Abstract
Defective autophagy is linked to diseases such as rheumatoid arthritis, lupus and inflammatory bowel disease (IBD). However, the mechanisms by which autophagy limits inflammation remain poorly understood. Here we found that loss of the autophagy-related gene Atg16l1 promoted accumulation of the adaptor TRIF and downstream signaling in macrophages. Multiplex proteomic profiling identified SQSTM1 and Tax1BP1 as selective autophagy-related receptors that mediated the turnover of TRIF. Knockdown of Tax1bp1 increased production of the cytokines IFN-β and IL-1β. Mice lacking Atg16l1 in myeloid cells succumbed to lipopolysaccharide-mediated sepsis but enhanced their clearance of intestinal Salmonella typhimurium in an interferon receptor–dependent manner. Human macrophages with the Crohn’s disease–associated Atg16l1 variant T300A exhibited more production of IFN-β and IL-1β. An elevated interferon-response gene signature was observed in patients with IBD who were resistant to treatment with an antibody to the cytokine TNF. These findings identify selective autophagy as a key regulator of signaling via the innate immune system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Nicholson, J. K. et al. Host-gut microbiota metabolic interactions. Science 336, 1262–1267 (2012).
Kabat, A. M. et al. The autophagy gene Atg16l1 differentially regulates Treg and TH2 cells to control intestinal inflammation. eLife 5, e12444 (2016).
Martinez, J. et al. Noncanonical autophagy inhibits the autoinflammatory, lupus-like response to dying cells. Nature 533, 115–119 (2016).
Ktistakis, N. T. & Tooze, S. A. Digesting the expanding mechanisms of autophagy. Trends Cell. Biol. 26, 624–635 (2016).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12, 1–222 (2016).
Shpilka, T., Weidberg, H., Pietrokovski, S. & Elazar, Z. Atg8: an autophagy-related ubiquitin-like protein family. Genome Biol. 12, 226 (2011).
Khaminets, A., Behl, C. & Dikic, I. Ubiquitin-dependent and independent signals in selective autophagy. Trends Cell. Biol. 26, 6–16 (2016).
Harley, J. B. et al. Genome-wide association scan in women with systemic lupus erythematosus identifies susceptibility variants in ITGAM, PXK, KIAA1542 and other loci. Nat. Genet. 40, 204–210 (2008).
Lapaquette, P., Guzzo, J., Bretillon, L. & Bringer, M. A. Cellular and molecular connections between autophagy and inflammation. Mediat. Inflamm. 2015, 398483 (2015).
Deretic, V. Autophagy as an innate immunity paradigm: expanding the scope and repertoire of pattern recognition receptors. Curr. Opin. Immunol. 24, 21–31 (2012).
Gay, N. J., Symmons, M. F., Gangloff, M. & Bryant, C. E. Assembly and localization of Toll-like receptor signalling complexes. Nat. Rev. Immunol. 14, 546–558 (2014).
O’Neill, L. A., Golenbock, D. & Bowie, A. G. The history of Toll-like receptors — redefining innate immunity. Nat. Rev. Immunol. 13, 453–460 (2013).
Kagan, J. C. et al. TRAM couples endocytosis of Toll-like receptor 4 to the induction of interferon-beta. Nat. Immunol. 9, 361–368 (2008).
Blander, J. M. A long-awaited merger of the pathways mediating host defence and programmed cell death. Nat. Rev. Immunol. 14, 601–618 (2014).
Delgado, M. A., Elmaoued, R. A., Davis, A. S., Kyei, G. & Deretic, V. Toll-like receptors control autophagy. EMBO J. 27, 1110–1121 (2008).
Suet Ting Tan, R. et al. The synergy in cytokine production through MyD88-TRIF pathways is co-ordinated with ERK phosphorylation in macrophages. Immunol. Cell. Biol. 91, 377–387 (2013).
Tong, A. J. et al. A stringent systems approach uncovers gene-specific mechanisms regulating inflammation. Cell 165, 165–179 (2016).
Saitoh, T. et al. Loss of the autophagy protein Atg16L1 enhances endotoxin-induced IL-1beta production. Nature 456, 264–268 (2008).
Liu, S. et al. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science 347, aaa2630 (2015).
Oshiumi, H., Matsumoto, M., Funami, K., Akazawa, T. & Seya, T.TICAM-1, an adaptor molecule that participates in Toll-like receptor 3-mediated interferon-β induction. Nat. Immunol. 4, 161–167 (2003).
Jounai, N. et al. The Atg5 Atg12 conjugate associates with innate antiviral immune responses. Proc. Natl Acad. Sci. USA 104, 14050–14055 (2007).
Mathew, R. et al. Functional role of autophagy-mediated proteome remodeling in cell survival signaling and innate immunity. Mol. Cell. 55, 916–930 (2014).
Grimm, W. A. et al. The Thr300Ala variant in ATG16L1 is associated with improved survival in human colorectal cancer and enhanced production of type I interferon. Gut 65, 456–464 (2016).
Tal, M. C. et al. Absence of autophagy results in reactive oxygen species-dependent amplification of RLR signaling. Proc. Natl Acad. Sci. USA 106, 2770–2775 (2009).
Kagan, J. C., Magupalli, V. G. & Wu, H. SMOCs: supramolecular organizing centres that control innate immunity. Nat. Rev. Immunol. 14, 821–826 (2014).
Li, J. et al. The RIP1/RIP3 necrosome forms a functional amyloid signaling complex required for programmed necrosis. Cell 150, 339–350 (2012).
Lamour, G. et al. Mapping the broad structural and mechanical properties of amyloid fibrils. Biophys. J. 112, 584–594 (2017).
Funami, K., Sasai, M., Oshiumi, H., Seya, T. & Matsumoto, M. Homo-oligomerization is essential for Toll/interleukin-1 receptor domain-containing adaptor molecule-1-mediated NF-κB and interferon regulatory factor-3 activation. J. Biol. Chem. 283, 18283–18291 (2008).
Gentle, I. E. et al. TIR-domain-containing adapter-inducing interferon-β (TRIF) forms filamentous structures, whose pro-apoptotic signalling is terminated by autophagy. FEBS J. 284, 1987–2003 (2017).
Rogov, V., Dötsch, V., Johansen, T. & Kirkin, V. Interactions between autophagy receptors and ubiquitin-like proteins form the molecular basis for selective autophagy. Mol. Cell. 53, 167–178 (2014).
Iha, H. et al. Inflammatory cardiac valvulitis in TAX1BP1-deficient mice through selective NF-κB activation. EMBO J. 27, 629–641 (2008).
Parvatiyar, K., Barber, G. N. & Harhaj, E. W. TAX1BP1 and A20 inhibit antiviral signaling by targeting TBK1-IKKi kinases. J. Biol. Chem. 285, 14999–15009 (2010).
Whang, M. I. et al. The ubiquitin binding protein TAX1BP1 mediates autophagasome induction and the metabolic transition of activated T cells. Immunity 46, 405–420 (2017).
Tan, Y. & Kagan, J. C. A cross-disciplinary perspective on the innate immune responses to bacterial lipopolysaccharide. Mol. Cell. 54, 212–223 (2014).
Hampe, J. et al. A genome-wide association scan of nonsynonymous SNPs identifies a susceptibility variant for Crohn disease in ATG16L1. Natl Genet. 39, 207–211 (2007).
Dejager, L. et al. Pharmacological inhibition of type I interferon signaling protects mice against lethal sepsis. J. Infect. Dis. 209, 960–970 (2014).
Robinson, N. et al. Type I interferon induces necroptosis in macrophages during infection with Salmonella enterica serovar Typhimurium. Natl Immunol. 13, 954–962 (2012).
Owen, K. A., Anderson, C. J. & Casanova, J. E. Salmonella suppresses the TRIF-dependent type I interferon response in macrophages. mBio 7, e02051–e15 (2016).
Sotolongo, J. et al. Host innate recognition of an intestinal bacterial pathogen induces TRIF-dependent protective immunity. J. Exp. Med. 208, 2705–2716 (2011).
Lassen, K. G. et al. Atg16L1 T300A variant decreases selective autophagy resulting in altered cytokine signaling and decreased antibacterial defense. Proc. Natl Acad. Sci. USA 111, 7741–7746 (2014).
Murthy, A. et al. A Crohn’s disease variant in Atg16l1 enhances its degradation by caspase 3. Nature 506, 456–462 (2014).
Noble, C. L. et al. Regional variation in gene expression in the healthy colon is dysregulated in ulcerative colitis. Gut 57, 1398–1405 (2008).
Noble, C. L. et al. Characterization of intestinal gene expression profiles in Crohn’s disease by genome-wide microarray analysis. Inflamm. Bowel Dis. 16, 1717–1728 (2010).
Arijs, I. et al. Mucosal gene expression of antimicrobial peptides in inflammatory bowel disease before and after first infliximab treatment. PLoS. One 4, e7984 (2009).
Toedter, G. et al. Gene expression profiling and response signatures associated with differential responses to infliximab treatment in ulcerative colitis. Am. J. Gastroenterol. 106, 1272–1280 (2011).
Deretic, V., Saitoh, T. & Akira, S. Autophagy in infection, inflammation and immunity. Nat. Rev. Immunol. 13, 722–737 (2013).
Gomes, L. C. & Dikic, I. Autophagy in antimicrobial immunity. Mol. Cell. 54, 224–233 (2014).
Yang, Q. et al. TRIM32-TAX1BP1-dependent selective autophagic degradation of TRIF negatively regulates TLR3/4-mediated innate immune responses. PLoS. Pathog. 13, e1006600 (2017).
Kirkpatrick, D. S. et al. Phosphoproteomic characterization of DNA damage response in melanoma cells following MEK/PI3K dual inhibition. Proc. Natl Acad. Sci. USA 110, 19426–19431 (2013).
Zhuang, G. et al. Phosphoproteomic analysis implicates the mTORC2-FoxO1 axis in VEGF signaling and feedback activation of receptor tyrosine kinases. Sci. Signal. 6, ra25 (2013).
Choi, M. et al. MSstats: an R package for statistical analysis of quantitative mass spectrometry-based proteomic experiments. Bioinformatics 30, 2524–2526 (2014).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2007).
Irizarry, R. A. et al. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 31, e15 (2003).
Ritchie, M. E. et al. A comparison of background correction methods for two-colour microarrays. Bioinformatics 23, 2700–2707 (2007).
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Bourgon, R., Gentleman, R. & Huber, W. Independent filtering increases detection power for high-throughput experiments. Proc. Natl Acad. Sci. USA 107, 9546–9551 (2010).
Bueno, R. et al. Comprehensive genomic analysis of malignant pleural mesothelioma identifies recurrent mutations, gene fusions and splicing alterations. Nat. Genet. 48, 407–416 (2016).
Lee, C. A. & Falkow, S. The ability of Salmonella to enter mammalian cells is affected by bacterial growth state. Proc. Natl Acad. Sci. USA 87, 4304–4308 (1990).
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010).
Srinivasan, K. et al. Untangling the brain’s neuroinflammatory and neurodegenerative transcriptional responses. Nat. Commun. 7, 11295 (2016).
Acknowledgements
We thank K. Cherry and E. Chua for assistance with animal husbandry; S. Gierke and C. Chalouni for technical expertise in microscopy; M. Zepeda for assistance with human donor studies; A. Scherl and L. Diehl for pathology support; and J. Arron and I. Mellman for critical evaluation of the manuscript.
Author information
Authors and Affiliations
Contributions
M.S and A.M. conceptualized the study and designed experiments; M.S, J.L. and A.M. conducted experiments; E.V. performed mass spectrometry data analysis; J.M.B. designed and performed mass spectrometry; I.P., A.W. and Y.K. performed in vivo LPS sepsis and S. Typhimurium infection studies; Y.S. performed RNA-Seq analysis; J.A.H. and M.K. analyzed clinical data sets; B.M. coordinated in vivo studies; D.S.K. coordinated mass spectrometry, guided data analysis and discussed the study; M.v.L.C. guided data analysis, discussed the study and critiqued the manuscript; and A.M. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
All authors except J.M.B. are employees of Genentech; J.M.B. is an employee of MedImmune.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Loss of Atg16l1 enhances TRIF-dependent IFN-β production by BMDMs.
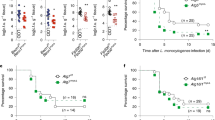
a, IFN-β and IL-1β levels (ELISA) in cell culture media of BMDMs treated with indicated TLR ligands for 6 hours. b, Immunoblots depicting TBK1 and IRF3 phosphorylation in Atg16l1 cKO BMDMs over 24 hours of LPS treatment. c, mRNA transcript levels of Ifnb1 in BMDMs treated with indicated TLR ligands for 3 hours. d, mRNA transcript levels of Il1b, Tnf and Il6 in BMDMs treated with indicated TLR ligands. e, IFN-α levels (ELISA) in cell culture media of BMDMs treated with indicated TLR ligands for 24 hours. f, Immunoblots of Ticam1 knockdown by lentiviral shRNA in BMDMs. g, IFN-β and TNF levels (ELISA) in cell culture media of BMDMs following TRIF knockdown as in (e) followed by LPS treatment for 24 hours. ELISA and qPCR data are pooled from 4 independent experiments, each performed in triplicate. Ticam1 knockdown data are pooled from 3 independent experiments, each performed in triplicate. Immunoblots are representative of 3 independent experiments. Bars in scatter plots depict mean. *P<0.05; **P<0.01; ***P<0.001 (Student’s two-sided t test without multiple comparisons correction).
Supplementary Figure 2 Elevated production of IFN-β and IL-1β by Atg16l1- or Atg5l-deficient BMDCs.
IFN-β, IL-1β and IL-6 levels (ELISA) in cell culture media of Control, Atg16l1-cKO and Atg5l-cKO BMDCs treated with indicated TLR ligands for 24 hours. Data are pooled from 3 independent experiments, each performed in triplicate. Bars in scatter plots depict mean. *P<0.05; **P<0.01 (Student’s two-sided t test without multiple comparisons correction).
Supplementary Figure 3 Depletion of the cytosolic RNA sensor RIG-I does not impact TLR4-mediated production of IFN-β by Atg16l1-deficient BMDMs.
a, Immunoblots of RIG-I following lentiviral shRNA-mediated knockdown of its encoding mRNA Ddx58. b, IFN-β and TNF levels (ELISA) in cell culture media of BMDMs following Ddx58 knockdown and LPS treatment for 24 hours. Immunoblots are representative of 3 independent experiments. ELISA data are pooled from 3 independent experiments, each performed in triplicate. Bars in scatter plots depict mean. *P<0.01 (Student’s two-sided t test without multiple comparisons correction).
Supplementary Figure 4 Accumulation of TRIF oligomers in Atg16l1-deficient BMDMs.
a, SDS-PAGE immunoblots of TRIF, MYD88 and αTubulin over 6 hours of LPS treatment. High molecular weight forms of TRIF are depicted as TRIF oligomers. b, SDS-PAGE following treatment of lysates with 150mM NaOH degrades TRIF and a majority of cellular protein in BMDMs. BMDMs were assessed over 9 hours of cycloheximide (CHX) chase. Coomassie Blue staining depicts total input protein. Immunoblots and gels are representative of 3 independent experiments (a) and 2 independent experiments (b).
Supplementary Figure 5 TMT 10-plex mass spectrometry reveals accumulation of autophagy receptors and cargo in Atg16l1-deficient BMDMs over 24 h of LPS treatment.
a, Venn diagram showing the overlap between the 6272 identified proteins in three biological replicates of the multiplexed proteomics assay. b, Histogram of the unique peptides per protein distribution for each replicate of the multiplexed proteomics assay showing excellent identification consistency. 1588 out of 6272 proteins did not meet quality filtering criteria due to insufficient data or low signal and were excluded from statistical testing. c, Waterfall plot of log2-fold-changes for each of the five pairwise comparisons (Atg16l1 cKO/ Control for each post LPS treatment time point). Most proteins did not show appreciable changes (|Log2FC| > 1) between genotypes. d, Heat map of additional proteins showing differential abundance (|log2-fold-change| > 1, no p-value threshold) in at least one timepoint of the 24-hour time course, classified by down (blue)- or up-regulation (red) in Atg16l1 deficient BMDMs and the impact of Atg16l1 deficiency on basal and signal-induced autophagy. e, Line plots of log2-scaled protein abundance changes over 24 hours of LPS stimulation for all core autophagy machinery proteins identified by multiplexed proteomics. Complete listing of all identified macroautophagy-associated proteins identified by TMT-MS is provided in Supplementary Table 3. f, Boxplots depicting a significant mean-shift of interferon beta response-associated proteins (GO003546) in Atg16l1 deficient BMDMs. Box plot limits represent the first and third quartile, the center line indicates the median, and whiskers denote the largest and smallest values no more than 1.5 times the interquartile range from the limits. Data beyond whiskers are outliers. *P<0.001 (Student’s one-sided t test – “IFN-β greater than other”). g, TNF levels in cell culture media of Control and Atg16l1 cKO BMDMs following Tax1bp1 knockdown and 24 hours of LPS treatment. Data are pooled from 3 independent experiments, each performed in triplicate. Bars in scatter plots depict mean.
Supplementary Figure 6 Maintenance of the T300A variant (genotype GG) of Atg16l1 in healthy people across genetically distinct populations.
Charts depict relative frequency of reference/non-risk (AA), heterozygous (AG) and risk/T300A (GG) genotypes of Atg16l1 in 5 major human populations annotated by the 1000 Genomes Project.
Supplementary Figure 7 Validation cohort identifying IRG signature as a biomarker of IBD progression.
Heatmap - interferon-regulated genes in ulcerative colitis samples in GSE23597. Log2 normalized signal for each gene was standardized to give a mean of 0 and standard deviation of 1. Samples are ordered by infliximab response (R=Responder, blue; NR=Non-responder, red) and pre- v. post-treatment time point (Pre-tx=Pre-treatment, grey; Post-tx=Post-treatment, black). Box plot – IRG signature score for interferon responsive genes. Log2 relative signature scores are shown for infliximab responders and non-responders, both prior to and after treatment. Box plot limits represent the first and third quartile, the center line indicates the median, and whiskers denote the largest and smallest values no more than 1.5 times the interquartile range from the limits. R Pre-tx, n = 25; R Post-tx, n = 20; NR Pre-tx, n = 7; NR Post-tx, n = 7. *P< 0.05 (linear regression testing differences between responders and non-responders in pre-treatment samples, or pre- and post-treatment samples in responders and non-responders, separately).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1-7
Supplementary Table 1: Atg16l1 cKO vs. Control protein abundance changes post-LPS stimulus.
Time point matched pairwise comparisons between triplicates of Atg16l1 cKO and Control BMDMs for all 4,684 quantified proteins. The comparison is expressed as a log2 fold-change of the estimated protein abundance difference from TMT data between the respective conditions and the significance of that change. For each pairwise comparison between Control and Atg16l1 cKO samples and corresponding time points, the Fold Change (Log2) and the results of an ANOVA test were reported
Supplementary Table 2: Gene Ontology Pathway Enrichment groups.
Results of the Gene Ontology enrichment, calculated by GOstats, for all clusters depicted in Figure 3d. Clusters are defined as 1) Proteins basally upregulated in Atg16l1 cKO BMDMs; 2) Proteins upregulated in Atg16l1 cKO BMDMs following LPS treatment, and 3) Proteins downregulated in Atg16l1 cKO BMDMs.
Supplementary Table 3: Macroautophagy Components
Identified proteins classified into macro-autophagy related categories, indicating whether they are changing significantly in the multiplexed proteomics assay (heat map illustrated in Fig. 3c; core autophagy components listed in Supplementary Fig. 5e).
Rights and permissions
About this article
Cite this article
Samie, M., Lim, J., Verschueren, E. et al. Selective autophagy of the adaptor TRIF regulates innate inflammatory signaling. Nat Immunol 19, 246–254 (2018). https://doi.org/10.1038/s41590-017-0042-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-017-0042-6
This article is cited by
-
Unraveling the role of autophagy regulation in Crohn's disease: from genetic mechanisms to potential therapeutics
Advanced Biotechnology (2024)
-
Sepsis-induced immunosuppression: mechanisms, diagnosis and current treatment options
Military Medical Research (2022)
-
sGRP78 enhances selective autophagy of monomeric TLR4 to regulate myeloid cell death
Cell Death & Disease (2022)
-
Role of autophagy on cancer immune escape
Cell Communication and Signaling (2021)
-
Pattern recognition receptors in health and diseases
Signal Transduction and Targeted Therapy (2021)