Abstract
Ferroptosis is an iron-dependent form of cell death driven by oxidation of polyunsaturated fatty acid (PUFA) phospholipids. Large-scale genetic screens have uncovered a specialized role for PUFA ether phospholipids (ePLs) in promoting ferroptosis. Understanding of the enzymes involved in PUFA-ePL production, however, remains incomplete. Here we show, using a combination of pathway mining of genetic dependency maps, AlphaFold-guided structure predictions and targeted lipidomics, that the uncharacterized transmembrane protein TMEM164—the genetic ablation of which has been shown to protect cells from ferroptosis—is a cysteine active center enzyme that selectively transfers C20:4 acyl chains from phosphatidylcholine to lyso-ePLs to produce PUFA ePLs. Genetic deletion of TMEM164 across a set of ferroptosis-sensitive cancer cell lines caused selective reductions in C20:4 ePLs with minimal effects on C20:4 diacyl PLs, and this lipid profile produced a variable range of protection from ferroptosis, supportive of an important but contextualized role for C20:4 ePLs in this form of cell death.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw sequencing data validating genomic alterations in 786-O base-edited and knockout cell lines are deposited at the Sequence Read Archive (accession number PRJNA902005). Data supporting the findings of this work are available within the paper and its Supplementary Information. The datasets, constructs and cell models generated and analyzed during the current study are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
van Meer, G., Voelker, D. R. & Feigenson, G. W. Membrane lipids: where they are and how they behave. Nat. Rev. Mol. Cell Biol. 9, 112–124 (2008).
Yang, W. S. et al. Peroxidation of polyunsaturated fatty acids by lipoxygenases drives ferroptosis. Proc. Natl Acad. Sci. USA 113, E4966–E4975 (2016).
Stockwell, B. R. et al. Ferroptosis: a regulated cell death nexus linking metabolism, redox biology, and disease. Cell 171, 273–285 (2017).
Yang, W. S. et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 156, 317–331 (2014).
Eaton, J. K. et al. Selective covalent targeting of GPX4 using masked nitrile-oxide electrophiles. Nat. Chem. Biol. 16, 497–506 (2020).
Zou, Y. et al. Plasticity of ether lipids promotes ferroptosis susceptibility and evasion. Nature 585, 603–608 (2020).
Cui, W., Liu, D., Gu, W. & Chu, B. Peroxisome-driven ether-linked phospholipids biosynthesis is essential for ferroptosis. Cell Death Differ. 28, 2536–2551 (2021).
Wainberg, M. et al. A genome-wide atlas of co-essential modules assigns function to uncharacterized genes. Nat. Genet. 53, 638–649 (2021).
Gallego-Garcia, A. et al. A bacterial light response reveals an orphan desaturase for human plasmalogen synthesis. Science 366, 128–132 (2019).
Werner, E. R. et al. The TMEM189 gene encodes plasmanylethanolamine desaturase which introduces the characteristic vinyl ether double bond into plasmalogens. Proc. Natl Acad. Sci. USA 117, 7792–7798 (2020).
Doll, S. et al. ACSL4 dictates ferroptosis sensitivity by shaping cellular lipid composition. Nat. Chem. Biol. 13, 91–98 (2017).
Dixon, S. J. et al. Human haploid cell genetics reveals roles for lipid metabolism genes in nonapoptotic cell death. ACS Chem. Biol. 10, 1604–1609 (2015).
Sun, W. Y. et al. Phospholipase iPLA2β averts ferroptosis by eliminating a redox lipid death signal. Nat. Chem. Biol. 17, 465–476 (2021).
Kramer, R. M. & Deykin, D. Arachidonoyl transacylase in human platelets. Coenzyme A-independent transfer of arachidonate from phosphatidylcholine to lysoplasmenylethanolamine. J. Biol. Chem. 258, 13806–13811 (1983).
Sugiura, T., Masuzawa, Y., Nakagawa, Y. & Waku, K. Transacylation of lyso platelet-activating factor and other lysophospholipids by macrophage microsomes. Distinct donor and acceptor selectivities. J. Biol. Chem. 262, 1199–1205 (1987).
Tsherniak, A. et al. Defining a cancer dependency map. Cell 170, 564–576 (2017).
Horibata, Y. & Hirabayashi, Y. Identification and characterization of human ethanolaminephosphotransferase1. J. Lipid Res. 48, 503–508 (2007).
Zou, Y. et al. A GPX4-dependent cancer cell state underlies the clear-cell morphology and confers sensitivity to ferroptosis. Nat. Commun. 10, 1617 (2019).
Zou, Y. et al. Cytochrome P450 oxidoreductase contributes to phospholipid peroxidation in ferroptosis. Nat. Chem. Biol. 16, 302–309 (2020).
Lee, H. C. et al. LPIAT1 regulates arachidonic acid content in phosphatidylinositol and is required for cortical lamination in mice. Mol. Biol. Cell 23, 4689–4700 (2012).
Reed, A. et al. LPCAT3 inhibitors remodel the polyunsaturated phospholipid content of human cells and protect from ferroptosis. ACS Chem. Biol. 17, 1607–1618 (2022).
Bourgeois, T. et al. Deletion of lysophosphatidylcholine acyltransferase 3 in myeloid cells worsens hepatic steatosis after a high-fat diet. J. Lipid Res. 62, 100013 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Parsons, W. H. et al. AIG1 and ADTRP are atypical integral membrane hydrolases that degrade bioactive FAHFAs. Nat. Chem. Biol. 12, 367–372 (2016).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Gabler, F. et al. Protein sequence analysis using the MPI Bioinformatics Toolkit. Curr. Protoc. Bioinformatics 72, e108 (2020).
Varadi, M. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2022).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Holm, L. Dali server: structural unification of protein families. Nucleic Acids Res. 50, W210–W215 (2022).
van Kempen, M. et al. Foldseek: fast and accurate protein structure search. Preprint at bioRxiv https://doi.org/10.1101/2022.02.07.479398 (2022).
Ayoub, R. & Lee, Y. RUPEE: a fast and accurate purely geometric protein structure search. PLoS ONE 14, e0213712 (2019).
Glab, B. et al. Cloning of glycerophosphocholine acyltransferase (GPCAT) from fungi and plants: a novel enzyme in phosphatidylcholine synthesis. J. Biol. Chem. 291, 25066–25076 (2016).
Tian, W., Chen, C., Lei, X., Zhao, J. & Liang, J. CASTp 3.0: computed atlas of surface topography of proteins. Nucleic Acids Res. 46, W363–W367 (2018).
Xu, Y. et al. CavityPlus: a web server for protein cavity detection with pharmacophore modelling, allosteric site identification and covalent ligand binding ability prediction. Nucleic Acids Res. 46, W374–W379 (2018).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Yan, H. F. et al. Ferroptosis: mechanisms and links with diseases. Signal Transduct. Target Ther. 6, 49 (2021).
Magtanong, L. et al. Context-dependent regulation of ferroptosis sensitivity. Cell Chem. Biol. 29, 1409–1418 (2022).
Zallot, R., Oberg, N. & Gerlt, J. A. The EFI web resource for genomic enzymology tools: leveraging protein, genome, and metagenome databases to discover novel enzymes and metabolic pathways. Biochemistry 58, 4169–4182 (2019).
Brown, S. D. & Babbitt, P. C. New insights about enzyme evolution from large scale studies of sequence and structure relationships. J. Biol. Chem. 289, 30221–30228 (2014).
Jonas, A. Lecithin cholesterol acyltransferase. Biochim. Biophys. Acta 1529, 245–256 (2000).
Ogura, Y., Parsons, W. H., Kamat, S. S. & Cravatt, B. F. A calcium-dependent acyltransferase that produces N-acyl phosphatidylethanolamines. Nat. Chem. Biol. 12, 669–671 (2016).
Bazan, J. F. & Fletterick, R. J. Viral cysteine proteases are homologous to the trypsin-like family of serine proteases: structural and functional implications. Proc. Natl Acad. Sci. USA 85, 7872–7876 (1988).
Higaki, J. N., Evnin, L. B. & Craik, C. S. Introduction of a cysteine protease active site into trypsin. Biochemistry 28, 9256–9263 (1989).
Baird, T. T. Jr, Wright, W. D. & Craik, C. S. Conversion of trypsin to a functional threonine protease. Protein Sci. 15, 1229–1238 (2006).
Richter, F. et al. Computational design of catalytic dyads and oxyanion holes for ester hydrolysis. J. Am. Chem. Soc. 134, 16197–16206 (2012).
Zoeller, R. A. et al. Plasmalogens as endogenous antioxidants: somatic cell mutants reveal the importance of the vinyl ether. Biochem. J. 338, 769–776 (1999).
Vinogradova, E. V. et al. An activity-guided map of electrophile-cysteine interactions in primary human T cells. Cell 182, 1009–1026 (2020).
Bar-Peled, L. et al. Chemical proteomics identifies druggable vulnerabilities in a genetically defined cancer. Cell 171, 696–709 (2017).
Snyder, F. & Wood, R. Alkyl and alk-1-enyl ethers of glycerol in lipids from normal and neoplastic human tissues. Cancer Res. 29, 251–257 (1969).
Albert, D. H. & Anderson, C. E. Ether-linked glycerolipids in human brain tumors. Lipids 12, 188–192 (1977).
Cajka, T., Smilowitz, J. T. & Fiehn, O. Validating quantitative untargeted lipidomics across nine liquid chromatography–high-resolution mass spectrometry platforms. Anal. Chem. 89, 12360–12368 (2017).
Koch, J. et al. Unequivocal mapping of molecular ether lipid species by LC–MS/MS in plasmalogen-deficient mice. Anal. Chem. 92, 11268–11276 (2020).
Huang, T. P., Newby, G. A. & Liu, D. R. Precision genome editing using cytosine and adenine base editors in mammalian cells. Nat. Protoc. 16, 1089–1128 (2021).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Acknowledgements
This work was supported by the National Institutes of Health (CA231991), the Damon Runyon Cancer Research Foundation (H.L.) and Lundbeck. We are grateful to J. Blankman, H. Reardon and M. Niphakis for helpful discussions throughout the performance of this research.
Author information
Authors and Affiliations
Contributions
A.R., T.W., J.F.B. and B.F.C. conceived of the project and wrote the paper. A.R. and T.W. generated genetically engineered cell lines and performed lipidomics, biochemical assays and western blotting experiments. J.F.B. performed computational structural analysis studies linking TMEM164 to the AIG1/ADTRP family of enzymes. H.L. assisted with targeted genomic sequencing and data analysis.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Eun-Woo Lee and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
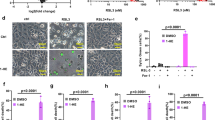
Extended Data Fig. 1 Characterization of sgTMEM164 786-O cells.
a, b, Confirmation of genetic alterations in 786-O cells at sgRNA sites targeted with (a) sgTMEM164-1 and (b) sgTMEM164-2 by next-generation sequencing analysis. c, d, Genetic deletion of TMEM164 in 786-O cells results in reduced sensitivity to ferroptosis induced by the GPX4 inhibitors ML210 (c) and RSL3 (d) measured at 24 h post-treatment with GPX4 inhibitors. Data represent mean values ± S.E.M. from two independent experiments.
Extended Data Fig. 2 Targeted lipidomic (LC-MS/MS) methods for measuring ePLs.
a, b, Detection of endogenous ePE-P(C18:0/C20:4) in 786-O cells by LC-MS/MS (a) and comparison to commercial ePE-P(C18:0/C20:4) standard (b). Left traces show extracted ion chromatogram of a feature corresponding to ePE-P(C18:0/C20:4) (750.5 → 303.3). Right traces show MS2 spectra for ePE-P(C18:0/C20:4). c,d, Targeted LC-MS/MS analysis of isobaric ePE-O and ePE-P species performed as described in the Methods section revealing that they are separated by approximately 1.5 min difference in retention time. Treatment of lipid extracts from 786-O cells with 10% (v/v) formic acid (c) or HCl (3 N) (d) for 30 min at 37 °C reveals ePE-P elutes after ePE-O as evidenced by a significant reduction in the ePE-P peak following HCl treatment. AUC, area under the curve.
Extended Data Fig. 3 Alterations in ePC and diacyl PC production in TMEM164-deficient cells.
a, b ePC (a) and PC (b) lipid measurements in sgCtrl and sgTMEM164 786-O cells. For a, due to technical limitations we did not distinguish between ePC-O and ePC-P lipids. Data represent mean values ± S.E.M from four independent experiments per group. P-values were derived using a Two-sided Student’s t-test performed relative to sgCtrl cells; only shown for lipids where sgCtrl and parental (WT) data did not differ.
Extended Data Fig. 4 Measurement of free fatty acids (FFAs), eLPE, and LPE in TMEM164-deficient cells.
FFA (a), eLPE (b) and LPE (c) lipid measurements in sgCtrl and sgTMEM164 786-O cells. Data represent mean values ± S.E.M from four independent experiments per group. P-values were derived using a Two-sided Student’s t-test performed relative to sgCtrl cells.
Extended Data Fig. 5 Integrated homology mapping and AlphaFold2 analysis identifies a predicted shared structure between TMEM164 and the AIG1/ADTRP lipid hydrolases.
a, Structure-based sequence alignment by DALI of AlphaFold2 models (http://alphafold.ebi.ac.uk) of the H.sapiens (UniProt Q5U3C3) and D.melanogaster (UniProt Q7JRB2) TMEM164 proteins, with representative proteins from the AIG1/ADTRP family (H.sapiens (Q9NVV5); S.cerevisiae (P38842)), prokaryotic YwaF (S.pneumoniae (Q8CYG1); B.subtilis (P25149)) and YpjA proteins (S.aureus (Q2FYH4); A.veneficus (F2KNL0)), and GPC1 enzymes (S.cerevisiae (P48236); A.thaliana (Q9FJB4). In some cases, models were built with ColabFold implementation of AlphaFold2 (http://github.com/sokrypton/ColabFold). The alignment was displayed and analyzed with Jalview (http://jalview.org), showing the six blocks of conserved structure corresponding to TM helices 1-6. Length and sequence-variable gaps in the alignment corresponding to loops in the superposed folds were abbreviated by residue stretches in parentheses. Positions of the variable nucleophile and invariant His residues (Cys123 and His181 in human TMEM164, respectively) are boxed and labeled. b, DALI-derived superpositions provide a measure of structural relatedness for the six main branches of the enzyme superfamily displayed in the phylogenetic hypertree in Fig. 4c (http://kinase.com/tools/HyperTree.html), with chains aligned in panel a noted by red circles. Z-score of 8.2 is provided for the most distant branch (YpjA). c, Matrix of DALI Z-scores displaying the clusters of more closely related AlphaFold2-derived structures from all-against-all superpositions performed with the DALI server (http://ekhidna2.biocenter.helsinki.fi), leading to reconstitution of the six branches of the enzyme superfamily. Five of these clusters have sequence alignments that can be retrieved from the PFAM database (http://pfam.xfam.org): TMEM164 (PF14808), AIG1/ADTRP (PF04750), GPC1 (PF10998), YwaF (PF09529) and YpjA (PF7187). d, Tube representations of aligned AlphaFold2 models showing positions of the catalytic dyad with black Ca spheres and preservation of the core 6TM fold (color-ramped from blue TM1 to red TM6) with N- or C-terminal extensions drawn in grey. Figures composed with PyMOL (http://pymol.org).
Extended Data Fig. 6 AlphaFold2 analysis of TMEM164 active site cavity.
a, The human TMEM164 structural model was processed by the ConSurf server (http://consurf.tau.ac.il) to map the sequence conservation profile of its underlying family to the 3D fold surface color-ramped from blue (conserved) to red (variable) patches. For side and top views, the ConSurf surfaces are aligned to the ribbon model of TMEM164 embedded with the CavityPlus-calculated volume in pink (server at http://pkumdl.cn), showing the positions of predicted portals to the lipid bilayer. In addition, the CavPharmer tool at CavityPlus was used to predict a pharmacophore skeleton for a prospective ligand (which could represent the lipid substrates or product of the enzymatic reaction) for TMEM164, showing a shape and chemical nature in agreement with a predicted function as a C20:4-preferring lyso-ePL acyltransferase. b, The program CAVER (https://loschmidt.chemi.muni.cz/caverweb) was used to visualize the tunnels leading from the central TMEM164 cavity that abuts the Cys123/His181 predicted catalytic dyad (shown in black stick format), to the portals that open to the lipid bilayer (tunnels 1 and 3, respectively in blue and orange, with lengths 12.8 Å and 16.7 Å) and to the outer leaflet of the membrane (tunnels 2 and 4, respectively in red and green, both with lengths of 15.1 Å). c, A top view of the six-transmembrane core of TMEM164 with docked C20:4 ePE, with acyl chains occupying tunnels 1 and 3, and head group cresting into the tunnel 2 space. Tunnels 1 and 3 may represent the principal entrance and exit routes for substrates and product, that place the headgroup in proximity of the active site.
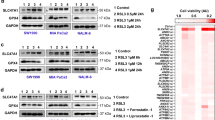
Extended Data Fig. 7 Characterization of C20:4 PC-dependent acyltransferase activity of TMEM164.
a, Western blot of membrane lysates from sgCtrl and sgTMEM164-2 786-O cells recombinantly expressing C-terminally FLAG-tagged WT-TMEM164 or a C123A-TMEM164 mutant. WT-TMEM164 lysates were serially diluted with mock lysate to identify a dilution (1:1) that matched the expression of C123A-TMEM164. Data are from a single experiment representative of two independent experiments. b, Membrane lysates from sgCtrl or sgTMEM164 786-O cells were treated with N-ethylmaleimide (NEM; 200 µM, 60 min) prior to exposure to diacyl PC(C18:0/C20:4-d8) and lyso-ePE-P(C18:0) substrates and measurement of ePE-P(C18:0/C20:4-d8) product. c, d, Measurement of (c) ePE-P(C18:0/C20:4-d8) and (d) diacyl PE(C17:1/C20:4-d8) formation in membrane lysates from sgCtrl and sgLPCAT3 786-O cells using diacyl PC(C18:0/C20:4-d8) and ePE-P(C18:0) or LPE(C17:1) as donor and acceptor substrate, respectively. For b-d, membrane lysates (1 µg/µL) were treated with 50 µM each of donor and acceptor substrates for 1 h at 37 °C prior to analysis. For b-d data represent mean values ± S.E.M from three independent experiments per group. P-values were derived using a Two-sided Student’s t-test performed relative to sgCtrl cells.
Extended Data Fig. 8 Characterization of sgTMEM164-C123Y base-edited cells.
a, Confirmation of base-editing of TMEM164 genomic DNA in 786-O cells at the sgRNA target site for sgTMEM164-C123Y base-edited cell population by next-generation sequencing analysis. b, Measurement of C20:4 (left) and additional (right) ePE-O lipids in sgCtrl and sgTMEM164-C123Y base-edited cell populations. c, Measurement of diacyl PE lipids in sgCtrl and sgTMEM164-C123Y base-edited 786-O cell populations. d, Cell viability of sgCtrl and sgTMEM164-C123Y base-edited cell populations treated with the indicated concentrations of the GPX4 inhibitor RSL3 measured at 24 h post-treatment. For b and c data represent mean values ± S.E.M from four independent experiments per group. For d, data represent mean values ± S.E.M. from two independent experiments. P-values were derived using a Two-sided Student’s t-test performed relative to sgCtrl cells.
Extended Data Fig. 9 C20:4 PE and non-C20:4 ePE-P measurements in TMEM164-deficient RKN and ES-2 cells.
a, b, Measurement of C20:4 PE lipids in sgCtrl and sgTMEM164-2 RKN (a) and ES-2 (b) cell lines. c, d, Measurement of non-C20:4 ePE-P lipids in sgCtrl and sgTMEM164-2 RKN (c) and ES-2 (d) cell lines. Data represent mean values ± S.E.M from four independent experiments per group. P-values were derived using a Two-sided Student’s t-test performed relative to sgCtrl cells.
Extended Data Fig. 10 TMEM164 catalytic mechanism of action.
Proposed catalytic mechanism for C20:4 PL-dependent acyltransferase activity of TMEM164. LPC, lysophosphatidylcholine.
Supplementary information
Supplementary Data 1
Supplementary datasets.
Source data
Source Data Fig. 1
Statistical source data related to Fig. 1.
Source Data Fig. 2
Statistical source data related to Fig. 2.
Source Data Fig. 4
Statistical source data related to Fig. 4.
Source Data Fig. 5
Statistical source data related to Fig. 5.
Source Data Fig. 5
Unprocessed western blots related to Fig. 5.
Source Data Extended Data Fig. 1
Statistical source data related to Extended Data Fig. 1.
Source Data Extended Data Fig. 3
Statistical source data related to Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Statistical source data related to Extended Data Fig. 4.
Source Data Extended Data Fig. 7
Statistical source data related to Extended Data Fig. 7.
Source Data Extended Data Fig. 7
Unprocessed western blots related to Extended Data Fig. 7.
Source Data Extended Data Fig. 8
Statistical source data related to Extended Data Fig. 8.
Source Data Extended Data Fig. 9
Statistical source data related to Extended Data Fig. 9.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Reed, A., Ware, T., Li, H. et al. TMEM164 is an acyltransferase that forms ferroptotic C20:4 ether phospholipids. Nat Chem Biol 19, 378–388 (2023). https://doi.org/10.1038/s41589-022-01253-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-022-01253-7
This article is cited by
-
Ferroptotic therapy in cancer: benefits, side effects, and risks
Molecular Cancer (2024)
-
The cell biology of ferroptosis
Nature Reviews Molecular Cell Biology (2024)
-
Helicobacter pylori CagA-mediated ether lipid biosynthesis promotes ferroptosis susceptibility in gastric cancer
Experimental & Molecular Medicine (2024)
-
A guideline on the molecular ecosystem regulating ferroptosis
Nature Cell Biology (2024)
-
An integrated view of lipid metabolism in ferroptosis revisited via lipidomic analysis
Experimental & Molecular Medicine (2023)