Abstract
Pharmacological activation of voltage-gated ion channels by ligands serves as the basis for therapy and mainly involves a classic gating mechanism that augments the native voltage-dependent open probability. Through structure-based virtual screening, we identified a new scaffold compound, Ebio1, serving as a potent and subtype-selective activator for the voltage-gated potassium channel KCNQ2 and featuring a new activation mechanism. Single-channel patch-clamp, cryogenic-electron microscopy and molecular dynamic simulations, along with chemical derivatives, reveal that Ebio1 engages the KCNQ2 activation by generating an extended channel gate with a larger conductance at the saturating voltage (+50 mV). This mechanism is different from the previously observed activation mechanism of ligands on voltage-gated ion channels. Ebio1 caused S6 helices from residues S303 and F305 to perform a twist-to-open movement, which was sufficient to open the KCNQ2 gate. Overall, our findings provide mechanistic insights into the activation of KCNQ2 channel by Ebio1 and lend support for KCNQ-related drug development.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The 3D cryo-EM density maps of Ebio1- and Ebio-S1-bound KCNQ2 structures have been deposited in the Electron Microscopy Data Bank under the accession numbers EMD-35487 and EMD-38041, respectively. Atomic coordinates for the atomic models of Ebio1- and Ebio-S1-bound KCNQ2 structures have been deposited in the PDB under the accession numbers 8IJK and 8X43, respectively. All stable reagents generated in this study are available from the lead contact without restriction. Plasmids and strains are available from the authors upon request. Source data are provided with this paper.
References
Alsaloum, M., Higerd, G. P., Effraim, P. R. & Waxman, S. G. Status of peripheral sodium channel blockers for non-addictive pain treatment. Nat. Rev. Neurol. 16, 689–705 (2020).
Santos, R. et al. A comprehensive map of molecular drug targets. Nat. Rev. Drug Discov. 16, 19–34 (2017).
Schewe, M. et al. A pharmacological master key mechanism that unlocks the selectivity filter gate in K+ channels. Science 363, 875–880 (2019).
Wulff, H. & Zhorov, B. S. K+ Channel modulators for the treatment of neurological disorders and autoimmune diseases. Chem. Rev. 108, 1744–1773 (2008).
Wulff, H., Castle, N. A. & Pardo, L. A. Voltage-gated potassium channels as therapeutic targets. Nat. Rev. Drug Discov. 8, 982–1001 (2009).
Wulff, H., Christophersen, P., Colussi, P., Chandy, K. G. & Yarov-Yarovoy, V. Antibodies and venom peptides: new modalities for ion channels. Nat. Rev. Drug Discov. 18, 339–357 (2019).
Peters, H. C., Hu, H., Pongs, O., Storm, J. F. & Isbrandt, D. Conditional transgenic suppression of M channels in mouse brain reveals functions in neuronal excitability, resonance and behavior. Nat. Neurosci. 8, 51–60 (2005).
Wang, H. S. et al. KCNQ2 and KCNQ3 potassium channel subunits: molecular correlates of the M-channel. Science 282, 1890–1893 (1998).
Soh, H., Springer, K., Doci, K., Balsbaugh, J. L. & Tzingounis, A. V. KCNQ2 and KCNQ5 form heteromeric channels independent of KCNQ3. Proc. Natl Acad. Sci. USA 119, e2117640119 (2022).
Kuba, H., Yamada, R., Ishiguro, G. & Adachi, R. Redistribution of Kv1 and Kv7 enhances neuronal excitability during structural axon initial segment plasticity. Nat. Commun. 6, 8815 (2015).
Schwarz, J. R. et al. KCNQ channels mediate IKs, a slow K+ current regulating excitability in the rat node of Ranvier. J. Physiol. 573, 17–34 (2006).
Orhan, G. et al. Dominant-negative effects of KCNQ2 mutations are associated with epileptic encephalopathy. Ann. Neurol. 75, 382–394 (2014).
Vanoye, C. G. et al. High-throughput evaluation of epilepsy-associated KCNQ2 variants reveals functional and pharmacological heterogeneity. Jci. Insight 7, e156314 (2022).
Miceli, F. et al. Genotype-phenotype correlations in neonatal epilepsies caused by mutations in the voltage sensor of KV7.2 potassium channel subunits. Proc. Natl Acad. Sci. USA 110, 4386–4391 (2013).
Perucca, P. & Perucca, E. Identifying mutations in epilepsy genes: Impact on treatment selection. Epilepsy Res. 152, 18–30 (2019).
Lopez, J. P. et al. Ketamine exerts its sustained antidepressant effects via cell-type-specific regulation of Kcnq2. Neuron 110, 2283–2298 (2022).
Li, S. B. et al. Hyperexcitable arousal circuits drive sleep instability during aging. Science 375, eabh3021 (2022).
Tsuboi, D. et al. Dopamine drives neuronal excitability via KCNQ channel phosphorylation for reward behavior. Cell Rep. 40, 111309 (2022).
Huang, X. et al. Human amyotrophic lateral sclerosis excitability phenotype screen: target discovery and validation. Cell Rep. 35, 109224 (2021).
Maljevic, S., Wuttke, T. V. & Lerche, H. Nervous system KV7 disorders: breakdown of a subthreshold brake. J. Physiol. 586, 1791–1801 (2008).
Peretz, A. et al. Targeting the voltage sensor of Kv7.2 voltage-gated K+ channels with a new gating-modifier. Proc. Natl Acad. Sci. USA 107, 15637–15642 (2010).
Borgini, M., Mondal, P., Liu, R. & Wipf, P. Chemical modulation of Kv7 potassium channels. RSC Med. Chem. 12, 483–537 (2021).
Zhang, Q. et al. Dynamic PIP2 interactions with voltage sensor elements contribute to KCNQ2 channel gating. Proc. Natl Acad. Sci. USA 110, 20093–20098 (2013).
Li, X. et al. Molecular basis for ligand activation of the human KCNQ2 channel. Cell Res. 31, 52–61 (2021).
Abbott, G. W. KCNQs: ligand- and voltage-gated potassium channels. Front. Physiol. 11, 583 (2020).
Zheng, Y. et al. Structural insights into the lipid and ligand regulation of a human neuronal KCNQ channel. Neuron 110, 237–247 (2022).
Li, T. et al. Structural basis for the modulation of human KCNQ4 by small-molecule drugs. Mol. Cell 81, 25–37 (2021).
Stafstrom, C. E., Grippon, S. & Kirkpatrick, P. Ezogabine (retigabine). Nat. Rev. Drug Discov. 10, 729–730 (2011).
Catterall, W. A., Lenaeus, M. J. & El-Din, T. M. G. Structure and pharmacology of voltage-gated sodium and calcium channels. Annu. Rev. Pharmacol. Toxicol. 60, 133–154 (2020).
Diaz-Franulic, I., Poblete, H., Miño-Galaz, G., González, C. & Latorre, R. Allosterism and structure in thermally activated transient receptor potential channels. Annu. Rev. Biophys. 45, 371–398 (2016).
Zaczek, R. et al. Two new potent neurotransmitter release enhancers, 10, 10-bis(4-pyridinylmethyl)-9(10H)-anthracenone and 10, 10-bis(2-fluoro-4-pyridinylmethyl)-9 (10H)-anthracenone: comparison to linopirdine. J. Pharmacol. Exp. Ther. 285, 724–730 (1998).
Tatulian, L. & Brown, D. A. Effect of the KCNQ potassium channel opener retigabine on single KCNQ2/3 channels expressed in CHO cells. J. Physiol. 549, 57–63 (2003).
Taylor, K. C. et al. Structure and physiological function of the human KCNQ1 channel voltage sensor intermediate state. eLife 9, e53901 (2020).
Sun, J. & MacKinnon, R. Structural basis of human KCNQ1 modulation and gating. Cell 180, 340–347 (2020).
Jiang, Y. et al. X-ray structure of a voltage-dependent K+ channel. Nature 423, 33–41 (2003).
Swartz, K. J. Sensing voltage across lipid membranes. Nature 456, 891–897 (2008).
Catterall, W. A. Structure and function of voltage-gated ion channels. Annu. Rev. Biochem. 64, 493–531 (1995).
Hou, P. et al. Two-stage electro-mechanical coupling of a KV channel in voltage-dependent activation. Nat. Commun. 11, 676 (2020).
Kalstrup, T. & Blunck, R. S4-S5 linker movement during activation and inactivation in voltage-gated K+ channels. Proc. Natl Acad. Sci. USA 115, E6751–E6759 (2018).
Jensen, M. Ø. et al. Mechanism of voltage gating in potassium channels. Science 336, 229–233 (2012).
Xiong, Q., Sun, H. & Li, M. Zinc pyrithione-mediated activation of voltage-gated KCNQ potassium channels rescues epileptogenic mutants. Nat. Chem. Biol. 3, 287–296 (2007).
Rodriguez-Menchaca, A. A. et al. PIP2 controls voltage-sensor movement and pore opening of Kv channels through the S4-S5 linker. Proc. Natl Acad. Sci. USA 109, E2399–E2408 (2012).
Naranjo, D., Moldenhauer, H., Pincuntureo, M. & Díaz-Franulic, I. Pore size matters for potassium channel conductance. J. Gen. Physiol. 148, 277–291 (2016).
Lansky, S. et al. A pentameric TRPV3 channel with a dilated pore. Nature 621, 206–214 (2023).
Henderson, S. W., Nourmohammadi, S., Ramesh, S. A. & Yool, A. J. Aquaporin ion conductance properties defined by membrane environment, protein structure, and cell physiology. Biophys. Rev. 14, 181–198 (2022).
Perszyk, R. E. et al. The negative allosteric modulator EU1794-4 reduces single-channel conductance and Ca2+ permeability of GluN1/GluN2A N-methyl-d-aspartate receptors. Mol. Pharmacol. 99, 399–411 (2021).
Carnevale, V. & Klein, M. L. Small molecule modulation of voltage gated sodium channels. Curr. Opin. Struct. Biol. 43, 156–162 (2017).
Canul-Sánchez, J. A. et al. Different agonists induce distinct single-channel conductance states in TRPV1 channels. J. Gen. Physiol. 150, 1735–1746 (2018).
Sansom, M. S. Ion-channel gating. Twist to open. Curr. Biol. 5, 373–375 (1995).
Taly, A., Hénin, J., Changeux, J. P. & Cecchini, M. Allosteric regulation of pentameric ligand-gated ion channels: an emerging mechanistic perspective. Channels 8, 350–360 (2014).
Valadié, H., Lacapcre, J. J., Sanejouand, Y. H. & Etchebest, C. Dynamical properties of the MscL of Escherichia coli: a normal mode analysis. J. Mol. Biol. 332, 657–674 (2003).
Yang, H. Y. et al. Inherent dynamics of the acid-sensing ion channel 1 correlates with the gating mechanism. PLoS Biol. 7, e1000151 (2009).
Clarke, O. B. et al. Domain reorientation and rotation of an intracellular assembly regulate conduction in Kir potassium channels. Cell 141, 1018–1029 (2010).
Bavro, V. N. et al. Structure of a KirBac potassium channel with an open bundle crossing indicates a mechanism of channel gating. Nat. Struct. Mol. Biol. 19, 158–163 (2012).
Powers, A. S. et al. Structural basis of efficacy-driven ligand selectivity at GPCRs. Nat. Chem. Biol. 19, 805–814 (2023).
Hollingsworth, S. A. et al. Cryptic pocket formation underlies allosteric modulator selectivity at muscarinic GPCRs. Nat. Commun. 10, 3289 (2019).
Huang, X. S. et al. Structural basis for high-voltage activation and subtype-specific inhibition of human NaV1.8. Proc. Natl Acad. Sci. USA 119, e2208211119 (2022).
Liao, P. et al. Selective activation of TWIK-related acid-sensitive K+ 3 subunit–containing channels is analgesic in rodent models. Sci. Transl. Med. 11, eaaw8434 (2019).
Zhang, Q. et al. Inhibiting Hv1 channel in peripheral sensory neurons attenuates chronic inflammatory pain and opioid side effects. Cell Res. 32, 461–476 (2022).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. GCTF: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996).
Sehnal, D. et al. MOLE 2.0: advanced approach for analysis of biomacromolecular channels. J. Cheminform. 5, 39 (2013).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinformatics 54, 5.6.1–5.6.37 (2016).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Berendsen, H. J. C., van der Spoel, D. & van Drunen, R. GROMACS: a message-passing parallel molecular dynamics implementation. Comput. Phys. Commun. 91, 43–56 (1995).
Vanommeslaeghe, K. & MacKerell, A. D. Automation of the CHARMM general force field (CGenFF) I: bond perception and atom typing. J. Chem. Inf. Model. 52, 3144–3154 (2012).
Acknowledgements
This work is funded by grants from the National Key Research and Development Program of China (grant nos. 2022YFE0205600 to H.Y. and P.H.; 2018YFA0508100 to J.G. and Q.Z. and 2020YFA0908501 to J.G.), the National Natural Science Foundation of China (grant nos. 82273857 to Q.Z. and 32171221 to P.H.), the Joint Funding of the Macau Science and Technology Development Fund and the Ministry of Science and Technology of the People’s Republic of China (grant no. 0006/2021/AMJ to P.H.), the Innovative Research Team of High-level Local Universities in Shanghai (grant no. SHSMU-ZDCX20211201 to H.Y.) and the East China Normal University Medicine and Health Joint Fund (grant no. 2022JKXYD07001 to H.Y. and Z.C.). J.G. is supported by the MOE Frontier Science Center for Brain Science & Brain-Machine Integration, Zhejiang University. Single-particle cryo-EM data were collected at Center of Cryo-Electron Microscopy at Zhejiang University. We are also grateful for the support of the East China Normal University Multifunctional Platform for Innovation (001 and 011).
Author information
Authors and Affiliations
Contributions
H.Y., Q.Z. and J.G. conceived the project, designed the research and supervised the study. S.Z. performed virtual screening. K.W., Y.L. and J.H. performed whole-cell electrophysiology tests. K.W. and J.L. performed single-channel data acquisition. S.Z. and K.W. performed electrophysiological experiment analysis. D.M., Z.Y. and X.L. performed sample preparation, collected the cryo-EM structural data and solved the structures. S.Z., Q.Z. and J.S. performed and analyzed MD simulations. Q.Z., L.M. and Y.Y. assisted in compounds preparation and synthesis. Z.C. and P.H. provided intellectual expertise and shared key methodologies. S.Z., D.M. and Q.Z. prepared the draft of the manuscript. H.Y., Q.Z. and J.G. wrote the manuscript. All authors read and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
S.Z., K.W., Y.L., L.M., Y.Y., J.S., J.G., Q.Z. and H.Y. are inventors of patent applications that cover the potential usage of Ebio1 and its derivatives. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Huaizong Shen and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
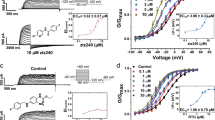
Extended Data Fig. 1 Virtual screening and biophysical characterization of small-molecules on KCNQ2 channel.
a, Chemical structures of initial 15 docking compounds, each with a different scaffold. b, Histogram showing the outward current potentiation effects of 15 candidates on the KCNQ2 channel. The effects of 10 μM candidates are checked at +50 mV. The dash line indicates a potentiation level of 1 (that is no effect). c, The effects of 10 μM candidates on the ΔV1/2 of KCNQ2 channel. NA means the results are not available. n = 3, 3, 2, 2, 2, 3, 3, 3, 3, 2, 3, 6, 3, 5 and 5 (b), n = 2, 2, 2, 4, 2, 2, 2, 3, 6, 2, 3 and 2 (c) biological replicates. Data are presented as the means ± s.e.m.
Extended Data Fig. 2 The selectivity of Ebio1 on several ion channels.
a, Summary of the effects of Ebio1 on TREK1, BK, hERG, CaV2.1, and NaV1.1 channels. Statistical analysis: one-way ANOVA with Dunnett’s test. b-f, The representative current traces of TREK1 (b), BK (c), hERG (d), CaV2.1 (e), and NaV1.1 (f) channels in the absence and the presence 10 μM Ebio1 are shown. n = 3, 3, 3, 2, 2 and 2 biological replicates (a). Data are presented as the means ± s.e.m.
Extended Data Fig. 3 Effects of Ebio1 on the activity of single KCNQ2/3, KCNQ4 and KCNQ5 channels.
a, Representative single-channel recordings from inside-out patches of KCNQ2/3, KCNQ4 and KCNQ5 at +50 mV in the absence and presence of 10 μM Ebio1 (left panel). The corresponding all-point amplitude histograms for the sweeps were fitted by Gaussian distributions (solid line in red) (right panel). b, Histogram showing the PO of KCNQ2/3, KCNQ4 and KCNQ5 at +50 mV in the absence and presence of 10 μM Ebio1. Statistical analysis: two-tailed t-test. n.s., not significant. c, Single-channel conductance of KCNQ2/3, KCNQ4 and KCNQ5 was fitted from two peak values in (a). Statistical analysis: two-tailed t-test. n.s., not significant. n = 4, 4, 3, 3, 3 and 3 (b), n = 4, 4, 3, 3, 3 and 3 (c) biological replicates. Data are presented as the means ± s.e.m.
Extended Data Fig. 4 Structure determination of human KCNQ2-Ebio1 complex.
a, Size-exclusion chromatography of KCNQ2-CaM on Superose 6 and SDS-PAGE analysis of the final sample. n = 3 independent experiments. b, Representative cryo-EM micrograph of KCNQ2-Ebio1. n = 3 independent experiments. c, Representative 2D classes. d, Flowchart of image processing for KCNQ2-Ebio1 particles. e, The density map of KCNQ2-Ebio1 is colored by local resolution. The local resolution is estimated with RELION 3.1 and generated in Chimera. f, The Gold-standard FSC curves of the final 3D reconstruction of KCNQ2-Ebio1, and the FSC curve for cross-validation between the map and the model of KCNQ2-Ebio1. g, EM densities for Ebio1 in the KCNQ2 channel. Gray meshes represent EM densities for Ebio1 and its surrounding residues. h, The weak and isolated densities are observed between VSD and PD in the map of KCNQ2-Ebio1 complex. The EM densities are contoured at the level of 0.01 in UCSF ChimeraX.
Extended Data Fig. 5 Structural changes induced by Ebio1.
a, The VSD structure of the Ebio1-bound KCNQ2. Only the S2-S4 helices are shown for clarity. The side chain of gating charges in S4 and the gating charge transfer center residue are shown in sticks and spheres. b, Comparison of Ebio1-bound VSD of KCNQ2 with open-state VSD of KCNQ2 (PDB code: 7CR0)24 and intermediate-state VSD of KCNQ1 (PDB code: 6MIE)33, which colored cyan, gray and purple, respectively. The side chain of gating charges in S4 and the gating charge transfer center residue are shown in sticks. c, Conformational change of the KCNQ2 channel complex in one KCNQ2-CaM subunit after Ebio1 bound. The conserved ‘EKR’ motif is colored red, which undergo structural rearrangement from a loop to a helix. S6 and HA helices of KCNQ2 are colored cyan. CaM is shown as helix and surface with its N-lobe in purple and C-lobe in pink. The rotational motion of CaM after Ebio1 bound is represented by a cartoon with a dash arrow.
Extended Data Fig. 6 MD simulations of the Ebio1- and RTG-bound KCNQ2 channel.
a, Simulation system II, RTG/KCNQ2 complex from the RTG-bound KCNQ2 cryo-EM structure. b, RMSD of RTG molecules against simulation time in the three independent repeats MD simulations of simulation system II. c, The number of hydrogen bonds formed between the -NH2 group of RTG and residue S303 (upper) or F305 (bottom) in the three independent repeats MD simulations of simulation system II. d, Representation of the channel pore diameter along the three independent repeats of RTG/KCNQ2 MD simulations of simulation system II. e, Simulation system III, Ebio1/KCNQ2 complex from the Ebio1-bound KCNQ2 cryo-EM structure. f, RMSD of Ebio1 molecules against simulation time in the three independent repeats MD simulations of simulation system III. g,h, Representation of the distance between residue S303 and F305 from adjacent subunit (g) or channel pore diameter (h) along the three independent repeats of Ebio1/KCNQ2 MD simulations of simulation system III.
Extended Data Fig. 7 The hydrogen bond restrained pore gate opening by affecting S6 helix motion.
a, Representative macroscopic current traces of KCNQ2S303A channel before (left traces) and after (right traces) application of 10 μM RTG or Ebio1. The holding potential was −80 mV. The KCNQ2S303A current was elicited by a series of voltage steps from −90 mV to +60 mV in 10 mV increments. b, Histogram showing the effects of 10 μM RTG or Ebio1 on the outward current amplitude of KCNQ2S303A at +50 mV. Statistical analysis: two-tailed t-test. c, Representative single-channel recordings from inside-out patches of KCNQ2S303A channel at +50 mV in the absence and presence of 10 μM RTG or Ebio1 (left panel). The corresponding all-point amplitude histograms for the sweeps were fitted by Gaussian distributions (solid line in red) (right panel). d, Histogram showing the PO of KCNQ2S303A at +50 mV in the absence and presence of 10 μM RTG or Ebio1. Statistical analysis: two-tailed t-test. n.s., not significant. e, Single-channel conductance of KCNQ2S303A was fitted from two peak values in (c). Statistical analysis: two-tailed t-test. n.s., not significant. n = 6, 7, 3 and 3 (b), n = 7, 7, 3 and 3 (d), n = 7, 7, 3 and 3 (e) biological replicates. Data are presented as the means ± s.e.m.
Extended Data Fig. 8 Small molecules RTG-S1 induce the dynamic rearrangement of the pore region of KCNQ2 channel.
a, RMSD of RTG-S1 molecules against simulation time in the three independent repeats MD simulations. b, Ensemble plot of the RTG-S1 molecule in the binding pocket of KCNQ2 channel during the MD simulations. c,d, Representation of the distance between residues S303 and F305 from the adjacent subunit (c) or channel pore diameter (d) along the three independent repeats of RTG-S1-dependent MD simulations. e, Typical KCNQ2 channel pore conformations of the two states (initial and final) from the RTG-S1-dependent trajectories. f, Superimposition of the gate region of the RTG-S1-dependent final state and KCNQ2-RTG (left) or KCNQ2-Ebio1 (right) cryo-EM structures.
Extended Data Fig. 9 Structure determination of human KCNQ2-Ebio1-S1 complex.
a, Size-exclusion chromatography of KCNQ2-CaM on Superose 6 and SDS-PAGE analysis of the final sample. n = 3 independent experiments. b, Representative cryo-EM micrograph of KCNQ2-Ebio1-S1. n = 3 independent experiments. c, Representative 2D classes. d, Flowchart of image processing for KCNQ2-Ebio1-S1 particles. e, The density map of KCNQ2-Ebio1-S1 is colored by local resolution. The local resolution is estimated with RELION 3.0 and generated in Chimera. f, EM densities for Ebio1-S1 in the KCNQ2 channel. Gray meshes represent EM densities for Ebio1-S1 and its surrounding residues. g, The Gold-standard FSC curves of the final 3D reconstruction of KCNQ2-Ebio1-S1, and the FSC curve for cross-validation between the map and the model of KCNQ2-Ebio1-S1. h, The weak and isolated densities are observed between VSD and PD in the map of KCNQ2-Ebio1-S1 complex. The EM densities are contoured at the level of 0.01 in UCSF ChimeraX.
Extended Data Fig. 10 Cryo-EM structures of human KCNQ2-Ebio1-S1 complex.
a, Structure model of the KCNQ2-Ebio1-S1 complex viewed parallel to the membrane (left) and top-down to the membrane (right) from the extracellular space. b, Comparison of Ebio1-S1-bound VSD of KCNQ2 with Ebio1-bound VSD of KCNQ2, which colored yellow and blue, respectively. The side chain of gating charges in S4 and the gating charge transfer center residue are shown in sticks.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2 and Figs. 1–12.
Supplementary Data 1
Statistical source data for Supplementary Figs. 1–3, 5, 8–10 and 12.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data
Source Data Extended .Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed gel.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 9
Unprocessed gel.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, S., Ma, D., Wang, K. et al. A small-molecule activation mechanism that directly opens the KCNQ2 channel. Nat Chem Biol (2024). https://doi.org/10.1038/s41589-023-01515-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41589-023-01515-y
This article is cited by
-
A new twist to increase ion flow
Nature Chemical Biology (2024)