Abstract
Chemically inducible systems represent valuable synthetic biology tools that enable the external control of biological processes. However, their translation to therapeutic applications has been limited because of unfavorable ligand characteristics or the immunogenicity of xenogeneic protein domains. To address these issues, we present a strategy for engineering inducible split protein regulators (INSPIRE) in which ligand-binding proteins of human origin are split into two fragments that reassemble in the presence of a cognate physiological ligand or clinically approved drug. We show that the INSPIRE platform can be used for dynamic, orthogonal and multiplex control of gene expression in mammalian cells. Furthermore, we demonstrate the functionality of a glucocorticoid-responsive INSPIRE platform in vivo and apply it for perturbing an endogenous regulatory network. INSPIRE presents a generalizable approach toward designing small-molecule responsive systems that can be implemented for the construction of new sensors, regulatory networks and therapeutic applications.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data supporting the findings of this study are available within the article or Supplementary Information. Plasmid maps, including complete and annotated GenBank files, are provided in Supplementary Data. A subset of key plasmids used in this study will be made available on Addgene. Source data are provided with this paper. Any other relevant data or reagents are available from the corresponding author upon reasonable request.
References
Stanton, B. Z., Chory, E. J. & Crabtree, G. R. Chemically induced proximity in biology and medicine. Science 359, eaa05902 (2018).
Derose, R., Miyamoto, T. & Inoue, T. Manipulating signaling at will: chemically-inducible dimerization (CID) techniques resolve problems in cell biology. Pflug. Arch. Eur. J. Physiol. 465, 409–417 (2013).
Fegan, A., White, B., Carlson, J. C. T. & Wagner, C. R. Chemically controlled protein assembly: techniques and applications. Chem. Rev. 110, 3315–3336 (2010).
Miyamoto, T. et al. Rapid and orthogonal logic gating with a gibberellin-induced dimerization system. Nat. Chem. Biol. 8, 465–470 (2012).
Liang, F.-S., Ho, W. Q. & Crabtree, G. R. Engineering the ABA plant stress pathway for regulation of induced proximity. Sci. Signal. 4, rs2 (2011).
Spencer, D., Wandless, T., Schreiber, S. & Crabtree, G. Controlling signal transduction with synthetic ligands. Science 262, 1019–1024 (1993).
Putyrski, M. & Schultz, C. Protein translocation as a tool: the current rapamycin story. FEBS Lett. 586, 2097–2105 (2012).
Ziegler, M. J. et al. Mandipropamid as a chemical inducer of proximity for in vivo applications. Nat. Chem. Biol. 18, 64–69 (2022).
Kang, S. et al. COMBINES-CID: an efficient method for de novo engineering of highly specific chemically induced protein dimerization systems. J. Am. Chem. Soc. 141, 10948–10952 (2019).
Glasgow, A. A. et al. Computational design of a modular protein sense-response system. Science 366, 1024–1028 (2019).
Foight, G. W. et al. Multi-input chemical control of protein dimerization for programming graded cellular responses. Nat. Biotechnol. 37, 1209–1216 (2019).
Fink, T. et al. Design of fast proteolysis-based signaling and logic circuits in mammalian cells. Nat. Chem. Biol. 15, 115–122 (2019).
Gao, Y. et al. Complex transcriptional modulation with orthogonal and inducible dCas9 regulators. Nat. Methods 13, 1043–1049 (2016).
Liu, R., Liu, C.-B., Mohi, M. G., Arai, K. & Watanabe, S. Analysis of mechanisms involved in the prevention of γ irradiation-induced apoptosis by hGM-CSF. Oncogene 19, 571–579 (2000).
Gallego, O. et al. Detection and characterization of protein interactions in vivo by a simple live-cell imaging method. PLoS ONE 8, e62195 (2013).
Barlow, A. D., Nicholson, M. L. & Herbert, T. P. Evidence for rapamycin toxicity in pancreatic β-cells and a review of the underlying molecular mechanisms. Diabetes 62, 2674–2682 (2013).
Sun, J. & Sadelain, M. The quest for spatio-temporal control of CAR T cells. Cell Res. 25, 1281–1282 (2015).
Schellekens, H. Factors influencing the immunogenicity of therapeutic proteins. Nephrol. Dial. Transplant. 20(Suppl 6), vi3–9 (2005).
Škrlec, K., Štrukelj, B. & Berlec, A. Non-immunoglobulin scaffolds: a focus on their targets. Trends Biotechnol. 33, 408–418 (2015).
Riddell, S. R. et al. T-cell mediated rejection of gene-modified HIV-specific cytotoxic T lymphocytes in HIV-infected patients. Nat. Med. 2, 216–223 (1996).
Hill, Z. B., Martinko, A. J., Nguyen, D. P. & Wells, J. A. Human antibody-based chemically induced dimerizers for cell therapeutic applications. Nat. Chem. Biol. 14, 112–117 (2018).
Feng, J. et al. A general strategy to construct small molecule biosensors in eukaryotes. eLife 4, e10606 (2015).
Dirnberger, D., Unsin, G., Schlenker, S. & Reichel, C. A small-molecule-protein interaction system with split-ubiquitin as sensor. ChemBioChem 7, 936–942 (2006).
Ottmann, O. et al. Long-term efficacy and safety of dasatinib in patients with chronic myeloid leukemia in accelerated phase who are resistant to or intolerant of imatinib. Blood Cancer J. 8, 88 (2018).
Cronstein, B. N. & Aune, T. M. Methotrexate and its mechanisms of action in inflammatory arthritis. Nat. Rev. Rheumatol. 16, 145–154 (2020).
Vandyke, K. et al. Therapeutic concentrations of dasatinib inhibit in vitro osteoclastogenesis. Leukemia 23, 994–997 (2009).
Joannon, P., Oviedo, I., Campbell, M. & Tordecilla, J. High-dose methotrexate therapy of childhood acute lymphoblastic leukemia: lack of relation between serum methotrexate concentration and creatinine clearance. Pediatr. Blood Cancer 43, 17–22 (2004).
Sladek, F. M. What are nuclear receptor ligands? Mol. Cell. Endocrinol. 334, 3–13 (2011).
He, Y. et al. Structures and mechanism for the design of highly potent glucocorticoids. Cell Res. 24, 713–726 (2014).
Barnes, P. J. Anti-inflammatory actions of glucocorticoids: molecular mechanisms. Clin. Sci. 94, 557–572 (1998).
Saponaro, F., Sestito, S., Runfola, M., Rapposelli, S. & Chiellini, G. Selective thyroid hormone receptor-beta (TRβ) agonists: new perspectives for the treatment of metabolic and neurodegenerative disorders. Front. Med. (Lausanne) 7, 331 (2020).
Jameera Begam, A., Jubie, S. & Nanjan, M. J. Estrogen receptor agonists/antagonists in breast cancer therapy: a critical review. Bioorg. Chem. 71, 257–274 (2017).
Grygiel-Górniak, B. Peroxisome proliferator-activated receptors and their ligands: nutritional and clinical implications – a review. Nutr. J. 13, 17 (2014).
Cuzzocrea, S. et al. Rosiglitazone, a ligand of the peroxisome proliferator-activated receptor-γ, reduces acute inflammation. Eur. J. Pharmacol. 483, 79–93 (2004).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Gao, Y. et al. Complex transcriptional modulation with orthogonal and inducible dCas9 regulators. Nat. Methods 13, 1043–1049 (2016).
Khalil, A. S. & Collins, J. J. Synthetic biology: applications come of age. Nat. Rev. Genet. 11, 367–379 (2010).
Ma, D., Peng, S. & Xie, Z. Integration and exchange of split dCas9 domains for transcriptional controls in mammalian cells. Nat. Commun. 7, 13056 (2016).
Lebar, T., Verbič, A., Ljubetič, A. & Jerala, R. Polarized displacement by transcription activator-like effectors for regulatory circuits. Nat. Chem. Biol. 15, 80–87 (2019).
Gao, X. J., Chong, L. S., Kim, M. S. & Elowitz, M. B. Programmable protein circuits in living cells. Science 361, 1252–1258 (2018).
Gong, S. et al. Dynamics and correlation of serum cortisol and corticosterone under different physiological or stressful conditions in mice. PLoS ONE 10, e0117503 (2015).
Tritos, N. A., Biller, B. M. K. & Swearingen, B. Management of Cushing disease. Nat. Rev. Endocrinol. 7, 279–289 (2011).
Agorastos, A. & Chrousos, G. P. The neuroendocrinology of stress: the stress-related continuum of chronic disease development. Mol. Psychiatry 27, 502–513 (2022).
Russell, G. & Lightman, S. The human stress response. Nat. Rev. Endocrinol. 15, 525–534 (2019).
Tinberg, C. E. et al. Computational design of ligand-binding proteins with high affinity and selectivity. Nature 501, 212–216 (2013).
Bick, M. J. et al. Computational design of environmental sensors for the potent opioid fentanyl. eLife 6, e28909 (2017).
Dolberg, T. B. et al. Computation-guided optimization of split protein systems. Nat. Chem. Biol. 17, 531–539 (2021).
Wu, H. D. et al. Rational design and implementation of a chemically inducible heterotrimerization system. Nat. Methods 17, 928–936 (2020).
Delfosse, V., Maire, A., Le, Balaguer, P. & Bourguet, W. A structural perspective on nuclear receptors as targets of environmental compounds. Acta Pharmacol. Sin. 36, 88–101 (2015).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Konc, J., Lešnik, S., Škrlj, B. & Janežič, D. ProBiS-Dock Database: a web server and interactive web repository of small ligand–protein binding sites for drug design. J. Chem. Inf. Model. 61, 4097–4107 (2021).
Brooks, B. R. et al. CHARMM: the biomolecular simulation program. J. Comput. Chem. 30, 1545–1614 (2009).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25, 402–408 (2001).
Acknowledgements
This research was funded by grants from the Slovenian research agency (P4-0176 (R.J.), N4-0080 (R.J.), J1-9173 (R.J.), J1-2471 (U.B.) and P2-0046 (U.B.).).
Author information
Authors and Affiliations
Contributions
E.R. and T.L. performed the experiments on cell culture. D.L. performed the experiments on mice. K.K., S.L. and U.B. performed the bioinformatics analysis. E.R. and T.L. designed and analyzed the experiments. E.R., T.L. and R.J. wrote the manuscript. R.J. conceived of the study.
Corresponding author
Ethics declarations
Competing interests
The authors E.R., T.L., and R.J. are inventors of the EPO Patent application LU102654 ‘Engineering Chemically Inducible Split Protein Actuators (INSPIRE)’, submitted by the National Institute of Chemistry. All other authors declare that they have no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Srivatsan Raman and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Design of LYN-based INSPIRE system.
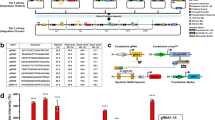
a, Left: crystal structure of Lyn kinase domain (LYN) in complex with dasatinib (DAS) (PDB ID 2zva). Splits are chosen in surface-exposed loops (cyan); α-helixes and β-strands are colored in dark and light gray, respectively. Right: aa sequence of LYN with additional annotations indicating split site locations and secondary structures. b, Schematic illustration showing the process of cloning split LYN fragments into reporter luciferase vectors (nLuc and cLuc). Split fragments were fused to split luciferase segments via a flexible gs10 linker. Additionally, constructs were tagged with either Myc- or HA-tag to confirm the expression of fusions via Western blot. c, Expression of N- (top) and C-terminal (bottom) split LYN fragments in HEK293T cells was verified with Western blot. Constitutively expressed mCitrine fluorescent protein was used as transfection control, and an empty plasmid vector (pcDNA3) was used as a negative control. The mCitrine transfection control was run on a separate gel. The transfection mixtures are provided in Supplementary Table 4. Data are representative of two independent experiments (n = 1). Original uncropped images are provided in the Source Data.
Extended Data Fig. 2 Analysis of split protein–ligand complex structural parameters.
a, Selected LBP is computationally split, and the contact number is calculated between nSplit–ligand (P1-L), nSplit–cSplit (P1-P2), and cSplit–ligand (P2-L). b–d, Calculated contact numbers for LYN in complex with DAS (PDB ID 2zva) (b), DHFR in complex with MTX (PDB ID 1u72) (c) and GR2 in complex with MOF (PDB ID 4e2j) (d). nSplit–ligand contacts (P1-L), cSplit–ligand (P2-L), and nSplit–cSplit (P1-P2) are represented by blue, yellow, and red lines, respectively. Arrows indicate the split site positions, and light shaded regions show the split sites showing the highest efficacy (split 267 for LYN, 173 for DHFR, and 175 for GR2). e, The results page of the ProteinSplit web application (http://elixir.fkkt.um.si/ProteinSplitIndex.html). f, Percentage of database proteins with the minimum number of α-helices or β-sheets located in the shorter segment up to a possible cut indicated on the x-axis. The proteins, where the shorter segment contains 1, 2, or 3 α-helixes, β-sheets, or combinations of both, represent 80% of proteins from various organisms and 84% of human proteins. Source data are provided as a Source Data file.
Extended Data Fig. 3 Design of DHFR-based INSPIRE system.
a, Left: crystal structure of dihydrofolate reductase (DHFR) in complex with methotrexate (MTX) (PDB ID 1u72). Splits are chosen in surface-exposed loops (orange); α-helixes and β-strands are colored in dark and light gray, respectively. Right: aa sequence of DHFR with additional annotations indicating split site locations and secondary structures. b, Expression of N- (left) and C-terminal (right) split DHFR fragments in HEK293T cells verified with Western blot. Constitutively expressed mCitrine fluorescent protein was used as transfection control, and an empty plasmid vector (pcDNA3) was used as a negative control. The mCitrine transfection control was run on a separate gel. The transfection mixtures are provided in Supplementary Table 4. Data are representative of two independent experiments (n = 1). Original uncropped images are provided in the Source Data file.
Extended Data Fig. 4 Design of GR2-based INSPIRE system.
a, Left: crystal structure of ligand-binding domain (LBD) of glucocorticoid receptor 2 (GR2) in complex with mometasone furoate (MOF) (PDB ID 4e2j). Splits are chosen in surface-exposed loops (magenta); α-helixes and β-strands are colored in dark and light gray, respectively. Right: aa sequence of GR2, with additional annotations indicating split site locations and secondary structures. b, Expression of N- (left) and C-terminal (right) split GR2 fragments in HEK293T cells verified with Western blot. Constitutively expressed mCitrine fluorescent protein was used as transfection control, and an empty plasmid vector (pcDNA3) was used as a negative control. The mCitrine transfection control was run on a separate gel. Transfection mixtures are provided in Supplementary Table 4. Data are representative of two independent experiments (n = 1). Original uncropped images are provided in the Source Data file.
Extended Data Fig. 5 INSPIRE-mediated activation of a reporter gene.
a, Schematic presentation of the transcriptional activator INSPIRE systems (TA-INSPIRE). Ligand-induced dimerization of split protein fragments results in the expression of firefly luciferase gene (fLuc) on a reporter plasmid with 10 binding sites for gRNA upstream of the minimal promoter (Pmin). b–g, Transcriptional activation of the fLuc reporter gene by employing LYN267- (b), DHFR173- (c), GR2175- (d), TRβ412- €, ERβ446- (f), and PPARγ429- (g) based TA-INSPIRE systems after stimulation of HEK293T cells with the indicated ligands. Results are shown as fold activation in reporter gene expression compared to non-stimulated mock cells transfected with reporter plasmid only. Detailed descriptions of the genetic components and transfection mixtures are provided in Supplementary Table 4. Plots shows the means ± s.d. of n = 8 biological replicates combined from two independent experiments. See Supplementary Table 8 for sample sizes (n) and P values. Source data are provided as a Source Data file.
Extended Data Fig. 6 INSPIRE-mediated activation of individual endogenous genes.
a,b, Activation of individual endogenous genes by employing GR2- (a) or ERβ- (b) based TA-INSPIRE systems. HEK293T cells were transfected with individual gRNAs targeting TTN, ASCL1, NEUROD1, and TUNAR endogenous genes and stimulated with indicated ligands. Results are shown as fold activation in reporter gene expression compared to non-stimulated mock cells transfected with an empty plasmid (pcDNA3.1). Detailed descriptions of the genetic components and transfection mixtures are provided in Supplementary Table 4. Plots shows the means ± s.d. of n ≤ 6 biological replicates combined from two independent experiments. Conditions were compared using a two-sided unpaired t-test with Welch correction (****P < 0.0001; ***P < 0.001; **P < 0.01; ns > 0.05). See Supplementary Table 8 for sample sizes (n) and P values. Source data are provided as a Source Data file.
Extended Data Fig. 7 Characterization of reversibility of TA-INSPIRE systems.
a–g, Reversibility of FKBP-FRB-based CID system (a) and LYN267- (b), DHFR173- (c), GR2175- (d), TRβ412- (e), ERβ446- (f), and PPARγ429- (g) based TA-INSPIRE systems after ligand-mediated induction of reporter gene expression. Cells were continuously stimulated with the indicated ligands for 72 h (ON conditions) or stimulated for 24 h and then washed and cultured without the ligand (ON–OFF conditions). Results are shown as fold activation in reporter gene expression compared to non-stimulated mock cells transfected with only reporter plasmid. Detailed descriptions of the genetic components and transfection mixtures are provided in Supplementary Table 4. Plots shows the means ± s.d. of n = 8 biological replicates combined from two independent experiments. See Supplementary Table 8 for sample sizes (n) and P values. Source data are provided as a Source Data file.
Extended Data Fig. 8 Orthogonality of inducible GR2- and ERβ-based INSPIRE components and construction of logic gates by orthogonal TA-INSPIRE.
a, Orthogonality of GR2- and ERβ-based INSPIRE components (nSplit and cSplit fragments) was tested in HEK293T cells by co-transfected individual split fragments fused to split luciferase. Orthogonality was assessed by measuring luciferase activity after cell stimulation with indicated synthetic ligands (MOF and 4-OHT) or natural ligands (COR and EST). b, Schematic presentation of the transcriptional activation system used in (c) to simultaneously modulate expression of two reporter genes (BFP and mCit) using orthogonal GR2- and ERβ-based TA-INSPIRE. pMin represents a minimal promoter, and mCit and BFP represent the mCitrine and TagBFP fluorescent protein genes, respectively. c, Conditional activation of reporter genes mCit and BFP after stimulation of the cells with either synthetic ligands MOF (1 µM) and 4-OHT (10 µM) (left) or natural ligands COR (100 µM) and EST (10 µM) (right), detected with fluorescent confocal microscopy. Scale bars represent 100 µm. Microscopic images are representative of two independent experiments (n = 1 biologically independent sample) and four separate observations within the same experiment. d, Construction of functional OR gate, AND gate, and switching between transcriptional activation and repression (ACT/REP) utilizing GR2- and ERβ-based TA-INSPIRE in combination with TALE DNA-binding domain (see schematics in Fig. 4b–d). Cells were stimulated with natural ligands COR (100 µM) and EST (10 µM) as indicated. Results are shown as fold activation in reporter gene expression compared to non-stimulated mock cells transfected with only reporter plasmid. Detailed descriptions of the genetic components and transfection mixtures are provided in Supplementary Table 4. Plots shows the means ± s.d. of n = 8 biological replicates combined from two independent experiments. See Supplementary Table 8 for sample sizes (n) and P values. Source data are provided as a Source Data file.
Extended Data Fig. 9 Application of GR2-INSPIRE system for ligand-mediated reconstitution of split TEV protease.
a, Schematic presentation of the split-tobacco etch virus protease (TEVp) reconstitution utilizing INSPIRE (nSplit and cSplit). Ligand-mediated reconstitution of split TEVp leads to the cleavage and activation of cyclic luciferase reporter (cycLuc). b, Ligand-mediated induction of split TEVp reconstitution utilizing INSPIRE systems after stimulation of HEK293T cells with COR (left) or MOF (right). Results are shown as fold activation in reporter gene expression compared to non-stimulated mock cells transfected with only reporter plasmid. Detailed descriptions of the genetic components and transfection mixtures are provided in Supplementary Table 4. Plots shows the means ± s.d. of n = 8 biological replicates combined from two independent experiments. See Supplementary Table 8 for sample sizes (n) and P values. Source data are provided as a Source Data file.
Extended Data Fig. 10 Evaluation of GR2-INSPIRE responsiveness to corticosterone (CORT) and prescreen of the effective gRNAs targeting the mouse 11β-HSD2 gene.
a, The dose-response curve of GR2175-based INSPIRE system for corticosterone (CORT), utilizing dual luciferase assay in HEK293T cells. b, Activation of the mouse 11β-HSD2 gene in Neuro2a cells transfected with dCas9-VPR and indicated gRNAs. The light shaded region indicates the gRNA with the highest fold activation. Results are shown as fold activation in endogenous gene expression compared to mock cells transfected with an empty plasmid (pcDNA3). Detailed descriptions of the genetic components and transfection mixtures are provided in Supplementary Table 4. Plots shows the means ± s.d. of n = 6 biological replicates combined from two independent experiments. Conditions were compared using a two-sided unpaired t-test with Welch correction (****P < 0.0001; **P < 0.01; ns > 0.05). See Supplementary Table 8 for sample sizes (n) and P values. Source data are provided as a Source Data file.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3, Tables 1–8 and Note.
Supplementary Data
Archive of plasmid maps.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rihtar, E., Lebar, T., Lainšček, D. et al. Chemically inducible split protein regulators for mammalian cells. Nat Chem Biol 19, 64–71 (2023). https://doi.org/10.1038/s41589-022-01136-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-022-01136-x
This article is cited by
-
Integrated compact regulators of protein activity enable control of signaling pathways and genome-editing in vivo
Cell Discovery (2024)
-
Irreversible light-activated SpyLigation mediates split-protein assembly in 4D
Nature Protocols (2024)
-
Target-dependent RNA polymerase as universal platform for gene expression control in response to intracellular molecules
Nature Communications (2023)
-
A split and inducible adenine base editor for precise in vivo base editing
Nature Communications (2023)