Abstract
The hierarchy of human hemopoietic progenitor cells that produce lymphoid and granulocytic–monocytic (myeloid) lineages is unclear. Multiple progenitor populations produce lymphoid and myeloid cells, but they remain incompletely characterized. Here we demonstrated that lympho-myeloid progenitor populations in cord blood — lymphoid-primed multi-potential progenitors (LMPPs), granulocyte-macrophage progenitors (GMPs) and multi-lymphoid progenitors (MLPs) — were functionally and transcriptionally distinct and heterogeneous at the clonal level, with progenitors of many different functional potentials present. Although most progenitors had the potential to develop into only one mature cell type (‘uni-lineage potential’), bi- and rarer multi-lineage progenitors were present among LMPPs, GMPs and MLPs. Those findings, coupled with single-cell expression analyses, suggest that a continuum of progenitors execute lymphoid and myeloid differentiation, rather than only uni-lineage progenitors’ being present downstream of stem cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Dykstra, B. et al. Long-term propagation of distinct hematopoietic differentiation programs in vivo. Cell Stem Cell 1, 218–229 (2007).
Challen, G. A., Boles, N. C., Chambers, S. M. & Goodell, M. A. Distinct hematopoietic stem cell subtypes are differentially regulated by TGF-beta1. Cell Stem Cell 6, 265–278 (2010).
Benz, C. et al. Hematopoietic stem cell subtypes expand differentially during development and display distinct lymphopoietic programs. Cell Stem Cell 10, 273–283 (2012).
Chen, J. Y. et al. Hoxb5 marks long-term haematopoietic stem cells and reveals a homogenous perivascular niche. Nature 530, 223–227 (2016).
Oguro, H., Ding, L. & Morrison, S. J. SLAM family markers resolve functionally distinct subpopulations of hematopoietic stem cells and multipotent progenitors. Cell Stem Cell 13, 102–116 (2013).
Sanjuan-Pla, A. et al. Platelet-biased stem cells reside at the apex of the haematopoietic stem-cell hierarchy. Nature 502, 232–236 (2013).
Yamamoto, R. et al. Clonal analysis unveils self-renewing lineage-restricted progenitors generated directly from hematopoietic stem cells. Cell 154, 1112–1126 (2013).
Sun, J. et al. Clonal dynamics of native haematopoiesis. Nature 514, 322–327 (2014).
Busch, K. et al. Fundamental properties of unperturbed haematopoiesis from stem cells in vivo. Nature 518, 542–546 (2015).
Sawai, C. M. et al. Hematopoietic stem cells are the major source of multilineage hematopoiesis in adult animals. Immunity 45, 597–609 (2016).
Yu, V. W. et al. Epigenetic memory underlies cell-autonomous heterogeneous behavior of hematopoietic stem cells. Cell 167, 1310–1322 (2016).
Wilson, A. et al. Hematopoietic stem cells reversibly switch from dormancy to self-renewal during homeostasis and repair. Cell 135, 1118–1129 (2008).
Cabezas-Wallscheid, N. et al. Identification of regulatory networks in HSCs and their immediate progeny via integrated proteome, transcriptome, and DNA methylome analysis. Cell Stem Cell 15, 507–522 (2014).
Pietras, E. M. et al. Functionally distinct subsets of lineage-biased multipotent progenitors control blood production in normal and regenerative conditions. Cell Stem Cell 17, 35–46 (2015).
Majeti, R., Park, C. Y. & Weissman, I. L. Identification of a hierarchy of multipotent hematopoietic progenitors in human cord blood. Cell Stem Cell 1, 635–645 (2007).
Notta, F. et al. Distinct routes of lineage development reshape the human blood hierarchy across ontogeny. Science 351, aab2116 (2016).
Adolfsson, J. et al. Identification of Flt3+ lympho-myeloid stem cells lacking erythro-megakaryocytic potential a revised road map for adult blood lineage commitment. Cell 121, 295–306 (2005).
Månsson, R. et al. Molecular evidence for hierarchical transcriptional lineage priming in fetal and adult stem cells and multipotent progenitors. Immunity 26, 407–419 (2007).
Guo, G. et al. Mapping cellular hierarchy by single-cell analysis of the cell surface repertoire. Cell Stem Cell 13, 492–505 (2013).
Naik, S. H. et al. Diverse and heritable lineage imprinting of early haematopoietic progenitors. Nature 496, 229–232 (2013).
Doulatov, S. et al. Revised map of the human progenitor hierarchy shows the origin of macrophages and dendritic cells in early lymphoid development. Nat. Immunol. 11, 585–593 (2010).
Goardon, N. et al. Coexistence of LMPP-like and GMP-like leukemia stem cells in acute myeloid leukemia. Cancer Cell 19, 138–152 (2011).
Kohn, L. A. et al. Lymphoid priming in human bone marrow begins before expression of CD10 with upregulation of L-selectin. Nat. Immunol. 13, 963–971 (2012).
Görgens, A. et al. Multipotent hematopoietic progenitors divide asymmetrically to create progenitors of the lymphomyeloid and erythromyeloid lineages. Stem Cell Rep. 5, 154–155 (2015).
Velten, L. et al. Human haematopoietic stem cell lineage commitment is a continuous process. Nat. Cell Biol. 19, 271–281 (2017).
Perié, L., Duffy, K. R., Kok, L., de Boer, R. J. & Schumacher, T. N. The branching point in erythro-myeloid differentiation. Cell 163, 1655–1662 (2015).
Berardi, A. C. et al. Individual CD34+CD38lowCD19–CD10– progenitor cells from human cord blood generate B lymphocytes and granulocytes. Blood 89, 3554–3564 (1997).
Ichii, M. et al. The density of CD10 corresponds to commitment and progression in the human B lymphoid lineage. PLoS One 5, e12954 (2010).
Farlik, M. et al. DNA methylation dynamics of human hematopoietic stem cell differentiation. Cell Stem Cell 19, 808–822 (2016).
Lee, J. et al. Restricted dendritic cell and monocyte progenitors in human cord blood and bone marrow. J. Exp. Med. 212, 385–399 (2015).
Pronk, C. J. et al. Elucidation of the phenotypic, functional, and molecular topography of a myeloerythroid progenitor cell hierarchy. Cell Stem Cell 1, 428–442 (2007).
Galy, A., Travis, M., Cen, D., Chen, B. & Human, T. Human T, B, natural killer, and dendritic cells arise from a common bone marrow progenitor cell subset. Immunity 3, 459–473 (1995).
Manz, M. G., Miyamoto, T., Akashi, K. & Weissman, I. L. Prospective isolation of human clonogenic common myeloid progenitors. Proc. Natl. Acad. Sci. USA 99, 11872–11877 (2002).
Reinisch, A. et al. A humanized bone marrow ossicle xenotransplantation model enables improved engraftment of healthy and leukemic human hematopoietic cells. Nat. Med. 22, 812–821 (2016).
Novershtern, N. et al. Densely interconnected transcriptional circuits control cell states in human hematopoiesis. Cell 144, 296–309 (2011).
Wilson, N. K. et al. Combined single-cell functional and gene expression analysis resolves heterogeneity within stem cell populations. Cell Stem Cell 16, 712–724 (2015).
Haghverdi, L., Buettner, F. & Theis, F. J. Diffusion maps for high-dimensional single-cell analysis of differentiation data. Bioinformatics 31, 2989–2998 (2015).
Moignard, V. et al. Decoding the regulatory network of early blood development from single-cell gene expression measurements. Nat. Biotechnol. 33, 269–276 (2015).
Paul, F. et al. Transcriptional heterogeneity and lineage commitment in myeloid progenitors. Cell 163, 1663–1677 (2015).
Quek, L. et al. Genetically distinct leukemic stem cells in human CD34– acute myeloid leukemia are arrested at a hemopoietic precursor-like stage. J. Exp. Med. 213, 1513–1535 (2016).
Six, E. M. et al. Cytokines and culture medium have a major impact on human in vitro T-cell differentiation. Blood Cells Mol. Dis. 47, 72–78 (2011).
Calvo, J., BenYoucef, A., Baijer, J., Rouyez, M. C. & Pflumio, F. Assessment of human multi-potent hematopoietic stem/progenitor cell potential using a single in vitro screening system. PLoS One 7, e50495 (2012).
Woll, P. S. et al. Myelodysplastic syndromes are propagated by rare and distinct human cancer stem cells in vivo. Cancer Cell 25, 794–808 (2014).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Coifman, R. R. et al. Geometric diffusions as a tool for harmonic analysis and structure definition of data: diffusion maps. Proc. Natl. Acad. Sci. USA 102, 7426–7431 (2005).
Angerer, P. et al. destiny: diffusion maps for large-scale single-cell data in R. Bioinformatics 32, 1241–1243 (2016).
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010).
Yates, A. et al. Ensembl 2016. Nucleic Acids Res. 44 D1, D710–D716 (2016).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Brennecke, P. et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods 10, 1093–1095 (2013).
Lun, A. T., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75(2016).
Acknowledgements
We thank J.C. Zuniga-Pflucker for OP9-DL4 cells used for initial experiments; and the High-Throughput Genomics Group at the Wellcome Trust Centre for Human Genetics (funded by Wellcome Trust (090532/Z/09/Z)) for generation of sequencing data. Supported by the MRC (MHU Award G1000729 and MRC Disease Team Award 4050189188 to P.V.; PhD studentship to Z.A. and F.H.; Single Cell Award MR/M00919X/1), CRUK (Program Grant C7893/A12796 to P.V.; Program Grant C1163/A21762 to B.G.; Development Fund CRUKDF0176-DK to D.K. and P.V.; and CRUK Development Fund C5255/A20758 to B.S. and P.V.), Bloodwise (Specialist Program 13001 and Project Grant 12019), the Oxford Partnership Comprehensive Biomedical Research Centre (NIHR BRC Funding scheme), the US National Institutes of Health (R01CA188055 and U01HL099999 to R.M.), the New York Stem Cell Foundation (Robertson Investigator, R.M.), the Leukemia and Lymphoma Society (Scholar Award to R.M.) and the Austrian Science Fund (FWF) (Erwin-Schroedinger Research fellowship to A.R.).
Author information
Authors and Affiliations
Contributions
D.K. and B.S. contributed equally to this work and are listed alphabetically; F.H. and A.R. contributed equally to this work; D.K., B.S., Z.A. and P.V. designed the experiments; D.K., B.S., Z.A., A.R., M.S., L.Q. and N.G. performed experiments and analyzed data; F.H., G.O., Z.A., E.R. and S.T. performed bioinformatics and statistical analysis; J.D. and B.U. prepared samples; J.C., E.S., F.P., C.P., R.M. and B.G. provided reagents and materials; D.K., B.S. and P.V. wrote the paper; and all authors edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Immunophenotypic separation of eight distinct human hemopoietic stem–progenitor cell (HSPC) populations and in vitro functional lympho-myeloid potential of LMPPs, MLPs and GMPs
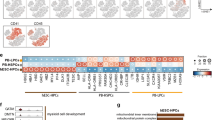
(a) Representative plot of flow sorting of human CB HSPCs. Population frequencies shown are the mean from 44 CB donors calculated as a percentage of Lin-CD34+ compartment. HSC, hemopoietic stem cell; MPP, multipotent progenitor; LMPP, lymphoid-primed multi-potential progenitor; MLP, multi-lymphoid progenitor; CMP, common myeloid progenitor; GMP, granulocyte-monocyte progenitor; MEP, megakaryocyte-erythroid progenitor; B/NK, B-NK cell progenitor. (b) Representative flow cytometric analysis of the B cell, NK, monocytic and granulocytic output of CB LMPP, MLP and GMP cultured for 2 weeks on SGF15/2. Frequencies shown are mean from 3 CB donors calculated as a percentage of human CD45+ cells. (c) Representative flow cytometric analysis of T cell output of CB LMPP, MLP and GMP cultured for 5 weeks on SF7a. Frequencies shown are average from 5 CB donors calculated as a percentage of human CD45+ cells.
Supplementary Figure 2 LMPPs and GMPs are lympho-myeloid progenitors, while MLPs are mainly lymphoid progenitors in quantitative in vitro assays
(a) Experimental strategy for sorting, culture and analysis for quantitative in vitro assays in SGF15/2. Similar strategy was used for the other culture conditions. (b) LDA and single cell in vitro culture conditions used in the study. (c) Representative flow cytometric profiles of the outputs from the quantitative in vitro assays. Plots are gated on human CD45+ cells. (d) LDA plots showing the frequency (f: 1 in X cells can give rise to) of LMPP, MLP and GMP cells with B-cell, NK cell, monocytic and granulocytic potential. Plots are generated in R and the lines represent the estimates calculated using ELDA software. (e) Total cloning efficiency (left) of single LMPP, MLP and GMP in S7T2GM/G/M culture (LMPP: 69/96 cells, MLP: 4/52, GMP: 98/110). Significance defined using Fisher’s exact test. Cloning efficiency of lymphoid (Ly, middle) and myeloid lineages (My, right). Bars indicate total cloning efficiency; filled portion indicates the proportion of lymphoid (lymphoid plus mixed clones) or myeloid potential (myeloid plus mixed clones). Mean ± SD is shown. Significance is defined using student’s t-test. (f) Single-, (g) bi- and (h) multi-lineage single cell outputs in S7T2GM/G/M culture, presented as percentage of the positive wells. (i) Lymphoid (Ly), myeloid (My) and lympho-myeloid (Ly-My) outputs presented as percentage of all plated single cell LMPP, MLP and GMP wells in S7T2GM/G/M culture. (j) Summary of lineage outputs from single LMPP, MLP and GMP. For the LDA data were obtained from 4 CB donors, for the single cell assay in S7T2GM/G/M culture data were obtained from 2 CB donors.
Supplementary Figure 3 Human LMPPs, MLPs and GMPs have distinct differentiation potential in vivo
(a) Experimental strategy for generation of human ossicles in NSG mice and human progenitor transplantation. (b) Gating strategy for the sorting of human progenitors for in vivo transplantation. (c) Number of cells of LMPP, MLP and GMP populations injected per ossicle. (d) Correlation between the number of transplanted cells and the myeloid/lymphoid ratio of the output cells.
Supplementary Figure 4 Transcriptional profiling of human HSPCs in bulk shows the distinct transcriptional patterns of human progenitor populations
(a) Heatmap showing hierarchical clustering and the expression of all genes by HSPC populations. Expression values are normalized per gene (per row). (b) Plot comparing the eigenvalues for each principle component (PC) for the PC analysis (PCA) of HSPC using the top 300 ANOVA genes (black) and 300 randomly selected genes from a randomized expression matrix (red). (c) 3D PCA plot showing HSPC populations using the top 300 ANOVA genes. Percentage variance for each PC is shown. (d) PCA plots showing HSPC populations using between 1,000 and 10,000 genes with the highest variance across all HSPC populations. Percentage variance for each PC is shown. (e) Table of differentially expressed genes in one versus one comparisons of HSPCs. Genes are upregulated in a population column versus row. (f) Heatmap showing the expression of genes recently identified as being expressed by lineage primed hemopoietic cells25. Genes are color-coded according to their classification. Expression values are normalized per gene. (g-h) Heatmap showing the expression of (g) hemopoietic-related transcription factors that were differentially expressed between the MLP and GMP and (h) cytokine and chemokine related genes by the LMPP, GMP and MLP. Genes affiliated with the lymphoid or myeloid lineages are color-coded (lymphoid: orange, myeloid: green) and genes associated with immune function are labeled in black. Expression values are normalized per gene.
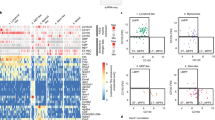
Supplementary Figure 5 Transcriptional heterogeneity of single lympho-myeloid progenitor cells (a) Heatmap showing hierarchical clustering on genes (rows) and single cells (columns), based on qRT-PCR gene expression analysis
(b) Top, cell type composition of the 3 clusters identified in the dendrogram in (a). Middle, differentially expressed genes between cluster pairs. Bottom, expected number of cells per cluster, based on the functional assays. (c) Diffusion map dimensionality reductions colored in by cell type (left) and cluster membership (right), based on qRT-PCR gene expression analysis. (d) Heatmap showing clustering of single LMPP, GMP and MLP using the 55 most highly and variably expressed genes between clusters. Single cell RNA-sequencing data from two donors were used to generate the clusters. The heat map shows clustering from one of two. Data from the other donor is in Fig. 5b. Log-normalised gene expression (rows) for each single cell (columns) is shown. (e) Cell type composition of the 3 clusters identified in the heatmap in (d) and Fig. 5b. (f) PCA plot colored by cell type (top) or cluster membership (bottom). For single cell qRT-PCR gene expression analysis: data from 4 CB donors. For single cell RNA sequencing analysis: data from 2 donors (donor 1 in Fig. 5, donor 2 in current figure).
Supplementary Figure 6 Further functional purification of the lympho-myeloid progenitor populations (a) CD10 fluorescence intensity in LMPPs in the 3 clusters identified from clustering of single cell RNA Seq data presented in . Wilcoxon rank sum test is used to define significance.
(b) Histograms showing expression of CD10 and CD45RA for LMPP with different functional output. The percentage cut-offs used to define the new sorting strategy for LMPPly and LMPPmix are shown. Thresholds were defined based on maximum CD10 and CD45RA expression of LMPPs with myeloid output. (c) Total cloning efficiency (left) of single MLP, LMPP, LMPPly, LMPPmix and GMP in SF7b culture (LMPPly: 26/108 cells, LMPPmix: 48/100). Significance defined using Fisher’s exact test. Cloning efficiency of lymphoid (Ly, middle) and myeloid lineages (My, right). Bars indicate total cloning efficiency; filled portion indicates the proportion of lymphoid (lymphoid plus mixed clones) or myeloid potential (myeloid plus mixed clones). Mean ± SD is shown. Significance is defined using students t-test. (d) Single-, (e) bi- and (f) multi-lineage outputs from single cells in SF7b culture, presented as percentage of the positive wells. (g) CD38 fluorescence intensity in GMPs in the 3 clusters identified from clustering of single cell RNA Seq data presented in Fig. 5b. Wilcoxon rank sum test is used to define significance. (h) Correlation between expression of selected genes and cell surface marker expression in GMPs. ρ indicates Spearman’s rank correlation coefficient values and p is corresponding p-value from cor.test function in R. (i) Histograms showing expression of CD38 for GMP with different functional output. The percentage cut-off used to define the new sorting strategy for GMP CD38hi and CD38mid are shown. For functional assay based on the revised LMPP sorting strategy (SF7b culture) data are from 6 CB donors; for LMPP, MLP and GMP controls data from 9 CB donors (including 3 CB donors used for Fig. 2f-j).
Supplementary Figure 7 Models of human lympho-myeloid differentiation
(a) Lineage output of lympho-myeloid progenitors (b-c) Models of human hemopoietic differentiation and lineage diversification. Dashed arrows - hierarchical relationships not experimentally validated. (d) Novel model of human lympho-myeloid differentiation. Multiple, rare, functionally distinct bi- and multi-lineage lympho-myeloid progenitors (LMPs) could differentiate either directly from hemopoietic stem cells (HSC) or multi-potent progenitors (MPP). We have used the term LMP, rather than LMPP (lymphoid primed multi-potential progenitor) to describe these lympho-myeloid progenitors, as not all LMPs will be lymphoid biased. Multi-lineage LMPs are rare. Lymphoid only and myeloid only progenitors are shown below. Bi-lineage progenitors are more frequent and uni-potent progenitors are most common. The hierarchical relationships between LMP and lymphoid and myeloid only bi- and uni-lineage progenitors remain to be determined. This model also leaves open the question of when commitment to either lymphoid or myeloid fate occurs. Erythro-megakaryocytic differentiation leading to a MEP (megakaryocyte-erythroid progenitor), erythroid (BFU-E) and megakaryocytic progenitor (MkP) could occur directly from either HSC or MPP.
Supplementary information
Supplementary Figures
Supplementary Figures 1–7
Rights and permissions
About this article
Cite this article
Karamitros, D., Stoilova, B., Aboukhalil, Z. et al. Single-cell analysis reveals the continuum of human lympho-myeloid progenitor cells. Nat Immunol 19, 85–97 (2018). https://doi.org/10.1038/s41590-017-0001-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-017-0001-2
This article is cited by
-
Loss of hematopoietic progenitors heterogeneity is an adverse prognostic factor in lower-risk myelodysplastic neoplasms
Leukemia (2024)
-
Hematopoietic stem cells with granulo-monocytic differentiation state overcome venetoclax sensitivity in patients with myelodysplastic syndromes
Nature Communications (2024)
-
Single-nucleus RNA sequencing and deep tissue proteomics reveal distinct tumour microenvironment in stage-I and II cervical cancer
Journal of Experimental & Clinical Cancer Research (2023)
-
Acute myeloid leukemia: from NGS, through scRNA-seq, to CAR-T. dissect cancer heterogeneity and tailor the treatment
Journal of Experimental & Clinical Cancer Research (2023)
-
Multi-omic single-cell velocity models epigenome–transcriptome interactions and improves cell fate prediction
Nature Biotechnology (2023)