Abstract
Metal-dependent formate dehydrogenases reduce CO2 with high efficiency and selectivity, but are usually very oxygen sensitive. An exception is Desulfovibrio vulgaris W/Sec-FdhAB, which can be handled aerobically, but the basis for this oxygen tolerance was unknown. Here we show that FdhAB activity is controlled by a redox switch based on an allosteric disulfide bond. When this bond is closed, the enzyme is in an oxygen-tolerant resting state presenting almost no catalytic activity and very low formate affinity. Opening this bond triggers large conformational changes that propagate to the active site, resulting in high activity and high formate affinity, but also higher oxygen sensitivity. We present the structure of activated FdhAB and show that activity loss is associated with partial loss of the metal sulfido ligand. The redox switch mechanism is reversible in vivo and prevents enzyme reduction by physiological formate levels, conferring a fitness advantage during O2 exposure.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data that support the findings of this study are available within the main text and its Supplementary Information file. The atomic coordinates and structure factors for the D. vulgaris H C872A variant structures have been deposited in the PDB under accession codes 8CM4, 8CM5, 8CM6 and 8CM7. Source data are provided with this paper.
References
Dalle, K. E. et al. Electro- and solar-driven fuel synthesis with first row transition metal complexes. Chem. Rev. 119, 2752–2875 (2019).
Wang, G. et al. Electrocatalysis for CO2 conversion: from fundamentals to value-added products. Chem. Soc. Rev. 50, 4993–5061 (2021).
Zhang, S., Fan, Q., Xia, R. & Meyer, T. J. CO2 reduction: from homogeneous to heterogeneous electrocatalysis. Acc. Chem. Res. 53, 2–11 (2019).
Shi, J. et al. Enzymatic conversion of carbon dioxide. Chem. Soc. Rev. 44, 5981–6000 (2015).
Yishai, O., Lindner, S. N., Gonzalez de la Cruz, J., Tenenboim, H. & Bar-Even, A. The formate bio-economy. Curr. Opin. Chem. Biol. 35, 1–9 (2016).
Mellmann, D., Sponholz, P., Junge, H. & Beller, M. Formic acid as a hydrogen storage material - development of homogeneous catalysts for selective hydrogen release. Chem. Soc. Rev. 45, 3954–3988 (2016).
Bulushev, D. & Ross, J. R. H. Towards sustainable production of formic acid from biomass for getting hydrogen and fuels. Chem. Sus. Chem. 11, 821–836 (2018).
Reda, T., Plugge, C. M., Abram, N. J. & Hirst, J. Reversible interconversion of carbon dioxide and formate by an electroactive enzyme. Proc. Natl Acad. Sci. USA 105, 10654–10658 (2008).
Stripp, S. T. et al. Second and outer coordination sphere effects in nitrogenase, hydrogenase, formate dehydrogenase, and CO dehydrogenase. Chem. Rev. 122, 11900–11973 (2022).
Schuchmann, K. & Müller, V. Autotrophy at the thermodynamic limit of life: a model for energy conservation in acetogenic bacteria. Nat. Rev. Microbiol. 12, 809–821 (2014).
Plugge, C. M., Zhang, W., Scholten, J. C. M. & Stams, A. J. M. Metabolic flexibility of sulfate-reducing bacteria. Front. Microbiol. 2, 81 (2011).
Sieber, J. R., Mcinerney, M. J. & Gunsalus, R. P. Genomic insights into syntrophy: the paradigm for anaerobic metabolic cooperation. Annu Rev. Microbiol. 66, 429–452 (2012).
Niks, D. & Hille, R. Molybdenum‐ and tungsten‐containing formate dehydrogenases and formylmethanofuran dehydrogenases: structure, mechanism, and cofactor insertion. Protein Sci. 28, 111–122 (2019).
Oliveira, A. R. et al. Toward the mechanistic understanding of enzymatic CO2 reduction. ACS Catal. 10, 3844–3856 (2020).
Dietrich, H. M. et al. Membrane-anchored HDCR nanowires drive hydrogen-powered CO2 fixation. Nature 607, 823–830 (2022).
Grimaldi, S., Schoepp-Cothenet, B., Ceccaldi, P., Guigliarelli, B. & Magalon, A. The prokaryotic Mo/W-bisPGD enzymes family: a catalytic workhorse in bioenergetic. Biochim. Biophys. Acta, Bioenerg. 1827, 1048–1085 (2013).
Hille, R. Molybdenum and tungsten in biology. Trends Biochem. Sci. 27, 360–367 (2002).
Schwarz, F. M., Schuchmann, K. & Müller, V. Hydrogenation of CO2 at ambient pressure catalyzed by a highly active thermostable biocatalyst. Biotechnol. Biofuels 11, 237 (2018).
da Silva, S. M., Pimentel, C., Valente, F. M. A. A., Rodrigues-Pousada, C. & Pereira, I. A. C. C. Tungsten and molybdenum regulation of formate dehydrogenase expression in Desulfovibrio vulgaris Hildenborough. J. Bacteriol. 193, 2909–2916 (2011).
da Silva, S. M. et al. Function of formate dehydrogenases in Desulfovibrio vulgaris Hildenborough energy metabolism. Microbiol. 159, 1760–1769 (2013).
Miller, M. et al. Interfacing formate dehydrogenase with metal oxides for the reversible electrocatalysis and solar‐driven reduction of carbon dioxide. Angew. Chem. Int. Ed. 58, 4601–4605 (2019).
Edwardes Moore, E. et al. A semi‐artificial photoelectrochemical tandem leaf witha CO2‐to‐formate efficiency approaching 1%. Angew. Chem. Int. Ed. 60, 26303–26307 (2021).
Szczesny, J. et al. Electroenzymatic CO2 fixation using redox polymer/enzyme-modified gas diffusion electrodes. ACS Energy Lett. 5, 321–327 (2020).
Alvarez-Malmagro, J. et al. Bioelectrocatalytic activity of W-formate dehydrogenase covalently immobilized on functionalized gold and graphite electrodes. ACS Appl. Mater. Interfaces 13, 11891–11900 (2021).
Antón-García, D. et al. Photoelectrochemical hybrid cell for unbiased CO2 reduction coupled to alcohol oxidation. Nat. Synth. 1, 77–86 (2022).
De Bok, F. A. M. M. et al. Two W-containing formate dehydrogenases (CO2-reductases) involved in syntrophic propionate oxidation by Syntrophobacter fumaroxidans. Eur. J. Biochem. 270, 2476–2485 (2003).
Schuchmann, K. & Müller, V. Direct and reversible hydrogenation of CO2 to formate by a bacterial carbon dioxide reductase. Science 342, 1382–1385 (2013).
Raaijmakers, H. et al. Gene sequence and the 1.8 A crystal structure gene sequence and the 1.8 A of the tungsten-containing formate dehydrogenase from Desulfovibrio gigas. Structure 10, 1261–1272 (2002).
Oliveira, A. R. et al. Spectroscopic and structural characterization of reduced Desulfovibrio vulgaris Hildenborough W-FdhAB reveals stable metal coordination during catalysis. ACS Chem. Biol. 17, 1901–1909 (2022).
Vilela-Alves, G. et al. Tracking W-formate dehydrogenase structural changes during catalysis and enzyme reoxidation. Int. J. Mol. Sci. 24, 476 (2023).
Grimaldi, S., Biaso, F., Burlat, B. & Guigliarelli, B. in Molybdenum and Tungsten Enzymes: Spectroscopic and Theoretical Investigations (eds Hille, R. et al.) 68–120 (The Royal Society of Chemistry, 2016).
Al-Attar, S. et al. Gating of substrate access and long-range proton transfer in Escherichia coli nitrate reductase A: the essential role of a remote glutamate residue. ACS Catal. 11, 14303–14318 (2021).
Iwaoka, M. & Isozumi, N. Hypervalent nonbonded interactions of a divalent sulfur atom. Implications in protein architecture and the functions. Molecules 2, 7266–7283 (2012).
Léger, C. & Bertrand, P. Direct electrochemistry of redox enzymes as a tool for mechanistic studies. Chem. Rev. 108, 2379–2438 (2008).
Chiu, J. & Hogg, X. P. J. Allosteric disulfides: sophisticated molecular structures enabling flexible protein regulation. J. Biol. Chem. 294, 2949–5908 (2019).
Hartmann, T. & Leimkühler, S. The oxygen-tolerant and NAD+-dependent formate dehydrogenase from Rhodobacter capsulatus is able to catalyze the reduction of CO2 to formate. FEBS J. 280, 6083–6096 (2013).
Yu, X., Niks, D., Mulchandani, A. & Hille, R. Efficient reduction of CO2 by the molybdenum-containing formate dehydrogenase from Cupriavidus necator (Ralstonia eutropha). J. Biol. Chem. 292, 16872–16879 (2017).
Graham, J. E. et al. How a formate dehydrogenase responds to oxygen: unexpected O2 insensitivity of an enzyme harboring tungstate, selenocysteine, and [4Fe-4S] clusters. ACS Catal. 12, 10449–10471 (2022).
Hogg, P. J. Disulfide bonds as switches for protein function. Trends Biochem. Sci. 28, 210–214 (2003).
Wensien, M. et al. A lysine-cysteine redox switch with an NOS bridge regulates enzyme function. Nature 593, 460–464 (2021).
Rabe von Pappenheim, F. et al. Widespread occurrence of covalent lysine–cysteine redox switches in proteins. Nat. Chem. Biol. 18, 368–375 (2022).
Schrapers, P. et al. Sulfido and cysteine ligation changes at the molybdenum cofactor during substrate conversion by formate dehydrogenase (FDH) from Rhodobacter capsulatus. Inorg. Chem. 54, 3260–3271 (2015).
Hakopian, S., Niks, D. & Hille, R. The air-inactivation of formate dehydrogenase FdsDABG from Cupriavidus necator. J. Inorg. Biochem. 231, 111788 (2022).
Agne, M., Appel, L., Seelmann, C. & Boll, M. Enoyl-coenzyme A respiration via formate cycling in syntrophic bacteria. mBio 13, 1–16 (2022).
Schink, B., Montag, D., Keller, A. & Müller, N. Hydrogen or formate: alternative key players in methanogenic degradation. Environ. Microbiol Rep. 9, 189–202 (2017).
Dong, G. & Ryde, U. Reaction mechanism of formate dehydrogenase studied by computational methods. J. Biol. Inorg. Chem. 1, 1243–1254 (2018).
Cypionka, H. Oxygen respiration by Desulfovibrio species. Annu. Rev. Microbiol. 54, 827–848 (2000).
Dolla, A., Fournier, M. & Dermoun, Z. Oxygen defense in sulfate-reducing bacteria. J. Biotechnol. 126, 87–100 (2006).
Mukhopadhyay, A. et al. Cell-wide responses to low-oxygen exposure in Desulfovibrio vulgaris Hildenborough. J. Bacteriol. 189, 5996–6010 (2007).
Fareleira, P. et al. Response of a strict anaerobe to oxygen: survival strategies in Desulfovibrio gigas. Microbiol. 149, 1513–1522 (2003).
Imlay, J. A., Sethu, R. & Rohaun, S. K. Evolutionary adaptations that enable enzymes to tolerate oxidative stress. Free Radic. Biol. Med. 140, 4–13 (2019).
Lu, Z. & Imlay, J. A. When anaerobes encounter oxygen: mechanisms of oxygen toxicity, tolerance and defence. Nat. Rev. Microbiol. 19, 774–785 (2021).
Baumgarten, A., Redenius, I., Kranczoch, J. & Cypionka, H. Periplasmic oxygen reduction by Desulfovibrio species. Arch. Microbiol. 176, 306–309 (2001).
Ramel, F. et al. Membrane-bound oxygen reductases of the anaerobic sulfate-reducing Desulfovibrio vulgaris Hildenborough: roles in oxygen defence and electron link with periplasmic hydrogen oxidation. Microbiol. Soc. 159, 2663–2673 (2013).
Fournier, M. et al. Response of the anaerobe Desulfovibrio vulgaris Hildenborough to oxidative conditions: proteome and transcript analysis. Biochimie 88, 85–94 (2006).
Vita, N., Hatchikian, E. C., Nouailler, M., Dolla, A. & Pieulle, L. Disulfide bond-dependent mechanism of protection against oxidative stress in pyruvate-ferredoxin oxidoreductase of anaerobic Desulfovibrio bacteria. Biochemistry 47, 957–964 (2008).
Rodríguez-Maciá, P. et al. Sulfide protects [FeFe] hydrogenases from O2. J. Am. Chem. Soc. 140, 9346–9350 (2018).
Marques, M. C., Coelho, R., De Lacey, A. L., Pereira, I. A. C. & Matias, P. M. The three-dimensional structure of [nifese] hydrogenase from Desulfovibrio vulgaris Hildenborough: a hydrogenase without a bridging ligand in the active site in its oxidised, ‘as-isolated’ state. J. Mol. Biol. 396, 893–907 (2010).
Felbek, C. et al. Mechanism of hydrogen sulfide-dependent inhibition of FeFe hydrogenase. ACS Catal. 11, 15162–15176 (2021).
Wittenborn, E. C. et al. The solvent-exposed Fe–S D-cluster contributes to oxygen-resistance in Desulfovibrio vulgaris Ni–Fe carbon monoxide dehydrogenase. ACS Catal. 10, 7328–7335 (2020).
Keller, K. L., Wall, J. D. & Chhabra, S. in Synthetic Biology, Part A Vol. 497 (ed. Voigt, C.) 503–517 (Elsevier Inc., 2011).
Meneghello, M. et al. Formate dehydrogenases reduce CO2 rather than HCO3−: an electrochemical demonstration. Angew. Chem. Int. Ed. 60, 9964–9967 (2021).
Fourmond, V. QSoas: a versatile software for data analysis. Anal. Chem. 88, 5050–5052 (2016).
Juanhuix, J. et al. Developments in optics and performance at BL13-XALOC, the macromolecular crystallography beamline at the ALBA Synchrotron. J. Synchrotron Radiat. 21, 679–689 (2014).
McCarthy, A. A. et al. ID30B—a versatile beamline for macromolecular crystallography experiments at the ESRF. J. Synchrotron Radiat. 25, 1249–1260 (2018).
Nurizzo, D. et al. The ID23-1 structural biology beamline at the ESRF. J. Synchrotron Radiat. 13, 227–238 (2006).
Von Stetten, D. et al. ID30A-3 (MASSIF-3)—a beamline for macromolecular crystallography at the ESRF with a small intense beam. J. Synchrotron Radiat. 27, 844–851 (2020).
Kabsch, W. XDS. Acta Crystallogr. D. Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D. Biol. Crystallogr. 69, 1204–1214 (2013).
Vonrhein, C. et al. Data processing and analysis with the autoPROC toolbox. Acta Crystallogr. D. Biol. Crystallogr. 67, 293–302 (2011).
Vonrhein, C. et al. Advances in automated data analysis and processing within autoPROC, combined with improved characterisation, mitigation and visualisation of the anisotropy of diffraction limits using STARANISO. Acta Cryst. A Found. Adv. 74, 43537 (2018).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D. Biol. Crystallogr. 67, 235–242 (2011).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D. Biol. Crystallogr. 67, 355–367 (2011).
Sievers, F. & Higgins, D. G. Clustal Omega for making accurate alignments of many protein sequences. Protein Sci. 27, 135–145 (2018).
Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview version 2-A multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Savojardo, C., Martelli, P. L., Fariselli, P. & Casadio, R. DeepSig: deep learning improves signal peptide detection in proteins. Bioinformatics 34, 1690–1696 (2018).
Acknowledgements
This work was financially supported by Fundação para a Ciência e Tecnologia (FCT, Portugal) through fellowship nos. SFRH/BD/116515/2016 (A.R.O.), DFA/BD/7897/2020 (R.R.M.) and COVID/BD/151766/2021 (A.R.O.), grant no. PTDC/BII-BBF/2050/2020 (I.A.C.P. and M.J.R.) and R&D units MOSTMICRO-ITQB (grant nos. UIDB/04612/2020 and UIDP/04612/2020) (I.A.C.P.) and UCIBIO (grant nos. UIDP/04378/2020 and UIDB/04378/2020) (M.J.R.), and Associated Laboratories LS4FUTURE (grant no. LA/P/0087/2020) (I.A.C.P.) and i4HB (grant no. LA/P/0140/2020) (M.J.R.). The European Union’s Horizon 2020 research and innovation program (grant no. 810856) is also acknowledged (I.A.C.P.). This work was also funded by the French national research agency (ANR – MOLYERE project, grant no. 16-CE-29-0010-01) (B.G.), and supported by the computing facilities of the Centre Régional de Compétences en Modélisation Moléculaire de Marseille. We thank the excellent technical assistance of João Carita from ITQB NOVA on microbial cell growth. We are also grateful to the EPR-MRS facilities of the Aix-Marseille University EPR centre and acknowledge the support of the European research infrastructure MOSBRI (grant no. 101004806) (B.G.) and the French research infrastructure INFRANALYTICS (FR2054) (B.G.). We also acknowledge the ESRF and ALBA Synchrotron for provision of synchrotron radiation facilities, and we thank the staff of the ESRF and EMBL Grenoble and ALBA for assistance and support in using beamlines ID23-1, ID30A-3, ID30B and XALOC.
Author information
Authors and Affiliations
Contributions
I.A.C.P. and A.R.O. conceived and designed biochemical experiments and sequence analysis. A.R.O. performed molecular biology experiments, protein purification, biochemical characterization, enzymatic assays, thermal shift assays, sequence analysis, EPR and electrochemical studies of WT, C845A and C872A variants and figure preparation. R.R.M. produced and characterized the M405A variant and contributed to CO2 reduction assays and figure preparation. C.M., M.J.R. and G.V.-A. designed the crystallography experiments and analyzed the crystal structures. C.M. and G.V.-A. crystallized the proteins, solved and refined all structures. G.V.-A. prepared all figures with crystal structures. K.K. helped with crystallization assays and first stages of refinement of the 8CM4 and 8CM5 structures. V.F. and C.L. designed electrochemical experiments, performed by V.F. and A.R.O. B.G. designed and analyzed EPR experiments, performed by B.G. and A.R.O. I.A.C.P., R.R.M. and N.P. designed in vivo experiments, performed by N.P. and R.R.M. I.A.C.P. and M.J.R. supervised and funded the project. A.R.O., I.A.C.P., M.J.R., C.M. and G.V.-A. wrote the manuscript with inputs from coauthors V.F., C.L. and B.G. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Russ Hille and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
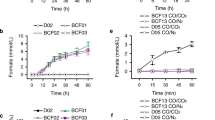
Extended Data Fig. 1 Steady-state formate oxidation (A, C, and E) and CO2 reduction kinetics (B, D and F).
Formate oxidation (A, C, and E) and CO2 reduction (B, D and F). A and B – As-isolated WT FdhAB assays without DTT and with DTT activation, respectively. C and D – C872A variant anaerobically isolated (no DTT activation). E and F – M405A variant anaerobically isolated (no DTT activation). The concentration of enzyme used in the assays was 14 nM for A, E and F, and 0.9-2 nM for B, C and D. Data are presented as mean values ± s.d. (n = 2 or 3 assay technical replicates). Lines represent the direct fitting of Michaelis-Menten equation to experimental data.
Extended Data Fig. 2 EPR spectra of WV species in WT FdhAB enzyme and in C872A and M405A variants.
Experimental spectra are in black, and simulations are shown below in red. a) resting WT sample poised at −468 mV by reduction with dithionite; b) DTT activated WT sample poised at −395 mV by reduction with formate; c) C872A sample poised at −443 mV by reduction with formate; d) C872A sample poised at −469 mV by reduction with dithionite; e) M405A reduced by dithionite. EPR conditions: temperature 80 K; microwave power 40 mW at 9.479 GHz; modulation amplitude 1 mT at 100 kHz. Simulated spectra result from the superimposition of WVF and WVD signals with parameters given in Table S2 with the following WVD /WVF ratio: a) 90/10; b) 5/95; c) 10/90; d) 80/20. For the sake of clarity, the radical signal at g=2.00 arising from mediators in redox titrations was not simulated.
Extended Data Fig. 3 FdhAB W active site.
FdhAB W active site in aerobic C872A_ox crystal structure (a, b, c) and C872A_anox anaerobic one (d). The results for three different labile group refinement strategies of the labile ligand, for the C872A-O2 model, are presented: (a) refinement with an oxygen replacing the sulfido ligand; (b) refinement with a sulfido ligand with an occupancy of 1; and (c) refinement with a sulfido ligand refined with an occupancy of 0.556. In (d) the W center for the C872Aanoxic-CHNH2O is shown. The pterin domains of both MGD cofactors and the sidechain of U192 are represented as light blue or blue sticks, respectively, for C872A-O2 and C872Aanoxic-CHNH2O, the W ion as a blue sphere and the sulfido ligand by a yellow stick. The 2Fo-Fc electron density map for each refinement is shown as a magenta mesh, at 1σ, and the Fo-Fc map is represented as a green mesh, for positive density, and as a red mesh, for negative density, both at 3σ.
Extended Data Fig. 4 Comparison of the active site in WT and C872A_anox structures.
Superposition of FdhAB WT (gray) and C872A_anox structures (blue) showing the new H193 conformation and respective hydrogen bonding network. The W ion is represented as a violet sphere and its two bound MGDs are shown as sticks. Distances are in Å.
Extended Data Fig. 5 A formamide molecule is present near the active site (C872A_anox structure) in the formate channel.
R441 and T450 are hydrogen bonding the formamide ligand. The W ion is represented as a violet sphere and its two bound MGDs are shown as sticks. The 2Fo-Fc electron density map is shown as a magenta mesh, at 1σ. Distances are in Å.
Extended Data Fig. 6 EPR spectrum of WV in reduced M405A variant.
a) M405A sample as-isolated anaerobically; b) after reduction with dithionite; c) after reduction with formate; d) formate-reduced sample after washing-off formate. EPR conditions: temperature 80 K; microwave power 40 mW; modulation amplitude 1 mT at 100 kHz.
Extended Data Fig. 7 Active site environment of the M405A variant.
Superposition of WT (grey) and M405A variant (yellow) is shown. Black arrows highlight the conformational changes. The W ion is represented as a violet sphere and its two bound MGDs are shown as sticks.
Extended Data Fig. 8 Rate of O2 inactivation.
Solid lines show the derivative of the logarithm of the current for a– DTT-activated WT FdhAB, b– Anaerobically purified WT FdhAB, c- C845A and d- C872A variants, after injection of 30 µM of O2. Dashed lines are the representative fits of the kinetic model in equation [3] (Supplementary Information).
Supplementary information
Supplementary Information
Supplementary Data 1–3, Figs. 1–3, Tables 1–7 and references.
Supplementary Data 1
Source Data for Supplementary Fig. 2.
Supplementary Data 2
Source Data for Supplementary Fig. 3.
Source data
Source Data Fig. 1
Statistical source data for Fig. 1.
Source Data Fig. 2
Source data for Fig. 2.
Source Data Fig. 4
Statistical source data for Fig. 4.
Source Data Fig. 5
Statistical source data for Fig. 5.
Source Data Table 1
Statistical source data for Table 1.
Source Data Extended Data Fig. 1
Statistical source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 6
Source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 8
Source data for Extended Data Fig. 8.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Oliveira, A.R., Mota, C., Vilela-Alves, G. et al. An allosteric redox switch involved in oxygen protection in a CO2 reductase. Nat Chem Biol 20, 111–119 (2024). https://doi.org/10.1038/s41589-023-01484-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-023-01484-2