Abstract
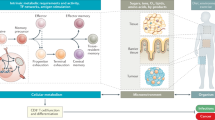
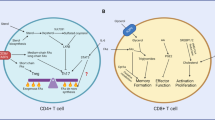
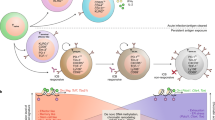
It has become increasingly clear that changes in metabolism are not just consequences of T cell activation but instead are also essential drivers of that process that shape the extent and nature of differentiation and function. The process of T cell exhaustion has been linked to the outcome of chronic immune responses in multiple contexts, including chronic infection, cancer and autoimmunity. Factors that regulate the development and maintenance of exhaustion are of increasing interest as targets of therapeutic modulation. Studies have shown T cell immunometabolism to be integral to the control and development of T cell exhaustion. Early metabolic changes are responsible for the later emergence of exhaustion, do not simply reflect changes secondary to chronic activation and are modifiable. Increased understanding of this metabolic control promises to improve the ability to modulate T cell immunity to chronic antigen stimulation in multiple contexts.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Marina Corral Spence/Springer Nature.

Marina Corral Spence/Springer Nature.

Marina Corral Spence/Springer Nature.

Marina Corral Spence/Springer Nature.
Similar content being viewed by others
References
MacIver, N. J., Michalek, R. D. & Rathmell, J. C. Metabolic regulation of T lymphocytes. Annu. Rev. Immunol. 31, 259–283 (2013).
Chang, C. H. et al. Posttranscriptional control of T cell effector function by aerobic glycolysis. Cell 153, 1239–1251 (2013).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Pearce, E. L. & Pearce, E. J. Metabolic pathways in immune cell activation and quiescence. Immunity 38, 633–643 (2013).
Anastasiou, D. et al. Inhibition of pyruvate kinase M2 by reactive oxygen species contributes to cellular antioxidant responses. Science 334, 1278–1283 (2011).
Zhao, S. et al. ATP-citrate lyase controls a glucose-to-acetate metabolic switch. Cell. Rep.. 17, 1037–1052 (2016).
van der Windt, G. J. et al. Mitochondrial respiratory capacity is a critical regulator of CD8+ T cell memory development. Immunity 36, 68–78 (2012).
Pearce, E. L. et al. Enhancing CD8 T-cell memory by modulating fatty acid metabolism. Nature 460, 103–107 (2009).
O’Sullivan, D. et al. Memory CD8+ T cells use cell-intrinsic lipolysis to support the metabolic programming necessary for development. Immunity 41, 75–88 (2014).
Buck, M. D. et al. Mitochondrial dynamics controls T cell fate through metabolic programming. Cell 166, 63–76 (2016).
van der Windt, G. J. et al. CD8 memory T cells have a bioenergetic advantage that underlies their rapid recall ability. Proc. Natl Acad. Sci. USA 110, 14336–14341 (2013).
Nomura, M. et al. Fatty acid oxidation in macrophage polarization. Nat. Immunol. 17, 216–217 (2016).
Buck, M. D., O’Sullivan, D. & Pearce, E. L. T cell metabolism drives immunity. J. Exp. Med. 212, 1345–1360 (2015).
Araki, K. et al. mTOR regulates memory CD8 T-cell differentiation. Nature 460, 108–112 (2009).
Wherry, E. J. T cell exhaustion. Nat. Immunol. 12, 492–499 (2011).
Pauken, K. E. & Wherry, E. J. Overcoming T cell exhaustion in infection and cancer. Trends Immunol. 36, 265–276 (2015).
McKinney, E. F. & Smith, K. G. T-cell exhaustion: understanding the interface of chronic viral and autoinflammatory diseases. Immunol. Cell. Biol. 94, 935–942 (2016).
Bengsch, B. et al. Bioenergetic insufficiencies due to metabolic alterations regulated by the inhibitory receptor PD-1 are an early driver of CD8+ T cell exhaustion. Immunity 45, 358–373 (2016).
Scharping, N. E. et al. The Tumor microenvironment represses t cell mitochondrial biogenesis to drive intratumoral T cell metabolic insufficiency and dysfunction. Immunity 45, 701–703 (2016).
Chang, C. H. et al. Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell 162, 1229–1241 (2015).
Wherry, E. J. et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity 27, 670–684 (2007).
Baitsch, L. et al. Exhaustion of tumor-specific CD8+ T cells in metastases from melanoma patients. J. Clin. Invest. 121, 2350–2360 (2011).
Gubin, M. M. et al. Checkpoint blockade cancer immunotherapy targets tumour-specific mutant antigens. Nature 515, 577–581 (2014).
Harris, A. L. Hypoxia—a key regulatory factor in tumour growth. Nat. Rev. Cancer 2, 38–47 (2002).
Doedens, A. L. et al. Hypoxia-inducible factors enhance the effector responses of CD8+ T cells to persistent antigen. Nat. Immunol. 14, 1173–1182 (2013).
Tyrakis, P. A. et al. S-2-hydroxyglutarate regulates CD8+ T-lymphocyte fate. Nature 540, 236–241 (2016).
Zhang, Y. et al. Enhancing CD8+ T cell fatty acid catabolism within a metabolically challenging tumor microenvironment increases the efficacy of melanoma immunotherapy. Cancer Cell. 32, 377–391 (2017).
Sukumar, M. et al. Inhibiting glycolytic metabolism enhances CD8+ T cell memory and antitumor function. J. Clin. Invest. 123, 4479–4488 (2013).
Phan, A. T. et al. Constitutive glycolytic metabolism supports CD8+ T cell effector memory differentiation during viral infection. Immunity 45, 1024–1037 (2016).
Paley, M. A. et al. Progenitor and terminal subsets of CD8+ T cells cooperate to contain chronic viral infection. Science 338, 1220–1225 (2012).
Ho, P. C. et al. Phosphoenolpyruvate is a metabolic checkpoint of anti-tumor T cell responses. Cell 162, 1217–1228 (2015).
Schwartz, R. H. T cell anergy. Annu. Rev. Immunol. 21, 305–334 (2003).
Geiger, R. et al. L-Arginine modulates T cell metabolism and enhances survival and anti-tumor activity. Cell 167, 829–842 (2016).
Sinclair, L. V. et al. Control of amino-acid transport by antigen receptors coordinates the metabolic reprogramming essential for T cell differentiation. Nat. Immunol. 14, 500–508 (2013).
Nakaya, M. et al. Inflammatory T cell responses rely on amino acid transporter ASCT2 facilitation of glutamine uptake and mTORC1 kinase activation. Immunity 40, 692–705 (2014).
Wang, R. et al. The transcription factor Myc controls metabolic reprogramming upon T lymphocyte activation. Immunity 35, 871–882 (2011).
Ghoneim, H. E. et al. De novo epigenetic programs inhibit PD-1 blockade-mediated T cell rejuvenation. Cell 170, 142–157 (2017).
Siska, P. J. et al. Mitochondrial dysregulation and glycolytic insufficiency functionally impair CD8 T cells infiltrating human renal cell carcinoma. JCI Insight 2, 93411 (2017).
Schurich, A. et al. Distinct metabolic requirements of exhausted and functional virus-specific CD8 T cells in the same host. Cell Rep. 16, 1243–1252 (2016).
Fisicaro, P. et al. Targeting mitochondrial dysfunction can restore antiviral activity of exhausted HBV-specific CD8 T cells in chronic hepatitis B. Nat. Med.. 23, 327–336 (2017).
Wherry, E. J. & Kurachi, M. Molecular and cellular insights into T cell exhaustion. Nat. Rev. Immunol. 15, 486–499 (2015).
Blackburn, S. D., Shin, H., Freeman, G. J. & Wherry, E. J. Selective expansion of a subset of exhausted CD8 T cells by αPD-L1 blockade. Proc. Natl Acad. Sci. USA 105, 15016–15021 (2008).
Oldstone, M. B. Anatomy of viral persistence. PLoS Pathog. 5, e1000523 (2009).
Chi, H. Regulation and function of mTOR signalling in T cell fate decisions. Nat. Rev. Immunol. 12, 325–338 (2012).
Rao, R. R., Li, Q., Odunsi, K. & Shrikant, P. A. The mTOR kinase determines effector versus memory CD8+ T cell fate by regulating the expression of transcription factors T-bet and Eomesodermin. Immunity 32, 67–78 (2010).
Rao, R. R., Li, Q., Gubbels Bupp, M. R. & Shrikant, P. A. Transcription factor Foxo1 represses T-bet-mediated effector functions and promotes memory CD8+ T cell differentiation. Immunity 36, 374–387 (2012).
Mueller, S. N. & Ahmed, R. High antigen levels are the cause of T cell exhaustion during chronic viral infection. Proc. Natl Acad. Sci. USA 106, 8623–8628 (2009).
Staron, M. M. et al. The transcription factor FoxO1 sustains expression of the inhibitory receptor PD-1 and survival of antiviral CD8+ T cells during chronic infection. Immunity 41, 802–814 (2014).
Blackburn, S. D. et al. Coregulation of CD8+ T cell exhaustion by multiple inhibitory receptors during chronic viral infection. Nat. Immunol. 10, 29–37 (2009).
Oestreich, K. J., Yoon, H., Ahmed, R. & Boss, J. M. NFATc1 regulates PD-1 expression upon T cell activation. J. Immunol. 181, 4832–4839 (2008).
Martinez, G. J. et al. The transcription factor NFAT promotes exhaustion of activated CD8+ T cells. Immunity 42, 265–278 (2015).
Patsoukis, N. et al. PD-1 alters T-cell metabolic reprogramming by inhibiting glycolysis and promoting lipolysis and fatty acid oxidation. Nat. Commun. 6, 6692 (2015).
Odorizzi, P. M., Pauken, K. E., Paley, M. A., Sharpe, A. & Wherry, E. J. Genetic absence of PD-1 promotes accumulation of terminally differentiated exhausted CD8+ T cells. J. Exp. Med. 212, 1125–1137 (2015).
Barber, D. L. et al. Restoring function in exhausted CD8 T cells during chronic viral infection. Nature 439, 682–687 (2006).
McKinney, E. F. & Smith, K. G. T cell exhaustion and immune-mediated disease-the potential for therapeutic exhaustion. Curr. Opin. Immunol. 43, 74–80 (2016).
Frauwirth, K. A. et al. The CD28 signaling pathway regulates glucose metabolism. Immunity 16, 769–777 (2002).
Klein Geltink, R. I. et al. Mitochondrial priming by CD28. Cell 171, 385–397 (2017).
Wieman, H. L., Wofford, J. A. & Rathmell, J. C. Cytokine stimulation promotes glucose uptake via phosphatidylinositol-3 kinase/Akt regulation of Glut1 activity and trafficking. Mol. Biol. Cell. 18, 1437–1446 (2007).
Zheng, Y., Delgoffe, G. M., Meyer, C. F., Chan, W. & Powell, J. D. Anergic T cells are metabolically anergic. J. Immunol. 183, 6095–6101 (2009).
Aubert, R. D. et al. Antigen-specific CD4 T-cell help rescues exhausted CD8 T cells during chronic viral infection. Proc. Natl Acad. Sci. USA 108, 21182–21187 (2011).
McKinney, E. F., Lee, J. C., Jayne, D. R. W., Lyons, P. A. & Smith, K. G. C. T-cell exhaustion, co-stimulation and clinical outcome in autoimmunity and infection. Nature 523, 612–616 (2015).
Pollizzi, K. N. et al. Asymmetric inheritance of mTORC1 kinase activity during division dictates CD8+ T cell differentiation. Nat. Immunol. 17, 704–711 (2016).
Pollizzi, K. N. et al. mTORC1 and mTORC2 selectively regulate CD8+ T cell differentiation. J. Clin. Invest. 125, 2090–2108 (2015).
Hedrick, S. M., Hess Michelini, R., Doedens, A. L., Goldrath, A. W. & Stone, E. L. FOXO transcription factors throughout T cell biology. Nat. Rev. Immunol. 12, 649–661 (2012).
Ouyang, W. et al. Foxo proteins cooperatively control the differentiation of Foxp3+ regulatory T cells. Nat. Immunol. 11, 618–627 (2010).
Ouyang, W. et al. Novel Foxo1-dependent transcriptional programs control Treg cell function. Nature 491, 554–559 (2012).
Ouyang, W., Beckett, O., Flavell, R. A. & Li, M. O. An essential role of the Forkhead-box transcription factor Foxo1 in control of T cell homeostasis and tolerance. Immunity 30, 358–371 (2009).
Kerdiles, Y. M. et al. Foxo1 links homing and survival of naive T cells by regulating L-selectin, CCR7 and interleukin 7 receptor. Nat. Immunol. 10, 176–184 (2009).
Riou, C. et al. Convergence of TCR and cytokine signaling leads to FOXO3a phosphorylation and drives the survival of CD4+ central memory T cells. J. Exp. Med. 204, 79–91 (2007).
You, H. et al. FOXO3a-dependent regulation of Puma in response to cytokine/growth factor withdrawal. J. Exp. Med. 203, 1657–1663 (2006).
Brunet, A. et al. Akt promotes cell survival by phosphorylating and inhibiting a Forkhead transcription factor. Cell 96, 857–868 (1999).
Sullivan, J. A., Kim, E. H., Plisch, E. H., Peng, S. L. & Suresh, M. FOXO3 regulates CD8 T cell memory by T cell-intrinsic mechanisms. PLoS Pathog. 8, e1002533 (2012).
Sullivan, J. A., Kim, E. H., Plisch, E. H. & Suresh, M. FOXO3 regulates the CD8 T cell response to a chronic viral infection. J. Virol. 86, 9025–9034 (2012).
van Grevenynghe, J. et al. Transcription factor FOXO3a controls the persistence of memory CD4+ T cells during HIV infection. Nat. Med. 14, 266–274 (2008).
Zhang, X. et al. FOXO1 is an essential regulator of pluripotency in human embryonic stem cells. Nat. Cell. Biol. 13, 1092–1099 (2011).
Hess Michelini, R., Doedens, A. L., Goldrath, A. W. & Hedrick, S. M. Differentiation of CD8 memory T cells depends on Foxo1. J. Exp. Med. 210, 1189–1200 (2013).
Ferber, E. C. et al. FOXO3a regulates reactive oxygen metabolism by inhibiting mitochondrial gene expression. Cell Death Differ. 19, 968–979 (2012).
Zhang, W. et al. FoxO1 regulates multiple metabolic pathways in the liver: effects on gluconeogenic, glycolytic, and lipogenic gene expression. J. Biol. Chem. 281, 10105–10117 (2006).
Greer, E. L. et al. The energy sensor AMP-activated protein kinase directly regulates the mammalian FOXO3 transcription factor. J. Biol. Chem. 282, 30107–30119 (2007).
Wu, Z. et al. Mechanisms controlling mitochondrial biogenesis and respiration through the thermogenic coactivator PGC-1. Cell 98, 115–124 (1999).
Finck, B. N. & Kelly, D. P. PGC-1 coactivators: inducible regulators of energy metabolism in health and disease. J. Clin. Invest. 116, 615–622 (2006).
Daitoku, H., Yamagata, K., Matsuzaki, H., Hatta, M. & Fukamizu, A. Regulation of PGC-1 promoter activity by protein kinase B and the forkhead transcription factor FKHR. Diabetes 52, 642–649 (2003).
Olmos, Y. et al. Mutual dependence of Foxo3a and PGC-1α in the induction of oxidative stress genes. J. Biol. Chem. 284, 14476–14484 (2009).
Long, S. A. et al. Partial exhaustion of CD8 T cells and clinical response to teplizumab in new-onset type 1 diabetes. Sci. Immunol. 1, eaai7793 (2016).
Freitag, J., Berod, L., Kamradt, T. & Sparwasser, T. Immunometabolism and autoimmunity. Immunol. Cell. Biol. 94, 925–934 (2016).
Zhao, J. et al. A missense variant in NCF1 is associated with susceptibility to multiple autoimmune diseases. Nat. Genet. 49, 433–437 (2017).
Gelderman, K. A., Hultqvist, M., Holmberg, J., Olofsson, P. & Holmdahl, R. T cell surface redox levels determine T cell reactivity and arthritis susceptibility. Proc. Natl Acad. Sci. USA 103, 12831–12836 (2006).
Tkachev, V. et al. Programmed death-1 controls T cell survival by regulating oxidative metabolism. J. Immunol. 194, 5789–5800 (2015).
Lee, J.C. et al. Genome-wide association study identifies distinct genetic contributions to prognosis and susceptibility in Crohn’s disease. Nat. Genet. 49, 262–268 (2017).
Lee, J. C. et al. Human SNP links differential outcomes in inflammatory and infectious disease to a FOXO3-regulated pathway. Cell 155, 57–69 (2013).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
McKinney, E.F., Smith, K.G.C. Metabolic exhaustion in infection, cancer and autoimmunity. Nat Immunol 19, 213–221 (2018). https://doi.org/10.1038/s41590-018-0045-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0045-y
This article is cited by
-
Upregulation of PD-1/PD-L1 and downregulation of immune signaling pathways lead to more severe visceral leishmaniasis in undernutrition mice
Parasites & Vectors (2024)
-
Ex vivo treatment with poly (I:C) alleviates the exhausted phenotype of tumor-infiltrating TCD8+ cells of gastric cancer patients
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
-
Glycolysis-cholesterol metabolic axis in immuno-oncology microenvironment: emerging role in immune cells and immunosuppressive signaling
Cell & Bioscience (2023)
-
Regulation of CD8+ T memory and exhaustion by the mTOR signals
Cellular & Molecular Immunology (2023)
-
T cell cholesterol efflux suppresses apoptosis and senescence and increases atherosclerosis in middle aged mice
Nature Communications (2022)