Abstract
G-protein-coupled receptors (GPCRs) are key regulators of human physiology and are the targets of many small-molecule research compounds and therapeutic drugs. While most of these ligands bind to their target GPCR with high affinity, selectivity is often limited at the receptor, tissue and cellular levels. Antibodies have the potential to address these limitations but their properties as GPCR ligands remain poorly characterized. Here, using protein engineering, pharmacological assays and structural studies, we develop maternally selective heavy-chain-only antibody (‘nanobody’) antagonists against the angiotensin II type I receptor and uncover the unusual molecular basis of their receptor antagonism. We further show that our nanobodies can simultaneously bind to angiotensin II type I receptor with specific small-molecule antagonists and demonstrate that ligand selectivity can be readily tuned. Our work illustrates that antibody fragments can exhibit rich and evolvable pharmacology, attesting to their potential as next-generation GPCR modulators.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The cryo-EM model and maps were deposited under the following accession numbers: AT118-H AT1R complex: PDB 8TH3, EMDB-41248; AT118-L AT1R complex: PDB 8TH4, EMDB-41249. Sequencing data can be found on GitHub: https://github.com/kruselab/AT118-library-sequencing. Source data are provided with this paper.
Code availability
The code for analyzing NGS sequencing data can be found on GitHub: https://github.com/kruselab/AT118-library-sequencing.
References
Mullard, A. 2022 FDA approvals. Nat. Rev. Drug Discov. 22, 83–88 (2023).
Hauser, A. S., Attwood, M. M., Rask-Andersen, M., Schioth, H. B. & Gloriam, D. E. Trends in GPCR drug discovery: new agents, targets and indications. Nat. Rev. Drug Discov. 16, 829–842 (2017).
Peterson, S. M. et al. Discovery and design of G protein-coupled receptor targeting antibodies. Expert Opin. Drug Discov. 18, 417–428 (2023).
Niwa, R. et al. Defucosylated chimeric anti-CC chemokine receptor 4 IgG1 with enhanced antibody-dependent cellular cytotoxicity shows potent therapeutic activity to T-cell leukemia and lymphoma. Cancer Res. 64, 2127–2133 (2004).
Smith, E. L. et al. GPRC5D is a target for the immunotherapy of multiple myeloma with rationally designed CAR T cells. Sci. Transl. Med. 11, eaau7746 (2019).
Irannejad, R. et al. Functional selectivity of GPCR-directed drug action through location bias. Nat. Chem. Biol. 13, 799–806 (2017).
Irannejad, R. et al. Conformational biosensors reveal GPCR signalling from endosomes. Nature 495, 534–538 (2013).
Stoeber, M. et al. A genetically encoded biosensor reveals location bias of opioid drug action. Neuron 98, 963–976 e965 (2018).
Kantor, E. D., Rehm, C. D., Haas, J. S., Chan, A. T. & Giovannucci, E. L. Trends in prescription drug use among adults in the United States from 1999–2012. JAMA 314, 1818–1831 (2015).
Bullo, M., Tschumi, S., Bucher, B. S., Bianchetti, M. G. & Simonetti, G. D. Pregnancy outcome following exposure to angiotensin-converting enzyme inhibitors or angiotensin receptor antagonists: a systematic review. Hypertension 60, 444–450 (2012).
Buse, M. G., Roberts, W. J. & Buse, J. The role of the human placenta in the transfer and metabolism of insulin. J. Clin. Invest. 41, 29–41 (1962).
McMahon, C. et al. Yeast surface display platform for rapid discovery of conformationally selective nanobodies. Nat. Struct. Mol. Biol. 25, 289–296 (2018).
McMahon, C. et al. Synthetic nanobodies as angiotensin receptor blockers. Proc. Natl Acad. Sci. USA 117, 20284–20291 (2020).
Hoefman, S., Ottevaere, I., Baumeister, J. & Sargentini-Maier, M. L. Pre-clinical intravenous serum pharmacokinetics of albumin binding and non-half-life extended nanobodies. Antibodies 4, 141–156 (2015).
Kelly, R. L. et al. High throughput cross-interaction measures for human IgG1 antibodies correlate with clearance rates in mice. mAbs 7, 770–777 (2015).
Harvey, E. P. et al. An in silico method to assess antibody fragment polyreactivity. Nat. Commun. 13, 7554 (2022).
Knauf, M. J. et al. Relationship of effective molecular size to systemic clearance in rats of recombinant interleukin-2 chemically modified with water-soluble polymers. J. Biol. Chem. 263, 15064–15070 (1988).
Sanchez, M. F., Els-Heindl, S., Beck-Sickinger, A. G., Wieneke, R. & Tampe, R. Photoinduced receptor confinement drives ligand-independent GPCR signaling. Science 371, eabb7657 (2021).
Martin, W. L., West, A. P. Jr., Gan, L. & Bjorkman, P. J. Crystal structure at 2.8 Å of an FcRn/heterodimeric Fc complex: mechanism of pH-dependent binding. Mol. Cell 7, 867–877 (2001).
Burvenich, I. J. et al. Cross-species analysis of Fc engineered anti-Lewis-Y human IgG1 variants in human neonatal receptor transgenic mice reveal importance of S254 and Y436 in binding human neonatal Fc receptor. mAbs 8, 775–786 (2016).
Lo, M. et al. Effector-attenuating substitutions that maintain antibody stability and reduce toxicity in mice. J. Biol. Chem. 292, 3900–3908 (2017).
Mukherjee, S. et al. Synthetic antibodies against BRIL as universal fiducial marks for single-particle cryoEM structure determination of membrane proteins. Nat. Commun. 11, 1598 (2020).
Bloch, J. S. et al. Development of a universal nanobody-binding Fab module for fiducial-assisted cryo-EM studies of membrane proteins. Proc. Natl Acad. Sci. USA 118, e2115435118 (2021).
Robertson, M. J. et al. Structure determination of inactive-state GPCRs with a universal nanobody. Nat. Struct. Mol. Biol. 29, 1188–1195 (2022).
Botte, M. et al. Cryo-EM structures of a LptDE transporter in complex with pro-macrobodies offer insight into lipopolysaccharide translocation. Nat. Commun. 13, 1826 (2022).
Wingler, L. M., McMahon, C., Staus, D. P., Lefkowitz, R. J. & Kruse, A. C. Distinctive activation mechanism for angiotensin receptor revealed by a synthetic nanobody. Cell 176, 479–490 e412 (2019).
Wingler, L. M. et al. Angiotensin and biased analogs induce structurally distinct active conformations within a GPCR. Science 367, 888–892 (2020).
Zhang, D. et al. Structural insights into angiotensin receptor signaling modulation by balanced and biased agonists. EMBO J. 42, e112940 (2023).
Ballesteros, J. A. W. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Methods Neurosci. 25, 366–428 (1995).
Feng, Y. H. et al. The docking of Arg2 of angiotensin II with Asp281 of AT1 receptor is essential for full agonism. J. Biol. Chem. 270, 12846–12850 (1995).
Zhang, H. et al. Structural basis for ligand recognition and functional selectivity at angiotensin receptor. J. Biol. Chem. 290, 29127–29139 (2015).
Zhang, H. et al. Structure of the angiotensin receptor revealed by serial femtosecond crystallography. Cell 161, 833–844 (2015).
Kenakin, T. New concepts in drug discovery: collateral efficacy and permissive antagonism. Nat. Rev. Drug Discov. 4, 919–927 (2005).
Valant, C., Felder, C. C., Sexton, P. M. & Christopoulos, A. Probe dependence in the allosteric modulation of a G protein-coupled receptor: implications for detection and validation of allosteric ligand effects. Mol. Pharmacol. 81, 41–52 (2012).
Maussang, D. et al. Llama-derived single variable domains (nanobodies) directed against chemokine receptor CXCR7 reduce head and neck cancer cell growth in vivo. J. Biol. Chem. 288, 29562–29572 (2013).
Jähnichen, S. et al. CXCR4 nanobodies (VHH-based single variable domains) potently inhibit chemotaxis and HIV-1 replication and mobilize stem cells. Proc. Natl Acad. Sci. USA 107, 20565–20570 (2010).
Boshuizen, R. S. et al. A combination of in vitro techniques for efficient discovery of functional monoclonal antibodies against human CXC chemokine receptor-2 (CXCR2). mAbs 6, 1415–1424 (2014).
Peng, L., Damschroder, M. M., Cook, K. E., Wu, H. & Dall’Acqua, W. F. Molecular basis for the antagonistic activity of an anti-CXCR4 antibody. mAbs 8, 163–175 (2016).
De Groof, T. W. M. et al. Targeting the latent human cytomegalovirus reservoir for T-cell-mediated killing with virus-specific nanobodies. Nat. Commun. 12, 4436 (2021).
Koth, C. M. et al. Molecular basis for negative regulation of the glucagon receptor. Proc. Natl Acad. Sci. USA 109, 14393–14398 (2012).
Hennen, S. et al. Structural insight into antibody-mediated antagonism of the glucagon-like peptide-1 receptor. Sci. Rep. 6, 26236 (2016).
Sarkar, K. et al. Modulation of PTH1R signaling by an ECD binding antibody results in inhibition of β-arrestin 2 coupling. Sci. Rep. 9, 14432 (2019).
Garces, F. et al. Molecular insight into recognition of the CGRPR complex by migraine prevention therapy Aimovig (erenumab). Cell Rep. 30, 1714–1723 e1716 (2020).
Cheng, R. K. Y. et al. Structural insight into allosteric modulation of protease-activated receptor 2. Nature 545, 112–115 (2017).
Wallukat, G. et al. Patients with preeclampsia develop agonistic autoantibodies against the angiotensin AT1 receptor. J. Clin. Invest. 103, 945–952 (1999).
Thway, T. M. et al. Antibodies from preeclamptic patients stimulate increased intracellular Ca2+ mobilization through angiotensin receptor activation. Circulation 110, 1612–1619 (2004).
Harris, J. A. et al. Selective G protein signaling driven by substance P-neurokinin receptor dynamics. Nat. Chem. Biol. 18, 109–115 (2022).
Asada, H. et al. Crystal structure of the human angiotensin II type 2 receptor bound to an angiotensin II analog. Nat. Struct. Mol. Biol. 25, 570–576 (2018).
Toyoda, Y. et al. Ligand binding to human prostaglandin E receptor EP(4) at the lipid-bilayer interface. Nat. Chem. Biol. 15, 18–26 (2019).
Hong, C. et al. Structures of active-state orexin receptor 2 rationalize peptide and small-molecule agonist recognition and receptor activation. Nat. Commun. 12, 815 (2021).
Toyoda, Y. et al. Structural basis of α1A-adrenergic receptor activation and recognition by an extracellular nanobody. Nat. Commun. 14, 3655 (2023).
Acelajado, M. C., Hughes, Z. H., Oparil, S. & Calhoun, D. A. Treatment of resistant and refractory hypertension. Circ. Res. 124, 1061–1070 (2019).
Proudfoot, A. E. Chemokine receptors: multifaceted therapeutic targets. Nat. Rev. Immunol. 2, 106–115 (2002).
Tao, Y. X. The melanocortin-4 receptor: physiology, pharmacology, and pathophysiology. Endocr. Rev. 31, 506–543 (2010).
Smith, J. S., Lefkowitz, R. J. & Rajagopal, S. Biased signalling: from simple switches to allosteric microprocessors. Nat. Rev. Drug Discov. 17, 243–260 (2018).
Yu, J. et al. Structural basis of μ-opioid receptor-targeting by a nanobody antagonist. Preprint at bioRxiv https://doi.org/10.1101/2023.12.06.570395 (2023).
Ma, Y. et al. Structure-guided discovery of a single-domain antibody agonist against human apelin receptor. Sci. Adv. 6, eaax7379 (2020).
Chavkin, C. & Goldstein, A. Specific receptor for the opioid peptide dynorphin: structure–activity relationships. Proc. Natl Acad. Sci. USA 78, 6543–6547 (1981).
Xu, Y. et al. Addressing polyspecificity of antibodies selected from an in vitro yeast presentation system: a FACS-based, high-throughput selection and analytical tool. Protein Eng. Des. Sel. 26, 663–670 (2013).
Gaspar, J. M. NGmerge: merging paired-end reads via novel empirically-derived models of sequencing errors. BMC Bioinformatics 19, 536 (2018).
Gietz, R. D. & Schiestl, R. H. High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 31–34 (2007).
Edelman, G. M. et al. The covalent structure of an entire γG immunoglobulin molecule. Proc. Natl Acad. Sci. USA 63, 78–85 (1969).
Barnea, G. et al. The genetic design of signaling cascades to record receptor activation. Proc. Natl Acad. Sci. USA 105, 64–69 (2008).
Staus, D. P. et al. Sortase ligation enables homogeneous GPCR phosphorylation to reveal diversity in β-arrestin coupling. Proc. Natl Acad. Sci. USA 115, 3834–3839 (2018).
Tsutsumi, N. et al. Structure of human Frizzled5 by fiducial-assisted cryo-EM supports a heterodimeric mechanism of canonical Wnt signaling. eLife 9, e58464 (2020).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
The PyMOL Molecular Graphics System, Version 3.0 (Schrödinger, 2024).
Morin, A. et al. Collaboration gets the most out of software. eLife 2, e01456 (2013).
Wu, E. L. et al. CHARMM-GUI membrane builder toward realistic biological membrane simulations. J. Comput. Chem. 35, 1997–2004 (2014).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Lee, J. et al. CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J. Chem. Theory Comput. 12, 405–413 (2016).
Jefferson, R. E. et al. Computational design of dynamic receptor-peptide signaling complexes applied to chemotaxis. Nat. Commun. 14, 2875 (2023).
McClendon, C. L., Hua, L., Barreiro, A. & Jacobson, M. P. Comparing conformational ensembles using the Kullback–Leibler divergence expansion. J. Chem. Theory Comput. 8, 2115–2126 (2012).
Acknowledgements
We thank R. Lefkowitz for allowing preliminary pharmacological experiments to be performed in his lab, M. Bao and X. Zhou for critical reading of the paper, A. Kossiakoff for the BAG2 anti-BRIL Fab and NabFab, and R. Walsh and M. Mayer for assistance and advice during cryo-EM data collection at the Harvard Center for Cryo-Electron Microscopy at Harvard Medical School. The SBGrid Consortium provided computation support of structural biology software. This work was funded by a Merck Fellowship from the Helen Hay Whitney Foundation (M.A.S.); a Hanna H. Gray Fellowship from HHMI (M.S.A.G.); NIH grants no. K99HD110612 (M.A.S.), no. DP5OD021345 (A.C.K.), no. R21HD101596 (A.C.K.), no. R01CA260415 (A.C.K.) and no. R01NS088566 (M.K.L.); the Vallee Foundation (A.C.K.); the Smith Family Foundation (A.C.K.); the Pew Charitable Trusts (L.M.W.); the Whitehead Foundation (L.M.W.); the New York Stem Cell Foundation (M.K.L); the William Randolph Hearst Fellowship (H.X.); the Swiss National Science Foundation (SNSF grants no. 31003A_182263 and no. 310030_208179) (P.B.); Swiss Cancer Research (grant no. KFS-4687-02-2019) (P.B.); funds from EPFL (P.B.); and the Ludwig Institute for Cancer Research (P.B.).
Author information
Authors and Affiliations
Contributions
M.A.S., D.P.S., C.M., L.M.W. and A.C.K. conceived of the project. M.A.S. generated the AT118 library and performed nanobody selections. J.D.H. and M.A.S. analyzed sequencing data. M.A.S., D.P.S. and L.M.W. purified all receptor constructs. M.A.S. and G.R.N. purified all nanobodies and Fab fragments. M.A.S., J.K. and L.M.W. performed radioligand binding assays. M.A.S. and G.R.N. performed flow cytometry binding and signaling assays. M.A.S. and D.P.S. designed AT118-Fc fusion constructs, which were purified by M.A.S. H.J. performed mouse blood pressure experiments under the supervision of H.A.R. H.X. carried out mouse placental transfer assays under the supervision of M.K.L. M.A.S. performed ELISA assays. M.A.S., S.M.S. and S.R. conceived and designed the cryo-EM screening strategy. M.A.S. and S.M.S. collected cryo-EM data. M.A.S., S.R. and M.S.A.G. processed cryo-EM data. M.A.S. modeled and analyzed the cryo-EM structures. S.Z. performed molecular dynamics simulations under the supervision of P.B. P.S. performed HPLC and HR-MS validation of small-molecule ligands. A.C.K. supervised all aspects of the research. M.A.S. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
A.C.K., C.M., L.M.W., D.P.S. and M.A.S. are co-inventors on a patent application for AT1R blocking nanobodies. A.C.K. is a cofounder and consultant for biotechnology companies Tectonic Therapeutic and Seismic Therapeutic, and also for the Institute for Protein Innovation, a nonprofit research institute. L.M.W. is a scientific advisor for Septerna. D.P.S. is a Septerna employee. C.M. is a Sanofi employee. P.B. holds patents and provisional patent applications in the field of engineered T-cell therapies and protein design. The other authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Haitao Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Engineering of high affinity AT1R nanobody antagonist with low non-specific binding.
a) Flowchart of nanobody selection. AT1R binders were enriched through two rounds of magnetic-activated cell sorting (MACS). Fluorescence-activated cell sorting (FACS) was used to isolate clone with low polyreactivity. A final FACS step enriched high-affinity AT1R binders. b) FACS round 1 plot. 1.38% of the population containing high-affinity AT1R binders with reduced polyspecificity were collected. c) FACS round 2 plot 0.7% of the population was collected containing high affinity AT1R binders. d) Binding of yeast-display library to FLAG-AT1R throughout each selection round. e) Distribution of AT1R-binding and polyreactive nanobodies in the yeast-display library throughout the selection process.
Extended Data Fig. 2 Effects of AT118-L and AT118-L-Fc fusion proteins on AT1R binding and signaling.
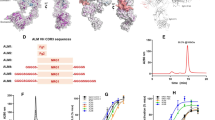
a) Binding of AT118-H variants displayed on yeast to detergent solubilized AT1R. Data are presented as mean ± SEM from three experiments. b) Non-specific binding of AT118-H variants displayed on yeast to biotinylated insect cell membrane polyspecificity reagent. Data are presented as mean ± SEM from three experiments. c–e) 3H-olmesartan competition experiments in Expi293 cell membranes containing AT1R with purified AT118-H variants. Variants containing D103N and S102G fail to displace olmesartan. The addition of V101D to D103N rescues the loss in pharmacological function. Data are presented as mean ± SEM from three experiments. Error is too small to be displayed if no bar is present. f) Accumulation of AT118-L F47T Y98F Fc fusion protein, that does not bind AT1R, in fetal serum. Data are presented as mean ± SEM from nine embryos from three separate litters for the control Fc (pMAS512, Supplementary Table 1) and eight embryos from two litters for the engineered Non-FcRn binding Fc (pMAS513, Supplementary Table 1).
Extended Data Fig. 3 Cryo-EM Construct Screening.
a) Fusion of AT118-H to the N-terminus of AT1R enhances total receptor expression. Data are presented as mean ± SEM from three experiments. b–f) Constructs screened for structure determination and representative 2-D class averages. b) AT118-AT1R fusion protein, c) Anti-nanobody Fab fragment bound to free AT118, but not AT118 in complex with AT1R23. d) MBP-AT118 in complex with AT1R25. e) AT118-AT1R-kappa opioid receptor (κOR) ICL3 fusion protein in complex nanobody 6 with an engineered alpaca framework, which binds κOR ICL3, anti-nanobody Fab, and anti-Fab nanobody23,24. f) AT118-AT1R-BRIL fusion protein in complex with anti-BRIL Fab and an anti-Fab nanobody22.
Extended Data Fig. 4 AT118-H AT1R Data Processing.
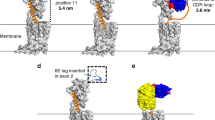
a) Size-exclusion chromatography trace b) SDS-PAGE gel under reducing conditions. c) cryo-EM data processing scheme, d) representative micrograph (n = 7,064, scale bar = 50 nm), and e) representative 2D class averages of AT118-H-AT1R-BRIL, anti-BRIL Fab, anti-Fab nanobody complex. Two independent purifications of this complex yielded similar size-exclusion and SDS-PAGE results. Fourier shell correlation (FSC) used to determine the f) global map and g) locally refined map resolutions. h) Local resolution estimate after local refinement.
Extended Data Fig. 5 Molecular Dynamics Simulations.
Distance landscape of a) AT118-H AT1R-BRIL and b) AT118-H AT1R between TM3 (residue 3.50) and TM6 (residue 6.34) and TM3 (residue 3.50) and TM7 (residue 7.53) displayed as individual runs (colored in purple, orange, blue, and yellow) and overall density. Active state (PDB ID: 6OS0) and inactive state (PDB ID: 4YAY) are provided for reference. Neither construct, AT1R or AT1R-BRIL in complex with AT118-H, visits an active like state. c) Examination of dihedral distributions of allosteric activation network within AT1R’s core for AT1R and AT1R-BRIL in complex with AT118-H plotted by Chi 1 angles. Kullback–Leibler (KL) divergence between the two constructs is zero, indicating that the BRIL fusion does not induce substantial conformational change in the activation network.
Extended Data Fig. 6 Binding of AT118-H and AT118-L with a broad panel of small molecule AT1R antagonists.
a) Molecular structures of small-molecule AT1R antagonists. b) Binding of AT118-H (orange) and AT118-L (purple) with a series of small-molecule AT1R antagonists. Error bars represent mean ± SEM from three independent experiments.
Extended Data Fig. 7 Allosteric effects of AT118-H and AT118-L.
a) Binding of AT118-H with modeled olmesartan. D103CDR3 of AT118-H would clash with olmesartan. Weak density for W842.60 is observed in the antagonist binding site in the orthosteric pocket of the AT118-H AT1R fusion protein structure. b) Binding of AT118-L with modeled ZD7155 (pink sticks, PDB 4YAY ref. 32). c) Allosteric effect of AT118-L on small molecule inhibition of AT1R activation. AT118-L potentiates the inhibitory effects of losartan (gray), but has no effect on olmesartan (green). Log EC50 data are expressed as mean ± SEM from three independent experiments. *p = 0.026 was determined with one-way repeated-measures ANOVA with Dunnett’s correction for multiple comparison.
Extended Data Fig. 8 AT118-L AT1R Data Processing.
a) Size exclusion trace, b) SDS-PAGE gel under reducing conditions, and c) cryo-EM data processing scheme of AT118-L AT1R-BRIL, anti-BRIL Fab, anti-Fab nanobody complex. One purification of this complex was performed and similar size-exclusion and SDS-PAGE results are in agreement with the analogous complex prepared in Extended Data Fig. 4. d) Fourier shell correlation (FSC) used to determine the global map resolution. e) FSC used to determine locally refined map resolution. f) Experimental density of losartan within the orthosteric binding pocket from locally refined map. g) Comparison of orthosteric binding pocket in AT118-L, Losartan, AT1R-BRIL complex (blue) and olmesartan bound AT1R (purple, PDB 4ZUD).
Extended Data Fig. 9 Antibody GPCR binding model.
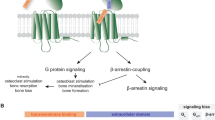
Nanobodies and other antibody fragments can adopt modular binding modes where one region mediates GPCR binding and another influences pharmacological function. The GPCR binding moiety can be formatted into a conventional antibody increasing avidity for the target GPCR or combined with secondary antibodies that recognize a tissue specific marker in a bispecific format. Pharmacokinetics, effector function, and tissue localization and delivery, can be further tuned by the antibodies constant Fc region.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3 and Tables 1–6.
Supplementary Data 1
Source data for Supplementary Table 2.
Source data
Source Data Fig. 1
Source data for Fig. 1.
Source Data Fig. 4
Source data for Fig. 4.
Source Data Fig. 6
Source data for Fig. 6.
Source Data Extended Data Fig. 2
Source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 6
Source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Source data for Extended Data Fig. 7.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Skiba, M.A., Sterling, S.M., Rawson, S. et al. Antibodies expand the scope of angiotensin receptor pharmacology. Nat Chem Biol (2024). https://doi.org/10.1038/s41589-024-01620-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41589-024-01620-6