Abstract
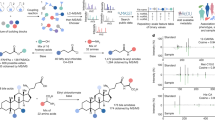
The microbiota generates diverse metabolites to modulate host physiology and disease, but their protein targets and mechanisms of action have not been fully elucidated. To address this challenge, we explored microbiota-derived indole metabolites and developed photoaffinity chemical reporters for proteomic studies. We identified many potential indole metabolite-interacting proteins, including metabolic enzymes, transporters, immune sensors and G protein-coupled receptors. Notably, we discovered that aromatic monoamines can bind the orphan receptor GPRC5A and stimulate β-arrestin recruitment. Metabolomic and functional profiling also revealed specific amino acid decarboxylase-expressing microbiota species that produce aromatic monoamine agonists for GPRC5A-β-arrestin recruitment. Our analysis of synthetic aromatic monoamine derivatives identified 7-fluorotryptamine as a more potent agonist of GPRC5A. These results highlight the utility of chemoproteomics to identify microbiota metabolite-interacting proteins and the development of small-molecule agonists for orphan receptors.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD040163. The metabolomics data have been searched in METLIN database and deposited to the XCMS Public with the dataset identifier 1529105. The RNA-seq data have been deposited to Gene Expression Omnibus with the accession number GSE225392. Human Microbiome Project Reference Genomes were downloaded via BioProject Accession number PRJNA28331. All other data supporting the findings of this study are available within the article and its Supplementary Information files, and from the corresponding author upon request. Source data are provided with this paper.
References
Donia, M. S. & Fischbach, M. A. HUMAN MICROBIOTA. Small molecules from the human microbiota. Science 349, 1254766 (2015).
Gill, S. K., Rossi, M., Bajka, B. & Whelan, K. Dietary fibre in gastrointestinal health and disease. Nat. Rev. Gastroenterol. Hepatol. 18, 101–116 (2021).
Liu, Y., Hou, Y., Wang, G., Zheng, X. & Hao, H. Gut microbial metabolites of aromatic amino acids as signals in host-microbe interplay. Trends Endocrinol. Metab. 31, 818–834 (2020).
Collins, S. L., Stine, J. G., Bisanz, J. E., Okafor, C. D. & Patterson, A. D. Bile acids and the gut microbiota: metabolic interactions and impacts on disease. Nat. Rev. Microbiol. 21, 236–247 (2023).
Zimmermann, M., Zimmermann-Kogadeeva, M., Wegmann, R. & Goodman, A. L. Mapping human microbiome drug metabolism by gut bacteria and their genes. Nature 570, 462–467 (2019).
Lam, K. C. et al. Microbiota triggers STING-type I IFN-dependent monocyte reprogramming of the tumor microenvironment. Cell 184, 5338–5356 (2021).
Rangan, K. J. et al. A secreted bacterial peptidoglycan hydrolase enhances tolerance to enteric pathogens. Science 353, 1434–1437 (2016).
Griffin, M. E. et al. Enterococcus peptidoglycan remodeling promotes checkpoint inhibitor cancer immunotherapy. Science 373, 1040–1046 (2021).
Dodd, D. et al. A gut bacterial pathway metabolizes aromatic amino acids into nine circulating metabolites. Nature 551, 648–652 (2017).
Dong, F. et al. Intestinal microbiota-derived tryptophan metabolites are predictive of Ah receptor activity. Gut Microbes 12, 1–24 (2020).
Scott, S. A., Fu, J. & Chang, P. V. Microbial tryptophan metabolites regulate gut barrier function via the aryl hydrocarbon receptor. Proc. Natl Acad. Sci. USA 117, 19376–19387 (2020).
Hezaveh, K. et al. Tryptophan-derived microbial metabolites activate the aryl hydrocarbon receptor in tumor-associated macrophages to suppress anti-tumor immunity. Immunity 55, 324–340 (2022).
Tintelnot, J. et al. Microbiota-derived 3-IAA influences chemotherapy efficacy in pancreatic cancer. Nature 615, 168–174 (2023).
Bhattarai, Y. et al. Gut microbiota-produced tryptamine activates an epithelial G-protein-coupled receptor to increase colonic secretion. Cell Host Microbe 23, 775–785 (2018).
Brubaker, S. W., Bonham, K. S., Zanoni, I. & Kagan, J. C. Innate immune pattern recognition: a cell biological perspective. Annu. Rev. Immunol. 33, 257–290 (2015).
Jia, W., Xie, G. & Jia, W. Bile acid-microbiota crosstalk in gastrointestinal inflammation and carcinogenesis. Nat. Rev. Gastroenterol. Hepatol. 15, 111–128 (2018).
Husted, A. S., Trauelsen, M., Rudenko, O., Hjorth, S. A. & Schwartz, T. W. GPCR-mediated signaling of metabolites. Cell Metab. 25, 777–796 (2017).
Zhao, X., Yang, X. & Hang, H. C. Chemoproteomic analysis of microbiota metabolite-protein targets and mechanisms. Biochemistry 61, 2822–2834 (2022).
Cheng, Y. & Lotan, R. Molecular cloning and characterization of a novel retinoic acid-inducible gene that encodes a putative G protein-coupled receptor. J. Biol. Chem. 273, 35008–35015 (1998).
Zhou, H. & Rigoutsos, I. The emerging roles of GPRC5A in diseases. Oncoscience 1, 765–776 (2014).
Lamas, B. et al. CARD9 impacts colitis by altering gut microbiota metabolism of tryptophan into aryl hydrocarbon receptor ligands. Nat. Med. 22, 598–605 (2016).
Chin, E. N. et al. Antitumor activity of a systemic STING-activating non-nucleotide cGAMP mimetic. Science 369, 993–999 (2020).
Pan, B. S. et al. An orally available non-nucleotide STING agonist with antitumor activity. Science 369, eaba6098 (2020).
Kroeze, W. K. et al. PRESTO-Tango as an open-source resource for interrogation of the druggable human GPCRome. Nat. Struct. Mol. Biol. 22, 362–369 (2015).
Williams, B. B. et al. Discovery and characterization of gut microbiota decarboxylases that can produce the neurotransmitter tryptamine. Cell Host Microbe 16, 495–503 (2014).
Pessione, E. et al. First evidence of a membrane-bound, tyramine and beta-phenylethylamine producing, tyrosine decarboxylase in Enterococcus faecalis: a two-dimensional electrophoresis proteomic study. Proteomics 9, 2695–2710 (2009).
Chen, H. et al. A forward chemical genetic screen reveals gut microbiota metabolites that modulate host physiology. Cell 177, 1217–1231 (2019).
Adibi, S. A. & Mercer, D. W. Protein digestion in human intestine as reflected in luminal, mucosal, and plasma amino acid concentrations after meals. J. Clin. Investig. 52, 1586–1594 (1973).
Chen, V., Griffin, M. E., Maguin, P., Varble, A. & Hang, H. C. RecT recombinase expression enables efficient gene editing in Enterococcus spp. Appl Environ. Microbiol 87, e0084421 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Chun, L., Zhang, W. H. & Liu, J. F. Structure and ligand recognition of class C GPCRs. Acta Pharmacol. Sin. 33, 312–323 (2012).
Zhong, S. et al. Lung tumor suppressor GPRC5A binds EGFR and restrains its effector signaling. Cancer Res. 75, 1801–1814 (2015).
Deng, J. et al. Knockout of the tumor suppressor gene Gprc5a in mice leads to NF-kappaB activation in airway epithelium and promotes lung inflammation and tumorigenesis. Cancer Prev. Res. 3, 424–437 (2010).
Popivanova, B. K. et al. Blocking TNF-alpha in mice reduces colorectal carcinogenesis associated with chronic colitis. J. Clin. Investig. 118, 560–570 (2008).
Cheng, Z. et al. Luciferase reporter assay system for deciphering GPCR pathways. Curr. Chem. Genomics 4, 84–91 (2010).
Olsen, R. H. J. et al. TRUPATH, an open-source biosensor platform for interrogating the GPCR transducerome. Nat. Chem. Biol. 16, 841–849 (2020).
Witherow, D. S., Garrison, T. R., Miller, W. E. & Lefkowitz, R. J. beta-Arrestin inhibits NF-kappaB activity by means of its interaction with the NF-kappaB inhibitor IkappaBalpha. Proc. Natl Acad. Sci. USA 101, 8603–8607 (2004).
Gao, H. et al. Identification of beta-arrestin2 as a G protein-coupled receptor-stimulated regulator of NF-kappaB pathways. Mol. Cell 14, 303–317 (2004).
Hirabayashi, Y. & Kim, Y. J. Roles of GPRC5 family proteins: focusing on GPRC5B and lipid-mediated signalling. J. Biochem. 167, 541–547 (2020).
Laschet, C., Dupuis, N. & Hanson, J. The G protein-coupled receptors deorphanization landscape. Biochem. Pharmacol. 153, 62–74 (2018).
Maini Rekdal, V., Bess, E. N., Bisanz, J. E., Turnbaugh, P. J. & Balskus, E. P. Discovery and inhibition of an interspecies gut bacterial pathway for Levodopa metabolism. Science 364, eaau6323 (2019).
Zhai, L. et al. Ruminococcus gnavus plays a pathogenic role in diarrhea-predominant irritable bowel syndrome by increasing serotonin biosynthesis. Cell Host Microbe 31, 33–44.e5 (2022).
Zhou, Y. et al. Increased Enterococcus faecalis infection is associated with clinically active Crohn disease. Medicine 95, e5019 (2016).
Seishima, J. et al. Gut-derived Enterococcus faecium from ulcerative colitis patients promotes colitis in a genetically susceptible mouse host. Genome Biol. 20, 252 (2019).
Cao, Y. et al. Commensal microbiota from patients with inflammatory bowel disease produce genotoxic metabolites. Science 378, eabm3233 (2022).
Kadara, H. et al. A Gprc5a tumor suppressor loss of expression signature is conserved, prevalent, and associated with survival in human lung adenocarcinomas. Neoplasia 12, 499–505 (2010).
Tao, Q. et al. Identification of the retinoic acid-inducible Gprc5a as a new lung tumor suppressor gene. J. Natl Cancer Inst. 99, 1668–1682 (2007).
Insel, P. A. et al. GPCRomics: GPCR expression in cancer cells and tumors identifies new, potential biomarkers and therapeutic targets. Front. Pharmacol. 9, 431 (2018).
Greenhough, A. et al. Cancer cell adaptation to hypoxia involves a HIF-GPRC5A-YAP axis. EMBO Mol. Med. 10, e8699 (2018).
Song, H. et al. NF-κB represses retinoic acid receptor-mediated GPRC5A transactivation in lung epithelial cells to promote neoplasia. JCI Insight 8, e153976 (2023).
Barnea, G. et al. The genetic design of signaling cascades to record receptor activation. Proc. Natl Acad. Sci. USA 105, 64–69 (2008).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 11, 2301–2319 (2016).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Ge, S. X., Jung, D. & Yao, R. ShinyGO: a graphical gene-set enrichment tool for animals and plants. Bioinformatics, 2628–2629 (2020).
Ribeiro, F. J. et al. Finished bacterial genomes from shotgun sequence data. Genome Res. 22, 2270–2277 (2012).
Hullahalli, K., Rodrigues, M., Nguyen, U. T. & Palmer, K. An attenuated CRISPR–Cas system in Enterococcus faecalis permits DNA acquisition. mBio 9, e00414–e00418 (2018).
Tautenhahn, R., Patti, G. J., Rinehart, D. & Siuzdak, G. XCMS online: a web-based platform to process untargeted metabolomic data. Anal. Chem. 84, 5035–5039 (2012).
Guijas, C. et al. METLIN: a technology platform for identifying knowns and unknowns. Anal. Chem. 90, 3156–3164 (2018).
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Acknowledgements
We thank H. Molina from Proteomics Resource Center at Rockefeller University for proteomics and HRMS, G. Siuzdak and L.T. Hoang from the Scripps Center for Metabolomics and Mass Spectrometry for untargeted metabolomics, S.R. Head and J. Shimashita from Genomics Core at Scripps Research for RNA-seq, P. Natarajan and A. Sundaresan from Center for Computational Biology and Bioinformatics and W. Li at Scripps Research for RNA-seq data analysis. We also thank G. Barnea (Brown University) for providing HTLA cells, B. Roth (University of North Carolina) for PRESTO-Tango plasmids, S. Kitamura and F. Martinez (Scripps Research) for sharing aromatic monoamines, M. Gilmore (Harvard Medical School) for E. faecium Com15, S. F. Brady (The Rockefeller University) for R. gnavus ATCC 29149, and C.-J. Guo (Weill Cornell Medical School) for C. sporogenes ATCC 15579. K.S. acknowledges support from Scripps Research Department of Immunology and Microbiology T32AI007244 grant. L.L.L acknowledges grant support from NIH-NINDS R01NS112482-02. H.C.H. acknowledges grant support from NIH-NIGMS R01GM087544.
Author information
Authors and Affiliations
Contributions
X.Z. and H.C.H. conceived the project. K.R.S. contributed to the generation of GPRC5A-KO cell lines and flow cytometry analysis of GPRC5A mutants, V.C. prepared TyrDC-KO E. faecium Com15, M.E.G. performed phylogenetic analysis of bacterial decarboxylases and L.L.L. provided the aromatic monoamine derivatives. X.Z. performed the experiments and interpreted the data. X.Z. and H.C.H. wrote the manuscript with input from K.R.S., V.C., M.E.G. and L.L.L.
Corresponding author
Ethics declarations
Competing interests
All authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Chemoproteomic analysis of indole-3-acetic acid-interacting proteins.
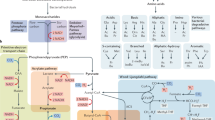
a, Scheme for in-gel fluorescence profiling and chemoproteomic analysis of microbiota metabolite protein targets in HT-29 cells using photoaffinity chemical reporters. b, In-gel fluorescence profiling of IAA-interacting proteins in HT-29 cells using indicated concentration of x-alk-IAA with or without UV irradiation. At least two independent experiments were performed. c, Volcano plot of the identified proteins from HT-29 cells by chemical proteomics using x-alk-IAA with UV irradiation compared to DMSO control (p value is 0.05, two folds change in four replicates from two sample t-test). Some of key hits are highlighted in GPCRs (green), transporters (blue) and immunity-associated proteins (red). d, Venn diagram for the statistically significant hits identified by chemical proteomics using two chemical reporters with or without UV irradiation. e, Gene ontology analysis of the major subcellular localization, Fisher’s exact test. f, KEGG pathway analysis of the statistically significant proteins identified in HT-29 cells using x-alk-IAA with UV irradiation from the hypergeometric test.
Extended Data Fig. 2 Chemoproteomic analysis of tryptamine-interacting proteins.
a, PRESTO-Tango assay for screening a collection of indole derivatives (100 µM) on ADRA2A, GPRC5A, GPR107 and GPR108. 0.1% DMSO were used as controls. b-c, ADRA2A and GPRC5A are crosslinked by IAA photoaffinity chemical reporters. Competitive labeling assay using 200 µM IAA or TA that compete with 20 µM x-alk-IAA in binding on ADRA2A or GPRC5A. d, In-gel fluorescence profiling of IAA- or TA-interacting proteins in A549 cells using indicated concentration of x-alk-IAA or x-alk-TA with or without UV irradiation. At least two independent experiments were performed. e, Volcano plot of the identified proteins from A549 cells by chemical proteomics using x-alk-IAA or x-alk-TA with UV irradiation (p value is 0.05, two folds change in four replicates). f, Pull-down of endogenous GPRC5A from A549 cells that were photo-crosslinked by x-alk-IAA or x-alk-TA followed by biotin labeling and neutravidin enrichment. At least two independent experiments were performed. Data indicates mean with SEM from three replicates (a,c). Two-way ANOVA using Holm-Sidak’s multiple comparisons test (a, c), ****P < 0.0001.
Extended Data Fig. 3 Targeted metabolomics of microbiota species.
a, LC-MS analysis of the concentration of aromatic monoamines in different bacterial cultures. Data indicates individual value of three replicates. b, Western blot analysis of the overexpression of His6-tagged E. faecium Com15 tyrosine decarboxylase (Efm_TyrDC), R. gnavus tryptophan decarboxylase (Rgs_TrpDC) or M. morganii glutamate/tyrosine decarboxylase (Mmi_Gln/TyrDC) in E. coli DH5α. c, Bacterial growth in minimal media supplemented with individual aAAs at 37 °C for 16 h. Data indicates mean with SEM from three replicates. d. LC-MS analysis of aromatic monoamines produced by overexpression of Efm_TyrDC, Rgs_TrpDC or Mmi_Gln/TyrDC in E. coli DH5α. Data indicates individual value of three replicates.
Extended Data Fig. 4 Enzyme kinetics of E. faecium Com15 tyrosine decarboxylase.
a, SDS-PAGE analysis of the purification of His6-tagged E. faecium Com15 tyrosine decarboxylase (Efm_TyrDC). At least two independent experiments were performed. b, LC-MS analysis of the time-dependent formation of tyramine from enzymatic transformation of tyrosine by 100 nM Efm_TyrDC at room temperature. c, LC-MS determination of tryptamine, tyramine, phenethylamine and histamine from enzymatic transformation of tryptamine, tyrosine, phenylalanine and histidine by 100 nM Efm_TyrDC at 37 °C for 10 min. d, PRESTO-Tango assay for Efm_TyrDC enzymatic products for GPRC5A activation. e, Michaeslis-Menten kinetics study of Efm_TyrDC catalyzed transformation of tyrosine and phenylalanine. Data indicates mean with SEM from three replicates (d). Two-way ANOVA using Sidak’s multiple comparisons test, ****p < 0.0001.
Extended Data Fig. 5 Genetic manipulation of E. faecium Com15 TyrDC.
a, Genetic deletion of segment of DNA disrupts TyrDC gene expression in E. faecium Com15. b, Agarose gel electrophoresis of the PCR products from wild-type and TyrDC gene-disrupted E. faecium Com15. c, Optical density measurement of bacterial growth including wild-type E. faecium Com15 (Efm_wt), E. faecium Com15 TyrDC knock-out (Efm_Δtdc), complementation of vector control (pKH12) or TyrDC (pKH12-tdc) in Efm_Δtdc at 37 °C for 16 h. d, PRESTO-Tango assay for examining bacterial cultures for GPRC5A activation. Bacterial growth media were filtered through 0.2 µM membrane followed by 3,000 MWKO membrane. Data indicates mean with SEM from three replicates (c,d).
Extended Data Fig. 6 Untargeted metabolomics of E. faecium Com15.
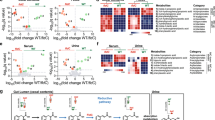
a, Principal component analysis (PCA) of the molecular features identified by the untargeted metabolomics of BHI media, E. faecium Com15 wild-type (Efm_WT) and tyrDC deletion strain (Efm_∆tdc) cultures. b, Volcano plot of the molecular features from Efm_WT and Efm_∆tdc strains. Dashed lines indicate q-value = 0.01, five folds change in five replicates. c, Fold change ranking of representative metabolites identified in Efm_WT in comparison with Efm_∆tdc cultures. d-g, Mass intensity of the representative metabolites identified in Efm_WT cultures. Data indicates mean with SEM from five replicates. h, PRESTO-Tango assay for the representative aromatic amines. Data indicates mean with SEM from three replicates.
Extended Data Fig. 7 Screening of aromatic monoamine derivatives.
a, PRESTO-Tango assay for screening a collection of tryptamine derivatives (100 µM) for GPRC5A activation. b, Dose-response curves of GPRC5A activation upon aromatic monoamines treatment. c, d, LDH cytotoxicity assay and MTT cell viability assay in mammalian cells. e, PRESTO-Tango assay for screening a collection of aromatic monoamines and phenylethylamine derivatives (100 µM) for GPRC5A activation. Data indicates mean with SEM from three replicates (a,b,c,d). One-way ANOVA using Dunnett’s multiple comparisons test.
Extended Data Fig. 8 PRESTO-Tango assay for GPCRs.
a, b, Evaluation of 100 µM of TA and 7FTA on class C orphan GPCRs (GPRC5A,B,C,D) and aminergic GPCRs (ADRA2A, DRD2, HTR4) activation. c, Dose-response curves of 7FTA on class C orphan GPCRs (GPRC5A,B,C,D) and aminergic GPCRs (ADRA2A, DRD2, HTR4) activation. d, Examination of endogenous neurotransmitters epinephrine (10 µM), dopamine (10 µM) and serotonin (100 µM) on GPRC5A activation e, Western blot analysis of FLAG-tagged GPCR-Tango constructs in HTLA cells. At least two independent experiments were performed. Data indicates mean with SEM from three replicates (a,b,c,d). Two-way ANOVA using Tukey’s multiple comparisons test (a-b), One-way ANOVA using Dunnett’s multiple comparisons test (b), ****p < 0.0001.
Extended Data Fig. 9 Assays for GPRC5A signaling.
a, NF-κB-dependent luciferase reporter assay in HEK293 cells transfected with different amount of vector, GRPC5A or ADRA2A plasmids followed by the treatment 10 ng/mL TNFα for 8 h. b, Dose response curves of 7FTA in the NF-κB-dependent luciferase reporter assay. HEK293 cells transfected with vector or GRPC5A plasmids were treated with increasing concentration of 7FTA with or without 10 ng/mL TNFα for 8 h. 0.1% DMSO was used as the vehicle control. c, Dose response curves of 7FTA in the G-protein activated luciferase reporter assay. HEK293 cells were transfected with GPRC5A plasmid and reporter plasmids for cAMP response element (CRE), nuclear factor of activated T-Cells (NFAT), or serum response element (SRE). Cells were then treated with increasing concentrations of 7FTA for 8 h. In CRE + forskolin samples, cells were also treated with 1 µM forskolin. d, BRET-based TRUPATH assay for GPRC5A in HEK293 cells upon DMSO or 100 µM 7FTA treatment. Data indicates mean with SEM from three replicates (a,b,c,d). Two-way ANOVA using Tukey’s multiple comparisons test (a-b), ****p < 0.0001.
Extended Data Fig. 10 RNAseq analysis of wild-type and GPRC5A-KO HT-29 cells.
a, Principal component analysis (PCA) plot of RNAseq data for wild-type and GPRC5A-KO HT-29 cells upon treatment with 0.1% DMSO, 50 µM 7FTA, 10 ng/mL TNFa with or without 7FTA. b, Volcano plot for the differential gene expression in wild-type HT-29 cells upon treatment with 0.1% DMSO or 50 µM 7FTA. Significant genes indicate an adjusted p value <0.05 and two folds change in three replicates. c, RT-qPCR analysis of respective fold gene expression level against ACTB for wild-type and GPRC5A-KO cells treatment with DMSO or 7FTA. Data represent fold gene expression in four replicates. d–f, RT-qPCR analysis of respective fold gene expression level for wild-type and GPRC5A-KO cells treated 0.1% DMSO, 10 ng/mL TNFα with or without 50 µM 7FTA for 8 h. Data indicates mean with SEM from four replicates (d,e) and three replicates (f). One-way ANOVA using Tukey’s multiple comparisons test, ****p < 0.0001.
Supplementary information
Supplementary Information
Supplementary Figs. 1–7, Supplementary Tables 1–4 and Supplementary Note.
Source data
Source Data Fig. 1
Unprocessed gels.
Source Data Fig. 2
Unprocessed gels.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Unprocessed gels.
Source Data Fig. 5
Unprocessed gels.
Source Data Extended Data Fig. 1
Unprocessed gels.
Source Data Extended Data Fig. 2
Unprocessed gels.
Source Data Extended Data Fig. 3
Unprocessed gels.
Source Data Extended Data Fig. 5
Unprocessed gels
Source Data Extended Data Fig. 8
Unprocessed gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, X., Stein, K.R., Chen, V. et al. Chemoproteomics reveals microbiota-derived aromatic monoamine agonists for GPRC5A. Nat Chem Biol 19, 1205–1214 (2023). https://doi.org/10.1038/s41589-023-01328-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-023-01328-z
This article is cited by
-
Decrypting orphan GPCR drug discovery via multitask learning
Journal of Cheminformatics (2024)
-
Dopamine receptor D2 confers colonization resistance via microbial metabolites
Nature (2024)