Abstract
Pathogens have co-evolved with mosquitoes to optimize transmission to hosts. Mosquito salivary-gland extract is known to modulate host immune responses and facilitate pathogen transmission, but the underlying molecular mechanisms of this have remained unknown. In this study, we identified and characterized a prominent 15-kilodalton protein, LTRIN, obtained from the salivary glands of the mosquito Aedes aegypti. LTRIN expression was upregulated in blood-fed mosquitoes, and LTRIN facilitated the transmission of Zika virus (ZIKV) and exacerbated its pathogenicity by interfering with signaling through the lymphotoxin-β receptor (LTβR). Mechanically, LTRIN bound to LTβR and ‘preferentially’ inhibited signaling via the transcription factor NF-κB and the production of inflammatory cytokines by interfering with the dimerization of LTβR during infection with ZIKV. Furthermore, treatment with antibody to LTRIN inhibited mosquito-mediated infection with ZIKV, and abolishing LTβR potentiated the infectivity of ZIKV both in vitro and in vivo. This study provides deeper insight into the transmission of mosquito-borne diseases in nature and supports the therapeutic potential of inhibiting the action of LTRIN to disrupt ZIKV transmission.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Caraballo, H. & King, K. Emergency department management of mosquito-borne illness: malaria, dengue, and West Nile virus. Emerg. Med. Pract. 16, 1–23 (2014). quiz 23–24.
Coutinho-Abreu, I. V., Guimarães-Costa, A. B. & Valenzuela, J. G. Impact of insect salivary proteins in blood feeding, host immunity, disease, and in the development of biomarkers for vector exposure. Curr. Opin. Insect Sci. 10, 98–103 (2015).
Schneider, B. S. & Higgs, S. The enhancement of arbovirus transmission and disease by mosquito saliva is associated with modulation of the host immune response. Trans. R. Soc. Trop. Med. Hyg. 102, 400–408 (2008).
Chagas, A. C. et al. Collagen-binding protein, Aegyptin, regulates probing time and blood feeding success in the dengue vector mosquito, Aedes aegypti. Proc. Natl. Acad. Sci. USA 111, 6946–6951 (2014).
Calvo, E., Mans, B. J., Andersen, J. F. & Ribeiro, J. M. C. Function and evolution of a mosquito salivary protein family. J. Biol. Chem. 281, 1935–1942 (2006).
Xu, X., Chang, B. W., Mans, B. J., Ribeiro, J. M. C. & Andersen, J. F. Structure and ligand-binding properties of the biogenic amine-binding protein from the saliva of a blood-feeding insect vector of Trypanosoma cruzi. Acta Crystallogr. D Biol. Crystallogr. 69, 105–113 (2013).
Xu, X. et al. Structure and function of a “yellow” protein from saliva of the sand fly Lutzomyia longipalpis that confers protective immunity against Leishmania major infection. J. Biol. Chem. 286, 32383–32393 (2011).
Ma, D. et al. Triplatin, a platelet aggregation inhibitor from the salivary gland of the triatomine vector of Chagas disease, binds to TXA(2) but does not interact with glycoprotein PVI. Thromb. Haemost. 107, 111–123 (2012).
Assumpcao, T. C. F. et al. Salivary antigen-5/CAP family members are Cu2+-dependent antioxidant enzymes that scavenge O2− and inhibit collagen-induced platelet aggregation and neutrophil oxidative burst. J. Biol. Chem. 288, 14341–14361 (2013).
Collin, N. et al. Lufaxin, a novel factor Xa inhibitor from the salivary gland of the sand fly Lutzomyia longipalpis, blocks PAR2 activation and inhibits inflammation and thrombosis in vivo. Arterioscler. Thromb. Vasc. Biol. 32, 2185–2198 (2012).
Surasombatpattana, P. et al. Aedes aegypti saliva contains a prominent 34-kDa protein that strongly enhances dengue virus replication in human keratinocytes. J. Invest. Dermatol. 134, 281–284 (2014).
Conway, M. J. et al. Aedes aegypti D7 saliva protein inhibits dengue virus infection. PLoS Negl. Trop. Dis. 10, e0004941 (2016).
Upadhyay, V. & Fu, Y.-X. Lymphotoxin signalling in immune homeostasis and the control of microorganisms. Nat. Rev. Immunol. 13, 270–279 (2013).
Ware, C. F. Network communications: lymphotoxins, LIGHT, and TNF. Annu. Rev. Immunol. 23, 787–819 (2005).
Onder, L. et al. Endothelial cell-specific lymphotoxin-β receptor signaling is critical for lymph node and high endothelial venule formation. J. Exp. Med. 210, 465–473 (2013).
Spahn, T. W., Eugster, H.-P., Fontana, A., Domschke, W. & Kucharzik, T. Role of lymphotoxin in experimental models of infectious diseases: potential benefits and risks of a therapeutic inhibition of the lymphotoxin-β receptor pathway. Infect. Immun. 73, 7077–7088 (2005).
Macho-Fernandez, E. et al. Lymphotoxin-β receptor signaling limits mucosal damage through driving IL-23 production by epithelial cells. Mucosal Immunol. 8, 403–413 (2015).
Wang, Y. et al. Lymphotoxin-β receptor signaling in intestinal epithelial cells orchestrates innate immune responses against mucosal bacterial infection. Immunity 32, 403–413 (2010).
Jones, A. V. et al. GWAS of self-reported mosquito bite size, itch intensity and attractiveness to mosquitoes implicates immune-related predisposition loci. Hum. Mol. Genet. 26, 1391–1406 (2017).
Wolf, M. J., Seleznik, G. M., Zeller, N. & Heikenwalder, M. The unexpected role of lymphotoxin-β receptor signaling in carcinogenesis: from lymphoid tissue formation to liver and prostate cancer development. Oncogene 29, 5006–5018 (2010).
Sudhamsu, J. et al. Dimerization of LTβR by LTα1β2 is necessary and sufficient for signal transduction. Proc. Natl. Acad. Sci. USA 110, 19896–19901 (2013).
Hoesel, B. & Schmid, J. A. The complexity of NF-κB signaling in inflammation and cancer. Mol. Cancer 12, 86 (2013).
Vorou, R. Zika virus, vectors, reservoirs, amplifying hosts, and their potential to spread worldwide: what we know and what we should investigate urgently. Int. J. Infect. Dis. 48, 85–90 (2016).
Ma, W., Li, S., Ma, S., Jia, L., Zhang, F. & Zhang, Y. et al. Zika virus causes testis damage and leads to male infertility in mice. Cell 167, 1511–1524 (2016).
Govero, J. et al. Zika virus infection damages the testes in mice. Nature 540, 438–442 (2016).
Nowakowski, T. J. et al. expression analysis highlights AXL as a candidate Zika virus entry receptor in neural stem cells. Cell Stem Cell 18, 591–596 (2016).
Verhelst, J., Hulpiau, P. & Saelens, X. Mx proteins: antiviral gatekeepers that restrain the uninvited. Microbiol. Mol. Biol. Rev. 77, 551–566 (2013).
Lazear, H. M. et al. A mouse model of Zika virus pathogenesis. Cell Host Microbe 19, 720–730 (2016).
Kruger, P. et al. Neutrophils: Between host defence, immune modulation, and tissue injury. PLoS Pathog. 11, e1004651 (2015).
Arcà, B., Lombardo, F., Struchiner, C. J. & Ribeiro, J. M. C. Anopheline salivary protein genes and gene families: an evolutionary overview after the whole genome sequence of sixteen Anopheles species. BMC Genomics 18, 153 (2017).
Pingen, M. et al. Host inflammatory response to mosquito bites enhances the severity of arbovirus infection. Immunity 44, 1455–1469 (2016).
Anisuzzaman, A., Islam, M. K., Alim, M. A. & Tsuji, N. Longistatin, an EF-hand Ca2+-binding protein from vector tick: identification, purification, and characterization. in Calcium-Binding Proteins and RAGE: From Structural Basics to Clinical Applications (ed. Heizmann, C. W.) 127–146 (Humana Press, Totowa, NJ, 2013).
Anisuzzaman, H. T. et al. Longistatin in tick saliva blocks advanced glycation end-product receptor activation. J. Clin. Invest. 124, 4429–4444 (2014).
Tripathi, S. et al. A novel Zika virus mouse model reveals strain specific differences in virus pathogenesis and host inflammatory immune responses. PLoS Pathog. 13, e1006258 (2017).
Tseng, P.-H. et al. Different modes of ubiquitination of the adaptor TRAF3 selectively activate the expression of type I interferons and proinflammatory cytokines. Nat. Immunol. 11, 70–75 (2010).
Yang, S. et al. Chemical punch packed in venoms makes centipedes excellent predators. Mol. Cell. Proteomics 11, 640–650 (2012).
Zhang, Z. et al. Mitochondrial DNA-LL-37 complex promotes atherosclerosis by escaping from autophagic recognition. Immunity 43, 1137–1147 (2015).
Ge, J., Gong, Y.-N., Xu, Y. & Shao, F. Preventing bacterial DNA release and absent in melanoma 2 inflammasome activation by a Legionella effector functioning in membrane trafficking. Proc. Natl. Acad. Sci. USA 109, 6193–6198 (2012).
Li, C. et al. Zika virus disrupts neural progenitor development and leads to microcephaly in mice. Cell Stem Cell 19, 120–126 (2016).
Liu, Y. et al. Evolutionary enhancement of Zika virus infectivity in Aedes aegypti mosquitoes. Nature 545, 482–486 (2017).
Juhn, J. et al. Spatial mapping of gene expression in the salivary glands of the dengue vector mosquito, Aedes aegypti. Parasit. Vectors 4, 1 (2011).
Qi, X. et al. Antagonistic regulation by the transcription factors C/EBPα and MITF specifies basophil and mast cell fates. Immunity 39, 97–110 (2013).
Acknowledgements
We thank F. Shao (National Institute of Biological Sciences, Beijing, China) for Ifnar–/– mice, and Y.-X.Fu (UT Southwestern Medical Center) for LTβR-immunoglobulin. Supported by National Key Research and Development Program of China (2017YFD0500300 and 2016YFC1201000), National Natural Science Foundation of China (21761142002, 31600721, 31200590, 31630075 and 31701134), Chinese Academy of Sciences (XDB13000000, KSZD-EW-Z-007, CXJJ-17-M141, Y4ZK111B01 and Y602381081) and Yunnan Province (2012BC009).
Author information
Authors and Affiliations
Contributions
L.J., C.S., X.H., O.R., H.Z., M.Y. and R.L. performed LTRIN identification, LTRIN preparation and data analysis; L.J., X.G., P.S., P.L., T.X., C.H., C.-F.Q., J.G., H.P., M.Z., G.C. and X.Q. performed ZIKV infection and data analysis; X.Q. wrote the manuscript with input from R.L. and L.J.; and X.Q. and R.L. designed the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 LTRIN purification from SGE of mosquitoes
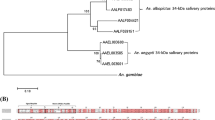
a, Representative salivary glands of mosquitoes. Salivary glands (SGs) from unfed (control) and fed A. aegypti were dissected for protein and RNA extraction. b, Salivary gland extract was separated by Sephadex G-75 gel filtration. Fraction I had immunosuppressive activities, which was selected for further purification. c, Fraction I was further separated by the Resource Q anion-exchange column. Peak IV (arrow indicated) was collected for SDS-PAGE analysis, and amino acid sequencing was performed by Edman degradation. d, Purified proteins from peak IV in (c) were separated on SDS-PAGE and stained with Coomassie Blue. The molecular weight of LTRIN was approximately 15-kDa. e, Protein sequence of A. aegypti LTRIN. Sequence of the peptide recovered from SGE of A. aegypti is given in red, and sequence of the recombinant protein purified from E. coli is underlined. f, Binding affinity analysis of LTRIN and LTβR. g, Binding affinity analysis of LTαβ2 and LTβR. Data are representative of three independent experiments. Scale bars, 300 μm for (a).
Supplementary Figure 2 LTRIN targets mouse LTβR
a, Homology analysis of human and mouse LTβRs. The extracellular domain is given in red. b, Dimerization analysis of mouse LTβR. Mouse BMDMs stimulated with mouse LTα and LTRIN for 1 h and cross-linked for 15 min with disuccinimidyl suberate, followed by lysing and western blot analysis. The ratio of mouse LTβR dimer to monomer was quantified (right). Data are representative of (left) or form (right) two independent experiments (b).
Supplementary Figure 3 Intracellular ZIKV staining in MSF, BMDM and HUVEC cells
a, Expression analysis of ZIKV in HUVEC and THP-1 cells during ZIKV infection (MOI 0.5) and ZIKV combined with LTRIN administration (ZV/IN, ZIKV at MOI 0.5 and LTRIN at 100 ng/mL) for 30 min. b, Confocal immunofluorescence analysis of ZIKV in MSFs and BMDMs during ZIKV infection (MOI 0.5) and ZIKV combined with LTRIN administration (ZIKV at MOI 0.5 and LTRIN at 100 ng/mL) for 12 hours. c, Confocal immunofluorescence analysis of ZIKV and AXL in LTRIN treated HUVEC cells and HUVEC cells with ZIKV infection (MOI 0.5) and ZIKV combined with LTRIN administration (ZIKV at MOI 0.5 and LTRIN at 100 ng/mL) for 12 hours. Scale bars, 20 μm for (b, c). *, P < 0.05; **, P < 0.01; ns, not significant (Student’s two-sided t-test without multiple-comparisons correction). Each symbol indicates an individual reaction in one experiment (a). Data are representative of (a, left in b and c) or from (right in b and c) three independent experiments.
Supplementary Figure 4 Effect of LTαβ on ZIKV infection
Expression analysis of ZIKV in THP-1 and HUVEC cells during ZIKV infection (MOI 0.5) and ZIKV combined with LTαβ2 administration (ZV/LTαβ2, ZIKV at MOI 0.5 and LTαβ2 at 1 nM) for indicated times. ns, not significant (Student’s two-sided t-test without multiple-comparisons correction). Each symbol indicates an individual reaction (technical replicate) in one experiment. Data are representative of two independent experiments.
Supplementary Figure 5 Effect of LTRIN on type I IFN response induced by ZIKV infection.
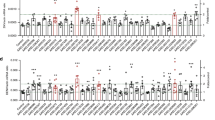
a,b, Expression analysis of ZIKV, Mx1, Mx2, Ifnb, Ifne, Cxcl9, Irf7, Isg15, Ifit1, Oasl2, and Usp18 in mouse BMDMs (a) and mouse MSFs (b) during ZIKV infection (MOI 0.5) and ZIKV combined with LTRIN administration (ZV/IN, ZIKV at MOI 0.5 and LTRIN at 100 ng/mL) for indicated times. c,d, Expression analysis of ZIKV, IFNB, and CXCL10 in THP-1 (c) and HUVEC cells (d) during ZIKV infection (MOI 0.5) and ZIKV combined with LTRIN administration (ZV/IN, ZIKV at MOI 0.5 and LTRIN at 100 ng/mL) for indicated times. *, P < 0.05; **, P < 0.01; ***, P < 0.001; ns, not significant (Student’s two-sided t-test without multiple-comparisons correction). Each symbol indicates an individual reaction (technical replicate) in one experiment. Data are representative of three independent experiments (a, b, c, d).
Supplementary Figure 6 LTRIN promoted ZIKV infection in testis and inflammatory response in Ifnar–/– mice in the late time of ZIKV infection.
a, Expression analysis of Il6, Tnf, Il1a, and Cxcl1 in the liver and kidney from WT and Ifnar–/– mice infected with ZIKV (500 PFUs) or ZIKV (500 PFUs) combined with LTRIN (2 μg per mouse) administration (ZV/IN) for 7 days. b, IL-6 and TNF production were measured in the sera and liver from Ifnar–/– mice infected with ZIKV (500 PFUs) or ZIKV (500 PFUs) combined with LTRIN (2 μg per mouse) administration (ZV/IN) for 7 days. c, Six- to seven-week-old male Ifnar–/– (n=3) mice were injected subcutaneously with 500 PFUs of ZIKV or ZIKV combined with LTRIN (2 μg per mouse) and weighed daily over time. d, Viral loads of ZIKV in the testis from mice in (c) at day 7 after infection were detected by qRT-PCR and plaque assay. *, P < 0.05; **, P < 0.01; ***, P < 0.001; ns, not significant (Student’s two-sided t-test without multiple-comparisons correction). Each symbol indicates an individual mouse in one experiment (a, b, d). Data are from two independent experiments (a, n=4; b, n=2), are representative of two independent experiments (c, d, n=3).
Supplementary Figure 7 Effect of LTRIN on dendritic cells, B cells and T cells recruitment trigged by ZIKV infection.
Four- to five-week-old WT mice were subcutaneously injected with 500 PFUs of ZIKV or 500 PFUs of ZIKV combined with LTRIN (2 μg per mouse) (ZV/IN) and immune cell infiltration was analyzed at day 1 post infection. a-d, Flow cytometry of dendritic cells (a), B cells (b), CD4+ T cells (c) and CD8+ T cells (d) in the spleen of mice infected with ZIKV or ZIKV combined with LTRIN (ZV/IN) for 1 day. The percentage of total cells was quantified (right). ns, not significant (Student’s two-sided t-test without multiple-comparisons correction). Each symbol indicates an individual mouse in one experiment (a, b, c, d). Data are representative of two independent experiments (n=4).
Supplementary Figure 8 Model of mosquito salivary protein LTRIN exhibited augmentation for ZIKV infectivity.
ZIKV was transmitted to mammalian host in the context of saliva during mosquito bites. After injection, one of the salivary proteins LTRIN directly and specifically binds to the LTβR and inhibits its activation leading to attenuated early inflammatory immune response, and enhances ZIKV transmission and dissemination in the mammalian host.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1-8
Supplementary Table 1
Human protein microarray analysis to identify proteins that interact with LTRIN
Supplementary Table 2
Top list of protein microarray analysis of LTRIN-binding proteins. R, confirmed times
Supplementary Table 3
Real Time qPCR Primer Sequences
Supplementary Data
Full scans of all blots
Rights and permissions
About this article
Cite this article
Jin, L., Guo, X., Shen, C. et al. Salivary factor LTRIN from Aedes aegypti facilitates the transmission of Zika virus by interfering with the lymphotoxin-β receptor. Nat Immunol 19, 342–353 (2018). https://doi.org/10.1038/s41590-018-0063-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0063-9
This article is cited by
-
Sensing the danger in mosquito spit
The EMBO Journal (2024)
-
A mosquito salivary protein-driven influx of myeloid cells facilitates flavivirus transmission
The EMBO Journal (2024)
-
The tick saliva peptide HIDfsin2 promotes the tick-borne virus SFTSV replication in vitro by enhancing p38 signal pathway
Archives of Toxicology (2023)
-
Tick saliva and its role in pathogen transmission
Wiener klinische Wochenschrift (2023)
-
A leafhopper saliva protein mediates horizontal transmission of viral pathogens from insect vectors into rice phloem
Communications Biology (2022)