Abstract
The sensor RIG-I detects double-stranded RNA derived from RNA viruses. Although RIG-I is also known to have a role in the antiviral response to DNA viruses, physiological RNA species recognized by RIG-I during infection with a DNA virus are largely unknown. Using next-generation RNA sequencing (RNAseq), we found that host-derived RNAs, most prominently 5S ribosomal RNA pseudogene 141 (RNA5SP141), bound to RIG-I during infection with herpes simplex virus 1 (HSV-1). Infection with HSV-1 induced relocalization of RNA5SP141 from the nucleus to the cytoplasm, and virus-induced shutoff of host protein synthesis downregulated the abundance of RNA5SP141-interacting proteins, which allowed RNA5SP141 to bind RIG-I and induce the expression of type I interferons. Silencing of RNA5SP141 strongly dampened the antiviral response to HSV-1 and the related virus Epstein-Barr virus (EBV), as well as influenza A virus (IAV). Our findings reveal that antiviral immunity can be triggered by host RNAs that are unshielded following depletion of their respective binding proteins by the virus.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Goubau, D., Deddouche, S. & Reis e Sousa, C. Cytosolic sensing of viruses. Immunity 38, 855–869 (2013).
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Loo, Y. M. & Gale, M. Jr. Immune signaling by RIG-I-like receptors. Immunity 34, 680–692 (2011).
Wu, J. & Chen, Z. J. Innate immune sensing and signaling of cytosolic nucleic acids. Annu. Rev. Immunol. 32, 461–488 (2014).
Hornung, V. et al. 5′-Triphosphate RNA is the ligand for RIG-I. Science 314, 994–997 (2006).
Pichlmair, A. et al. RIG-I-mediated antiviral responses to single-stranded RNA bearing 5′-phosphates. Science 314, 997–1001 (2006).
Schlee, M. et al. Recognition of 5′ triphosphate by RIG-I helicase requires short blunt double-stranded RNA as contained in panhandle of negative-strand virus. Immunity 31, 25–34 (2009).
Rehwinkel, J. et al. RIG-I detects viral genomic RNA during negative-strand RNA virus infection. Cell 140, 397–408 (2010).
Baum, A., Sachidanandam, R. & García-Sastre, A. Preference of RIG-I for short viral RNA molecules in infected cells revealed by next-generation sequencing. Proc. Natl. Acad. Sci. USA 107, 16303–16308 (2010).
Runge, S. et al. In vivo ligands of MDA5 and RIG-I in measles virus-infected cells. PLoS Pathog. 10, e1004081 (2014).
Goubau, D. et al. Antiviral immunity via RIG-I-mediated recognition of RNA bearing 5′-diphosphates. Nature 514, 372–375 (2014).
Saito, T., Owen, D. M., Jiang, F., Marcotrigiano, J. & Gale, M. Jr. Innate immunity induced by composition-dependent RIG-I recognition of hepatitis C virus RNA. Nature 454, 523–527 (2008).
Ablasser, A. et al. RIG-I-dependent sensing of poly(dA:dT) through the induction of an RNA polymerase III-transcribed RNA intermediate. Nat. Immunol. 10, 1065–1072 (2009).
Chiu, Y. H., Macmillan, J. B. & Chen, Z. J. RNA polymerase III detects cytosolic DNA and induces type I interferons through the RIG-I pathway. Cell 138, 576–591 (2009).
Chan, Y. K. & Gack, M. U. Viral evasion of intracellular DNA and RNA sensing. Nat. Rev. Microbiol. 14, 360–373 (2016).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Unterholzner, L. et al. IFI16 is an innate immune sensor for intracellular DNA. Nat. Immunol. 11, 997–1004 (2010).
Peri, P. et al. Herpes simplex virus type 1 Us3 gene deletion influences toll-like receptor responses in cultured monocytic cells. Virol. J. 5, 140 (2008).
Szymański, M., Barciszewska, M. Z., Erdmann, V. A. & Barciszewski, J. 5S rRNA: structure and interactions. Biochem. J. 371, 641–651 (2003).
Cui, S. et al. The C-terminal regulatory domain is the RNA 5′-triphosphate sensor of RIG-I. Mol. Cell 29, 169–179 (2008).
Hiscott, J. Triggering the innate antiviral response through IRF-3 activation. J. Biol. Chem. 282, 15325–15329 (2007).
Civril, F. et al. The RIG-I ATPase domain structure reveals insights into ATP-dependent antiviral signalling. EMBO Rep. 12, 1127–1134 (2011).
Glaunsinger, B. A. & Ganem, D. E. Messenger RNA turnover and its regulation in herpesviral infection. Adv. Virus Res. 66, 337–394 (2006).
Ciganda, M. & Williams, N. Eukaryotic 5S rRNA biogenesis. Wiley Interdiscip. Rev. RNA 2, 523–533 (2011).
Johnson, D. C. & Baines, J. D. Herpesviruses remodel host membranes for virus egress. Nat. Rev. Microbiol. 9, 382–394 (2011).
West, J. A. et al. The long noncoding RNAs NEAT1 and MALAT1 bind active chromatin sites. Mol. Cell 55, 791–802 (2014).
Niazi, A. K. et al. Targeting nucleic acids into mitochondria: progress and prospects. Mitochondrion 13, 548–558 (2013).
Kwong, A. D. & Frenkel, N. Herpes simplex virus-infected cells contain a function(s) that destabilizes both host and viral mRNAs. Proc. Natl. Acad. Sci. USA 84, 1926–1930 (1987).
Simonin, D., Diaz, J. J., Massé, T. & Madjar, J. J. Persistence of ribosomal protein synthesis after infection of HeLa cells by herpes simplex virus type 1. J. Gen. Virol. 78, 435–443 (1997).
Smiley, J. R. Herpes simplex virus virion host shutoff protein: immune evasion mediated by a viral RNase? J. Virol. 78, 1063–1068 (2004).
Strahle, L., Garcin, D. & Kolakofsky, D. Sendai virus defective-interfering genomes and the activation of interferon-beta. Virology 351, 101–111 (2006).
Gack, M. U. et al. Influenza A virus NS1 targets the ubiquitin ligase TRIM25 to evade recognition by the host viral RNA sensor RIG-I. Cell Host Microbe 5, 439–449 (2009).
Ayllon, J. & García-Sastre, A. The NS1 protein: a multitasking virulence factor. Curr. Top. Microbiol. Immunol. 386, 73–107 (2015).
Malathi, K., Dong, B., Gale, M. Jr. & Silverman, R. H. Small self-RNA generated by RNase L amplifies antiviral innate immunity. Nature 448, 816–819 (2007).
Nabet, B. Y. et al. Exosome RNA unshielding couples stromal activation to pattern recognition receptor signaling in cancer. Cell 170, 352–366 (2017).
Schlee, M. & Hartmann, G. Discriminating self from non-self in nucleic acid sensing. Nat. Rev. Immunol. 16, 566–580 (2016).
Barrat, F. J., Elkon, K. B. & Fitzgerald, K. A. Importance of nucleic acid recognition in inflammation and autoimmunity. Annu. Rev. Med. 67, 323–336 (2016).
Gack, M. U. et al. TRIM25 RING-finger E3 ubiquitin ligase is essential for RIG-I-mediated antiviral activity. Nature 446, 916–920 (2007).
Shapira, S. D. et al. A physical and regulatory map of host-influenza interactions reveals pathways in H1N1 infection. Cell 139, 1255–1267 (2009).
Marquitz, A. R., Mathur, A., Shair, K. H. & Raab-Traub, N. Infection of Epstein-Barr virus in a gastric carcinoma cell line induces anchorage independence and global changes in gene expression. Proc. Natl. Acad. Sci. USA 109, 9593–9598 (2012).
Tischer, B. K., Smith, G. A. & Osterrieder, N. En passant mutagenesis: a two step markerless red recombination system. Methods Mol. Biol. 634, 421–430 (2010).
Poon, A. P. & Roizman, B. Differentiation of the shutoff of protein synthesis by virion host shutoff and mutant gamma (1)34.5 genes of herpes simplex virus 1. Virology 229, 98–105 (1997).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucleic Acids Res. 44, W3–W10 (2016). W1.
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Andrews, S. FastQC: a quality control tool for high throughput sequence data. Available from: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2010).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Dillies, M. A. et al. A comprehensive evaluation of normalization methods for Illumina high-throughput RNA sequencing data analysis. Brief. Bioinform. 14, 671–683 (2013).
R Core Team. R: A language and environment for statistical computing. (R Foundation for Statistical Computing, Vienna, Austria, 2013).
Wickham, H. Ggplot2: Elegant Graphics for Data Analysis (Springer, New York, 2009).
Gack, M. U. et al. Roles of RIG-I N-terminal tandem CARD and splice variant in TRIM25-mediated antiviral signal transduction. Proc. Natl. Acad. Sci. USA 105, 16743–16748 (2008).
Lin, R., Génin, P., Mamane, Y. & Hiscott, J. Selective DNA binding and association with the CREB binding protein coactivator contribute to differential activation of alpha/beta interferon genes by interferon regulatory factors 3 and 7. Mol. Cell. Biol. 20, 6342–6353 (2000).
Hatzivassiliou, E., Cardot, P., Zannis, V. I. & Mitsialis, S. A. Ultraspiracle, a Drosophila retinoic X receptor alpha homologue, can mobilize the human thyroid hormone receptor to transactivate a human promoter. Biochemistry 36, 9221–9231 (1997).
Dillon, P. J. et al. Tousled-like kinases modulate reactivation of gammaherpesviruses from latency. Cell Host Microbe 13, 204–214 (2013).
Cui, C. et al. Prediction and identification of herpes simplex virus 1-encoded microRNAs. J. Virol. 80, 5499–5508 (2006).
Lässig, C. et al. ATP hydrolysis by the viral RNA sensor RIG-I prevents unintentional recognition of self-RNA. eLife 4, 1–20 (2015).
Maharaj, N. P., Wies, E., Stoll, A. & Gack, M. U. Conventional protein kinase C-α (PKC-α) and PKC-β negatively regulate RIG-I antiviral signal transduction. J. Virol. 86, 1358–1371 (2012).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Acknowledgements
We thank B. Roizman (University of Chicago) for HSV-1Δvhs mutant and revertant; Y. Kawaguchi (University of Tokyo) for the bacterial artificial chromosome clone of HSV-1; A. Garcia-Sastre (Mount Sinai) for IAV wild-type and ΔNS1 recombinant virus; N. Raab-Traub (UNC-Chapel Hill) for AGS-EBV cells; D. Knipe (Harvard University) for the antibody to ICP8; W. Azab for help with manuscript preparation; and K. Waraska and A. Diallo for discussions. Supported by the US National Institutes of Health (R21 AI133361 and R01 AI087846 to M.U.G.), the German Research Foundation (fellowship SP 1600/1-1 to K.M.J.S.; and HO2489-8 to K.-P.H.) and BioSysNet (K.-P.H. and C.L.).
Author information
Authors and Affiliations
Contributions
J.J.C. and M.U.G. conceived of the study, interpreted data and wrote the manuscript; J.J.C. performed and analyzed all experiments, except those in Figs. 1c–e and 4b and Supplementary Fig. 4b (performed and analyzed by K.M.J.S.), Supplementary Fig. 2b (performed and analyzed by K.M.J.S. and M.v.G.) and Supplementary Fig. 7 (performed and analyzed by M.v.G.); C.L. and K.-P.H. performed the in vitro ATP hydrolysis experiment; and T.H. and N.O. provided HSV-1mut.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 RIG-I, cGAS and IFI16 sense HSV-1 infection in a temporally distinct manner.
(a) Left, middle left: qRT-PCR analysis of IFNB1 and TNF mRNA in primary NHLF cells transfected with either non-targeting control siRNA (si.Ctrl) or RIG-I-, cGAS-, or IFI16-specific siRNAs (si.RIG-I, si.cGAS, si.IFI16) for 30 h and then infected with HSV-1 (MOI 0.1) for the indicated times. Results were normalized to 18S rRNA, and fold induction is shown relative to mock-infected control cells. Right three panels: Knockdown efficiency of endogenous RIG-I (DDX58), cGAS (MB21D1) and IFI16 was confirmed by qRT-PCR. (b) qRT-PCR analysis of IFNB1, IFIH1, OASL1, RSAD2, IFIT2, and CCL5 mRNA in Ddx58 –/– and Ddx58 +/+ MEFs infected with HSV-1 (MOI 1 each) for the indicated times. Results were normalized and shown as in (a). Data represent mean and s.d. of n = 3 biological replicates, and are representative of at least two independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001 (unpaired t-test); NS, statistically not significant; ND, not detected.
Supplementary Figure 2 Endogenous RNA5SP141 purified from cells is immunostimulatory.
(a) Equal pulldown (PD) of FLAG–RIG-I and FLAG-GFP from the experiment shown in Figures 1a and 1b was confirmed by immunoblot (IB) with anti-FLAG. (b) qRT-PCR analysis of IFNB1 mRNA in primary NHLF cells transfected for 20 h with RNA purified by streptavidin-PD from total cellular RNA using 3′-biotinylated locked nucleic acid (LNA) oligos specific for RNA5SP141. RNAs purified using LNA oligos designed to capture a nonsense ‘scramble’ sequence and the highly abundant small nuclear RNA RNU1-1 that is not known to be immunostimulatory served as negative controls. (c) Validation of the LNA oligos targeting RNA5SP141 (LNA_RNA5SP141), RNU1-1 (LNA_RNU1-1), or ‘scramble’ RNA (LNA_Scramble). Representative enrichment of RNU1-1 and RNA5SP141 by streptavidin-PD as described in (b). Data are representative of two independent experiments (mean and s.d. of n = 3 biological replicates in (b), n = 3 technical replicates in (c)). **P < 0.01 (unpaired t-test). NS, statistically not significant.
Supplementary Figure 3 In vitro–transcribed RNA5SP141 is immunostimulatory.
(a) Predicted secondary structure of RNA5SP141 modeled using the Vienna RNAfold web server. MFE, minimum free energy. 5′-ppp, 5′-triphosphate. (b) Forward (F) and reverse (R) DNA oligonucleotides used to construct in vitro transcription templates for RNA5SP141, parental 5S rRNA, and ‘scrambled’ RNA. (c) Virtual gel image of the indicated in vitro–transcribed RNAs analyzed by Agilent 2100 Bioanalyzer. (d) IFNB1 transcripts in primary NHLF cells that were either mock-transfected or transfected with the indicated amounts of in vitro–transcribed RNA5SP141 or RABVLe (positive control), determined by qRT-PCR analysis at 18 h post-transfection. Data in (d) represent mean and s.d. of n = 3 biological replicates, and are representative of three independent experiments.
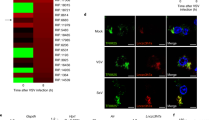
Supplementary Figure 4 RNA5SP141 expression during HSV-1 infection.
(a) Left: qRT-PCR analysis of IFNB1 mRNA in primary NHLF cells transfected with either non-targeting control siRNA (si.Ctrl), or siRNAs targeting RIG-I (si.RIG-I) or MDA5 (si.MDA5), followed 30 h later by transfection for 16 h with 1 pmol of the indicated in vitro–transcribed RNAs, or 0.05 μg/ml HMW-poly(I:C) which served as a control. Middle, right: Knockdown efficiency of endogenous RIG-I (DDX58) and MDA5 (IFIH1) was confirmed by qRT-PCR. (b) Relative expression of RNA5SP141 in HEK 293T cells infected with HSV-1WT (MOI 1) for 16 h, as compared to uninfected cells, determined by RNAseq analysis. Red boundaries represent ± 2-fold change in gene expression. (c) qRT-PCR analysis of RNA5SP141 transcripts in primary NHLF cells infected with HSV-1WT (MOI 1) for the indicated times. (d,e) qRT-PCR analysis of IL6 (d) and IL8 (e) transcripts in primary NHLF cells that were mock-infected or infected with HSV-1Δvhs or HSV-1WT (revertant) (MOI 10 each) for 16 h. Results were normalized to 18S rRNA, and fold induction is shown relative to values for mock-infected control cells. Data are representative of at least two independent experiments (mean and s.d. of n = 3 biological replicates in (a) and (c-e)).
Supplementary Figure 5 Depletion of RNA5SP141 dampens the cytokine response induced by infection with HSV-1 but not that induced by infection with SeV.
(a) Left: qRT-PCR analysis of TNF mRNA in primary NHLF cells transfected with either non-targeting control siRNA (si.Ctrl), or siRNAs targeting RIG-I (si.RIG-I) or RNA5SP141 (si.5SP141) for 72 h and then infected with HSV-1WT (MOI 0.1) for 16 h. Middle, right: Knockdown efficiency of endogenous RIG-I (DDX58) and RNA5SP141 was confirmed by qRT-PCR. (b) ELISA of IFN-β (left) and CCL5 (right) in the supernatants of primary NHLF cells that were transfected with either non-targeting control siRNA (si.Ctrl), or siRNAs targeting RIG-I (si.RIG-I) or RNA5SP141 (si.5SP141) for 72 h, followed by infection with SeV (50 HAU/ml) for 24 h. Data are representative of two independent experiments (mean and s.d. of n = 3 biological replicates in (a), and n = 2 biological replicates in (b)). *P < 0.05, **P < 0.01, ***P < 0.001 (unpaired t-test). NS, statistically not significant; ND, not detected.
Supplementary Figure 6 Knockdown of RIG-I and RNA5SP141 with gapmers.
Representative knockdown efficiency of endogenous RIG-I (DDX58) (left) and RNA5SP141 (right) in primary NHLF cells achieved by transfection of two different gapmers targeting RIG-I (Gap RIG-I_1 and Gap RIG-I_2) or RNA5SP141 (Gap 5SP141_1 and Gap 5SP141_2), determined by qRT-PCR at 72 h post-transfection. Non-targeting control gapmer (Gap NT) served as control. Data represent mean and s.d. of n = 3 technical replicates, and are representative of two independent experiments.
Supplementary Figure 7 Cytokine induction after reactivation of EBV in AGS-EBV cells.
(a-d) IFNB1, TNF, IL6, and IL8 transcripts in AGS-EBV cells treated with 2.5 mM sodium butyrate (NaB) for the indicated times to induce EBV reactivation, assessed by qRT-PCR analysis. (e) Efficient EBV reactivation was confirmed by determining the transcript amounts of EBV early gene BMRF1 (EA-D) by qRT-PCR analysis. Data represent mean and s.d. of n = 3 technical replicates.
Supplementary Figure 8 Silencing of RNA5SP141 diminishes the antiviral response induced by infection with IAV.
(a) Representative knockdown efficiency of RIG-I (DDX58) and RNA5SP141 in primary NHLF cells achieved by transfection of siRNAs targeting RIG-I (si.RIG-I) or RNA5SP141 (si.5SP141), assessed by qRT-PCR at 72 h post-transfection. (b) IFNB1, IFIT2, and CCL5 transcripts in HEK 293T cells transfected with non-targeting control siRNA (si.Ctrl), or siRNAs targeting RIG-I or RNA5SP141 (si.RIG-I or si.5SP141) followed by infection with IAV (MOI 0.1) for 16 h, determined by qRT-PCR. (c) Representative knockdown efficiency of RIG-I (DDX58) and RNA5SP141 in HEK 293T cells achieved by transfection of siRNAs targeting RIG-I (si.RIG-I) or RNA5SP141 (si.5SP141), assessed by qRT-PCR at 72 h post-transfection. (d) qRT-PCR analysis of IFNB1 mRNA in HEK 293T cells transfected with siRNAs as in (b) and then infected with IAVΔNS1 (MOI 0.1) for 16 h. Data represent mean and s.d. of n = 3 technical replicates (a,c) or n = 3 biological replicates (b,d), and are representative of two (a-c) or three (d) independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001 (unpaired t-test).
Supplementary information
Integrated Supplementary Figures
Supplementary Figures 1–8
Rights and permissions
About this article
Cite this article
Chiang, J.J., Sparrer, K.M.J., van Gent, M. et al. Viral unmasking of cellular 5S rRNA pseudogene transcripts induces RIG-I-mediated immunity. Nat Immunol 19, 53–62 (2018). https://doi.org/10.1038/s41590-017-0005-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-017-0005-y
This article is cited by
-
DUSP4 modulates RIG-I- and STING-mediated IRF3-type I IFN response
Cell Death & Differentiation (2024)
-
Epstein-Barr virus infection: the micro and macro worlds
Virology Journal (2023)
-
Exploiting RIG-I-like receptor pathway for cancer immunotherapy
Journal of Hematology & Oncology (2023)
-
Cellular state landscape and herpes simplex virus type 1 infection progression are connected
Nature Communications (2023)
-
Cellular functions of eukaryotic RNA helicases and their links to human diseases
Nature Reviews Molecular Cell Biology (2023)