzheimer’s disease. Nature 520, E7–E8 (2015).
Heilmann, S. et al. PLD3 in non-familial Alzheimer’s disease. Nature 520, E3–E5 (2015).
Yoshikawa, F. et al. Phospholipase D family member 4, a transmembrane glycoprotein with no phospholipase D activity, expression in spleen and early postnatal microglia. PLoS One 5, e13932 (2010).
Osisami, M., Ali, W. & Frohman, M. A. A role for phospholipase D3 in myotube formation. PLoS One 7, e33341 (2012).
Bernardi, A. & Bernardi, G. Studies on acid hydrolases. IV. Isolation and characterization of spleen exonuclease. Biochim. Biophys. Acta 155, 360–370 (1968).
Hilmoe, R. J. Purification and properties of spleen phosphodiesterase. J. Biol. Chem. 235, 2117–2121 (1960).
Willenborg, D. O., Fordham, S. A., Staykova, M. A., Ramshaw, I. A. & Cowden, W. B. IFN-γ is critical to the control of murine autoimmune encephalomyelitis and regulates both in the periphery and in the target tissue: a possible role for nitric oxide. J. Immunol. 163, 5278–5286 (1999).
Behrens, E. M. et al. Repeated TLR9 stimulation results in macrophage activation syndrome-like disease in mice. J. Clin. Invest. 121, 2264–2277 (2011).
Ferlazzo, G. et al. Distinct roles of IL-12 and IL-15 in human natural killer cell activation by dendritic cells from secondary lymphoid organs. Proc. Natl Acad. Sci. USA 101, 16606–16611 (2004).
Ma, X. et al. The interleukin 12 p40 gene promoter is primed by interferon γ in monocytic cells. J. Exp. Med. 183, 147–157 (1996).
Razzell, W. E. Tissue and intracellular distribution of two phosphodiesterases. J. Biol. Chem. 236, 3028–3030 (1961).
Ipata, P. L. & Felicioli, R. A. A convenient spectrophotometric assay for phosphodiesterases, using dinucleoside-monophosphates as substrates. Eur. J. Biochem. 8, 174–179 (1969).
Krieg, A. M. CpG motifs in bacterial DNA and their immune effects. Annu. Rev. Immunol. 20, 709–760 (2002).
Chan, M. P. et al. DNase II-dependent DNA digestion is required for DNA sensing by TLR9. Nat. Commun. 6, 5853 (2015).
Tabeta, K. et al. The Unc93b1 mutation 3d disrupts exogenous antigen presentation and signaling via Toll-like receptors 3, 7 and 9. Nat. Immunol. 7, 156–164 (2006).
Pedersen, K. M., Finsen, B., Celis, J. E. & Jensen, N. A. Expression of a novel murine phospholipase D homolog coincides with late neuronal development in the forebrain. J. Biol. Chem. 273, 31494–31504 (1998).
Langenmayer, M. C. et al. Zinc deficiency-like syndrome in Fleckvieh calves: clinical and pathological findings and differentiation from bovine hereditary zinc deficiency. J. Vet. Intern. Med. 32, 853–859 (2018).
Ohto, U. et al. Toll-like receptor 9 contains two DNA binding sites that function cooperatively to promote receptor dimerization and activation. Immunity 48, 649–658 (2018).
Erdal, E., Haider, S., Rehwinkel, J., Harris, A. L. & McHugh, P. J. A prosurvival DNA damage-induced cytoplasmic interferon response is mediated by end resection factors and is limited by Trex1. Genes Dev. 31, 353–369 (2017).
Luecke, S. et al. cGAS is activated by DNA in a length-dependent manner. EMBO Rep. 18, 1707–1715 (2017).
Canna, S. W. & Behrens, E. M. Not all hemophagocytes are created equally: appreciating the heterogeneity of the hemophagocytic syndromes. Curr. Opin. Rheumatol. 24, 113–118 (2012).
Bracaglia, C. et al. Elevated circulating levels of interferon-γ and interferon-γ-induced chemokines characterise patients with macrophage activation syndrome complicating systemic juvenile idiopathic arthritis. Ann. Rheum. Dis. 76, 166–172 (2017).
Mouchess, M. L. et al. Transmembrane mutations in Toll-like receptor 9 bypass the requirement for ectodomain proteolysis and induce fatal inflammation. Immunity 35, 721–732 (2011).
Chen, J. H. et al. Pathology of the liver in familial hemophagocytic lymphohistiocytosis. Am. J. Surg. Pathol. 34, 852–867 (2010).
Sepulveda, F. E. & de Saint Basile, G. Hemophagocytic syndrome: primary forms and predisposing conditions. Curr. Opin. Immunol. 49, 20–26 (2017).
Wang, L. et al. DNA phosphorothioation is widespread and quantized in bacterial genomes. Proc. Natl Acad. Sci. USA 108, 2963–2968 (2011).
Bernardi, A. & Cantoni, G. L. Action of spleen exonuclease on transfer ribonucleic acid. J. Biol. Chem. 244, 1468–1476 (1969).
Mandecki, W. & Hayden, M. High-resolution polyacrylamide gel electrophoresis of oligonucleotides using L-histidine buffer. DNA 7, 57–62 (1988).
Slaymaker, I. M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016).
Vremec, D. & Shortman, K. The isolation and identification of murine dendritic cell populations from lymphoid tissues and their production in culture. Methods Mol. Biol. 415, 163–178 (2008).
Zhang, X., Goncalves, R. & Mosser, D.M. The isolation and characterization of murine macrophages. Curr. Prot. Immunol. (eds. Coligan, J.E. et al.) Ch. 14, Unit 14 11 (Wiley, Hoboken, NJ, USA, 2008).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Lattin, J. E. et al. Expression analysis of G protein-coupled receptors in mouse macrophages. Immunome Res. 4, 5 (2008).
Wu, C., Jin, X., Tsueng, G., Afrasiabi, C. & Su, A. I. BioGPS: building your own mash-up of gene annotations and expression profiles. Nucleic Acids Res. 44, D313–D316 (2016).
Lauterbach, H. et al. Mouse CD8α + DCs and human BDCA3 + DCs are major producers of IFN-λ in response to poly IC. J. Exp. Med. 207, 2703–2717 (2010).
Acknowledgements
We thank B. Beutler (University of Texas Southwestern Medical Center) for Unc93b3d/3d mice; Y. Doyon (Centre Hospitalier Universitaire de Québec) for plasmid eSpCas9(1.1)51; and S. Kupriyanov and G. Martin (The Scripps Research Institute Genetics Core) and S. Head and P. Natarajan (The Scripps Research Institute NGS Core) for technical assistance. This project was supported by the US National Institutes of Health (R21AI126011, R21AI101692 and R37AI059714).
Author information
Authors and Affiliations
Contributions
A.L.G. participated in many of the studies, identifying anomalies in PLD4- and PLD3-deficient mice and their myeloid cell cytokine responses and in the preparation of PLD3 and PLD4 proteins; D.H. elucidated the enzymatic function of PLD3 and PLD4, carried out all studies in 293 cells, and contributed to other studies; C.H. generated the Pld4fl/fl and Pld4–/– mutant mice and characterized many features of their phenotypes; A.M. cloned Pld4 cDNA and characterized Pld4 expression in the B cell lineage; V.T. and B.R.L. participated in the analysis of experimental autoimmune encephalomyelitis in Pld4–/– mice; P.D.S., T.R.B. and T.C.T. provided technical support; K.O. provided histological analysis of liver pathology; H.S.C., F.K., P.K., A. Zeitjian, R.L.S., M.B. and S.R.S. were student interns who participated in generating constructs for CRISPR-Cas9 mutagenesis and site-directed mutagenesis of Pld3 and Pld4; A. Zarpellon provided assistance with blood analysis using the IDEX Procyte DX system; B.C. assisted C.H. in the search for potential phospholipase activity of PLD4; E.S.P., M.O.K. and H.H. provided advice and assistance with DC studies (Fig. 1l–n) and data in Fig. 4d; J.T. and J.C.d.l.T. provided advice and assistance with RNA virus studies; M.D. assisted with PLD3 reagents and antibodies; L.T. provided advice on and assistance with the expression of PLD4 protein; A.L.G. and D.N. co-directed these studies; and A.L.G., D.H. and D.N. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Generation and analysis of conditional Pld4 knockout mouse mutation and Pld3 null mutants.
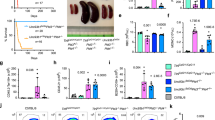
(a) Design of targeting construct in relation to the wild type Pld4 locus, including restriction enzyme sites and location of hybridization probe. FRT recombinase recognition sites flanking the neomycin resistance gene (neo) are indicated with brown triangles and loxP sites are blue triangles. The constructs were designed such that CRE recombinase deletes Pld4 exons 4-6 containing the first HKD domain, and introduces a frame-shift. (b,c) Southern blot analysis of targeted embryonic stem cells, identifying clone 28 as correctly targeted. Experiment was performed twice for each identified cell line. (d) Western blot analysis of mouse PLD4 protein expression. Spleen lysates were analyzed using a rabbit polyclonal anti-PLD4 antiserum. Experiment was performed once. (e) Flow cytometry analysis of PLD4 expression in spleen DCs. DCs (CD11c+TCRb–CD19–) were gated as shown in the upper panels into pDCs (SiglecH+CD45RA+), CD8a DCs (CD11b–CD8a+) and CD8– DCs (CD8–CD11b+) and evaluated for intracellular PLD4 using MAb16 (lower panels) comparing Pld4fl/fl (red) and Pld4–/– (blue). Experiment was performed twice. (f) Numbers of sorted DC subsets isolated from spleens from 10 mice of indicated genotype. Error bars indicated standard deviation between three sorts. (g) Elevated MHCII but not CD86 in peritoneal macrophages of Pld4–/– compared to Pld4+/– mice n= 6/group. Statistics were determined using a two-tailed Student’s T test assuming equal variance. (h) Proportions of splenic B cell subsets from indicated genotypes. Mice were 8-10 weeks old, 6 mice/group. Marginal zone subset statistics were determined using a two-tailed Student’s T test assuming equal variance. (i) Normalization of MZ B cell frequency in the absence of IFN-γ. n=7,9. Statistics were determined using a two-tailed Student’s T test assuming equal variance. (j-n) Generation and analysis of Pld3 mutants. (j) Exon 9 sequence changes in mutant alleles of Pld3 transmitted by different founder mice. Note some lines had two independent mutant alleles. (k) Immunoblot of PLD3 protein expression in lysates from thioglycolate-elicited macrophages harvested from C57BL/6, Pld4–/–, or Pld3 Crispr Founder mice. This experiment was performed once. (l-n) Effect of Pld3 or Pld4 genotype on spleen weight (l), splenic NK cell percentage (m), and MHCII expression on resident peritoneal macrophages (n). Pld4–/– compared to Pld4fl/fl mice n= 4/group, Pld3+/– compared to Pld3–/– n=5/group. Bars show means and standard deviation and each point represents an individual animal. Statistics were determined using a two-tailed Student’s T test assuming equal variance. This experiment was performed twice.
Supplementary Figure 2 Analysis of experimental autoimmune encephalomyelitis induced in the indicated mouse strains.
EAE was induced as described in Methods and the clinical severity score was tracked over the indicated days. Each figure represents an independent set of experiments. (a-c) represent a series of studies carried out by the laboratory of BL in a distinct animal room and time period compared to (d-f), which were carried out by AG and TB using distinct reagents and mouse rooms. In experiments (a-e) mice of the indicated genotypes were challenged with MOG on d0. In (f) lethally irradiated B6.CD45.1 mice were first reconstituted with Pld4–/– or Pld4fl/fl bone marrow 10 weeks prior to EAE induction. In all cases, mean and standard error of the mean are presented. In (e), P value reflects comparison between Ifng–/–Pld4–/– and B6.CD45.1 scores. Statistics were determined using a two-tailed paired Student’s T test assuming equal variance.
Supplementary Figure 3 Splenic sorted CD8+ DCs produce IL-12 in response to 2216PS and VACV70 independent of the STING pathway.
WT and Tmem173–/– sorted CD8+ DCs were plated in duplicates, stimulated with the indicated agents overnight and cytokine release measured. (a) IFN-λ production, measured by ELISA. (b) IL-12p70 production, measured by Luminex assay. (c) Comparison of IL-12p70 production (measured by ELISA) of CD8+ cDC sorted from mice deficient in IFN-γ alone or lacking both IFN-γ and PLD4. Duplicate cultures shown. Experiment performed twice. (d) Sorted splenic CD8–CD11bhi DCs produce IL-12p70 and IL-6 to indicated ligands. DCs plated in triplicate. Experiment performed twice. In all graphs, mean and standard deviation are shown. (c-d) Statistics were determined using a non-paired two-tailed Student’s T test assuming equal variance.
Supplementary Figure 4 LC-MS/MS analysis of bovine spleen phosphodiesterase II.
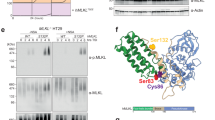
(a) An example of a peptide identification, with the peptide call in the upper left, the predicted fragments shown on the upper right, corresponding to the indicated m/z peaks in the chart. (b) Overview of the peptide matches to bovine PLD3, shown in red. The underlined peptide corresponds to the peptide shown in (a). (c) Complete list of all proteins identified by at least five unique peptides by LC-MS/MS analysis. PLD3 shown in bold. This experiment was performed once.
Supplementary Figure 5 Comparative efficiency of digestion of PLD4 and PLD3 for phosphorothioate (PS) and phosphodiester (PO) linked ODNs.
The indicated concentrations of mouse PLD3 or PLD4 were used to digest the indicated ODNs (2.5 μM) in 50 mM MEF pH 5.0, 125 mM NaCl at 37oC for 2 hours, electrophoresed on 20% polyacrylamide/TBE/urea gels and stained with Sybr gold. CpG-ODNs were provided in excess, 2.5 μM. This experiment was performed four times with similar results.
Supplementary Figure 6 Analysis of responses of Pld4–/– DCs, Pld3–/– thioglycolate-elicited macrophages and PLD3–/– human cell lines to TLR9 agonists.
(a-c) Responses of Pld4–/– DCs and Pld3–/– thioglycolate-elicited macrophages to 2216 fragments when complexed with Lyovec. Triplicate cultures of pDCs, CD8+ DCs and thioglycolate-elicited macrophages of the indicated genotypes were stimulated as in Fig 5 b-e except that the ligands were precomplexed with Lyovec at a concentration of 1 μg/ml. (a) CD8+ DCs; (b) pDCs, (c) macrophages. Mean and standard deviation are shown. This is representative of five experiments (a and b) and three experiments (c). Statistics were determined using an unpaired two-tailed Student’s T test assuming equal variance. (d) TLR9 responses of PLD3-deficient human cells to phosphorothioate (PS) or phosphodiester form of CpG-B (2006) or CpG-A (2216). PLD3 was knocked out in HEK-BluehTLR9 cells using CRISPR/Cas9, then individual, sequence-verified clones were assessed for TLR9 responses to the indicated ODNs. NFκB reporter activation was then measured. ODNs were used at 1 μM. Cells were plated in duplicates. Experiment performed three times with similar results. Statistics were determined using a two-tailed Student’s T test assuming equal variance comparing parental HEK-BluehTLR9 cells with PLD3KO.10. Mean and standard deviation are shown. (e) PLD3-deficient HEK-BluehTLR9 cells as in (d) stimulated with 2216 PS subfragments. Experiment was performed three times with similar results. Statistics were determined using a two-tailed Student’s T test assuming equal variance comparing parental HEK-BluehTLR9 cells with PLD3KO.10. Mean and standard deviation are shown. (f) IFN-α or IL-12p70 responses of Pld3+/– or Pld3–/– FLT3L-cultured DCs to listed TLR9 stimuli. Cells were plated in triplicates. Experiment was performed once. Mean and standard deviation are shown.
Supplementary Figure 7 Analysis of Pld3–/–Pld4–/– animals and irradiated recipients of bone marrow from Pld3–/–Pld4–/– animals.
(a) Weanling expected and observed genotype results from F2 crosses of either Pld3–/–Pld4+/– (left) or Pld3+/–Pld4–/– (right) parental breeding schemes. A Pearson’s Chi Square test was performed. (b) A 16 day-old litter from Pld3–/–Pld4+/– x Pld3+/–Pld4–/– breeding with genotype of pups shown. (c) Macroscopic image of liver from Pld3–/–Pld4–/– animal depicted in (b). (d) MHCII and CD86 expression levels of peritoneal macrophages isolated from bone marrow chimeric mice receiving bone marrow from either Pld3–/–Pld4+/– (red), Pld3+/–Pld4–/– (blue), or Pld3–/–Pld4–/– (green) donors. MHCII and CD86 expression levels of peritoneal macrophages from a control C57BL/6 animal (orange) are shown for comparison. Experiment performed once. Blood monocyte levels (e, i), serum cytokine IFN-γ (g), IFN-α (h), IL-6 (k) and chemokine CXCL10 (f), MCP3 (j) levels were determined from bone marrow chimeric mice receiving Pld3–/–Pld4+/– (dark grey), Pld3+/–Pld4–/– (light grey), or Pld3–/–Pld4–/– (white) bone marrow 8 weeks earlier (Dotted line represents level of sensitivity of assay). (e-k) Mean and individual mouse values are shown. Statistics were determined using a two-tailed Student’s T test assuming equal variance. Experiment was performed once.
Supplementary Figure 8 Gating strategies used to identify cellular subsets.
(a) DC subsets, (b) resident peritoneal macrophages, (c) isotype control staining for MHCII expression from resident peritoneal macrophages from control or Pld4–/– mice as gated in b, (d) bone marrow reconstituted B cells or (e) bone marrow reconstituted T cells.
Supplementary information
Supplementary Figures
Supplementary Figures 1-8 and Supplementary Tables 1-4
Rights and permissions
About this article
Cite this article
Gavin, A.L., Huang, D., Huber, C. et al. PLD3 and PLD4 are single-stranded acid exonucleases that regulate endosomal nucleic-acid sensing. Nat Immunol 19, 942–953 (2018). https://doi.org/10.1038/s41590-018-0179-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0179-y
This article is cited by
-
Cytosolic nucleic acid sensing and mitochondrial transcriptomic changes as early triggers of metabolic disease in db/db mice
Mammalian Genome (2024)
-
Phospholipase D4 as a signature of toll-like receptor 7 or 9 signaling is expressed on blastic T-bet + B cells in systemic lupus erythematosus
Arthritis Research & Therapy (2023)
-
Phospholipase D3 degrades mitochondrial DNA to regulate nucleotide signaling and APP metabolism
Nature Communications (2023)
-
Understanding nucleic acid sensing and its therapeutic applications
Experimental & Molecular Medicine (2023)
-
Regulation of the nucleic acid-sensing Toll-like receptors
Nature Reviews Immunology (2022)