Abstract
Transcriptome profiling is widely used to infer functional states of specific cell types, as well as their responses to stimuli, to define contributions to physiology and pathophysiology. Focusing on microglia, the brain’s macrophages, we report here a side-by-side comparison of classical cell-sorting-based transcriptome sequencing and the ‘RiboTag’ method, which avoids cell retrieval from tissue context and yields translatome sequencing information. Conventional whole-cell microglial transcriptomes were found to be significantly tainted by artifacts introduced by tissue dissociation, cargo contamination and transcripts sequestered from ribosomes. Conversely, our data highlight the added value of RiboTag profiling for assessing the lineage accuracy of Cre recombinase expression in transgenic mice. Collectively, this study indicates method-based biases, reveals observer effects and establishes RiboTag-based translatome profiling as a valuable complement to standard sorting-based profiling strategies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Gosselin, D. et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell 159, 1327–1340 (2014).
Heiman, M. et al. A translational profiling approach for the molecular characterization of CNS cell types. Cell 135, 738–748 (2008).
Sanz, E. et al. Cell-type-specific isolation of ribosome-associated mRNA from complex tissues. Proc. Natl Acad. Sci. USA 106, 13939–13944 (2009).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Jung, S. et al. Analysis of fractalkine receptor CX3CR1 function by targeted deletion and green fluorescent protein reporter gene insertion. Mol. Cell. Biol. 20, 4106–4114 (2000).
Davalos, D. et al. ATP mediates rapid microglial response to local brain injury in vivo. Nat. Neurosci. 8, 752–758 (2005).
Goldmann, T. et al. A new type of microglia gene targeting shows TAK1 to be pivotal in CNS autoimmune inflammation. Nat. Neurosci. 16, 1618–1626 (2013).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Orthgiess, J. et al. Neurons exhibit Lyz2 promoter activity in vivo: Implications for using LysM-Cre mice in myeloid cell research. Eur. J. Immunol. 46, 1529–1532 (2016).
Kim, K.-W. et al. In vivo structure/function and expression analysis of the CX3C chemokine fractalkine. Blood 118, e156–e167 (2011).
Fonseca, M. I. et al. Cell-specific deletion of C1qa identifies microglia as the dominant source of C1q in mouse brain. J. Neuroinflammation 14, 48 (2017).
Goldmann, T. et al. Origin, fate and dynamics of macrophages at central nervous system interfaces. Nat. Immunol. 17, 797–805 (2016).
Zeisel, A. et al. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
van den Brink, S. C. et al. Single-cell sequencing reveals dissociation-induced gene expression in tissue subpopulations. Nat. Methods 14, 935–936 (2017).
A-Gonzalez, N. et al. Phagocytosis imprints heterogeneity in tissue-resident macrophages. J. Exp. Med. 214, 1281–1296 (2017).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 22, 1775–1789 (2012).
Braunschweig, U. et al. Widespread intron retention in mammals functionally tunes transcriptomes. Genome Res. 24, 1774–1786 (2014).
Khong, A. et al. The stress granule transcriptome reveals principles of mRNA accumulation in stress granules. Mol. Cell 68, 808–820.e5 (2017).
Hubstenberger, A. et al. P-body purification reveals the condensation of repressed mRNA regulons. Mol. Cell 68, 144–157.e5 (2017).
Tebaldi, T. et al. Widespread uncoupling between transcriptome and translatome variations after a stimulus in mammalian cells. BMC Genomics 13, 220 (2012).
Bahar Halpern, K. et al. Nuclear retention of mRNA in mammalian tissues. Cell Rep. 13, 2653–2662 (2015).
Hirrlinger, J. et al. Split-cre complementation indicates coincident activity of different genes in vivo. PLoS One 4, e4286 (2009).
Wolf, Y. et al. Brown-adipose-tissue macrophages control tissue innervation and homeostatic energy expenditure. Nat. Immunol. 18, 665–674 (2017).
Boutej, H. et al. Diverging mRNA and protein networks in activated microglia reveal SRSF3 suppresses translation of highly upregulated innate immune transcripts. Cell Rep. 21, 3220–3233 (2017).
Wu, Y. E., Pan, L., Zuo, Y., Li, X. & Hong, W. Detecting activated cell populations using single-cell RNA-seq. Neuron 96, 313–329.e6 (2017).
Tilgner, H. et al. Deep sequencing of subcellular RNA fractions shows splicing to be predominantly co-transcriptional in the human genome but inefficient for lncRNAs. Genome Res. 22, 1616–1625 (2012).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001).
Jaitin, D. A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Acknowledgements
We thank all members of the Jung laboratory for discussion, the staff of the Weizmann Animal Facility, members of the FACS facility for expert advice, G. Friedlander for help with bioinformatics, and C. Glass for sharing sequencing data. S.J. was supported by the Israeli Science Foundation (887/11), an Infect-ERA grant, the European Research Council (Adv ERC 340345), and the Deutsche Forschungsgemeinschaft (CRC/TRR167 ‘NeuroMac’). I.B and M.G. were supported by the Deutsche Forschungsgemeinschaft DFG-SFB 1052/1: ‘Obesity mechanisms’ (projects A09 and B09).
Author information
Authors and Affiliations
Contributions
Z.H. and S.J. conceived the project and designed the experiments; Z.H., A.V. and J.O. performed the experiments; L.C.-M. performed RNA-seq and E.D. analyzed the data. I.U. and B.Z. performed bioinformatics analysis. S.Y., A.S., D.V., S.B.-H., I.B. and M.G. advised on experiments; Z.H. and S.J. wrote the paper; S.J. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Enrichment and de-enrichment of cell-specific genes in TAM-treated Cx3cr1CreER:Rpl22HA brains after IP.
Graphs showing normalized reads of example genes from heat map in Fig. 1c, showing cell-specific genes of Microglia (A), Astrocytes (B), Neurons (C) and Oligodendrocytes (D) enrichement in input, IP IgG control and IP HA samples of TAM-treated Cx3cr1CreER:Rpl22HA brains after IP. Each dot represents an individual mouse, n = 3, lines represent mean.
Supplementary Figure 2 Immunohistochemistry analysis of brains of reporter animals.
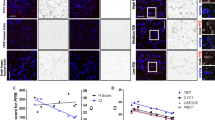
(A) Microscopic analysis of hypothalamus sections from Cx3cr1Cre:R26-YFP mice (left panel) and Cx3cr1CreER:R26-YFP mice (TAM treated (right panel) or not (middle panel), stained for IBA-1, GFP and DAPI, showing neuronal expression of GFP in Cx3cr1Cre brains and microglia-restricted GFP expression in Cx3cr1CreER brains. The animals analyzed are F1 offspring of the intercross of homozygote Cx3cr1Cre or Cx3cr1CreER animals and homozygote R26-YFP mice. Representative of 2 independent experiments. (B) Immuno-fluorescent staining of 30µm thick brain sections of cortex (upper panels) and cerebellum (lower panels) of Cx3cr1Cre:Rpl22HA (left) and Cx3cr1CreER:Rpl22HA animals, either untreated (middle) or TAM-treated (right), stained for IBA1, HA and NeuN, showing neuronal expression of HA in Cx3cr1Cre brains, and microglia-restricted HA expression in TAM-treated Cx3cr1CreER brains. Scale bar cortex = 100µm, scale bar cerebellum = 200µm. Representative of 2 independent experiments.
Supplementary Figure 3 Gating strategy for microglial isolation.
FACS dot plots representing gating strategy for sorting of microglia. Cells were gated for single cells by FSC-W and FSC-A; monocytes, neutrophils and dead cells were excluded by gating on Ly6C/G (Gr1)– DAPI– cells; final microglia gate was on CD11b+ CD45int cells.
Supplementary Figure 4 Microglial gene expression profiles are comparable across different retrieval methods.
Graphs showing normalized reads of example genes from cluster I of Fig. 2b, showing microglial genes at comparable reads number, indicating that retrieval methods are comparable. n = 3, lines represent mean.
Supplementary Figure 5 Microglia activation signature is robust, reproducible and induced by cell extraction from tissue.
(A) Heatmap clustered by K-means clustering, comparing direct IP to whole mRNA retrieved from sorted cells extracted with (+coll) or without (-coll) enzymatic digestion in the isolation protocol, revealing 4 clusters similar to Fig. 2b. n = 4 individual mice (one repeat of IP-HA was removed due to technical reasons). (B) Correlation plots of average of log2 normalized reads (n = 4) plotting sorted cells with enzymatic digestion on the Y axis and sorted cells without enzymatic digestion on the X axis (left panel) or IP on the X axis (right panel). (C) Correlation matrix combining two independent experiments, showing high reproducibility of the results.
Supplementary Figure 6 Analysis of microglia isolated from mice challenged with LPS.
(A) Heatmap of RNA-seq data comparing RiboTag to sorting with LPS treatment, taking significantly changed genes (fold change>2, p-value<0.05, as calculated by DESeq2 statistical analysis). n = 3 for PBS group, n = 4 for LPS group. (B) Graphs showing normalized reads of example genes taken from cluster I from Fig. S6A, showing similar levels of microglia signature genes among samples. Each dot represents an individual mouse, n = 3 for PBS group, n = 4 for LPS group, lines represent mean.
Supplementary Figure 7 Expression profiles of selected genes across brain cell populations.
Non-microglia mRNAs suggesting cargo contamination (Fig. 5f)Data obtained from (http://web.stanford.edu/group/barres_lab/brain_rnaseq.html) 9. (Zhang, Y. et al. J. Neurosci. 34, 11929–11947 (2014)).
Supplementary information
Supplementary Figures and Supplementary Table 1
Supplementary Figures 1–7 and Supplementary Table 1
Rights and permissions
About this article
Cite this article
Haimon, Z., Volaski, A., Orthgiess, J. et al. Re-evaluating microglia expression profiles using RiboTag and cell isolation strategies. Nat Immunol 19, 636–644 (2018). https://doi.org/10.1038/s41590-018-0110-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0110-6
This article is cited by
-
Cell cycle specific, differentially tagged ribosomal proteins to measure phase specific transcriptomes from asynchronously cycling cells
Scientific Reports (2024)
-
Microglia regulation of central nervous system myelin health and regeneration
Nature Reviews Immunology (2024)
-
Transcriptomics and translatomics identify a robust inflammatory gene signature in brain endothelial cells after ischemic stroke
Journal of Neuroinflammation (2023)
-
Cerebellar granule neurons induce Cyclin D1 before the onset of motor symptoms in Huntington’s disease mice
Acta Neuropathologica Communications (2023)
-
Translational profiling identifies sex-specific metabolic and epigenetic reprogramming of cortical microglia/macrophages in APPPS1-21 mice with an antibiotic-perturbed-microbiome
Molecular Neurodegeneration (2023)