Abstract
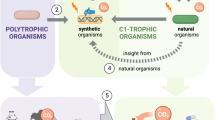
One-carbon (C1) substrates are preferred feedstocks for the biomanufacturing industry and have recently gained attention owing to their natural abundance, low production cost and availability as industrial by-products. However, native pathways to utilize these substrates are absent in most biotechnologically relevant microorganisms. Recent advances in synthetic biology, genome engineering and laboratory evolution are enabling the first steps towards the creation of synthetic C1-utilizing microorganisms. Here, we briefly review the native metabolism of methane, methanol, CO2, CO and formate, and how these C1-utilizing pathways can be engineered into heterologous hosts. In addition, this review analyses the potential, the challenges and the perspectives of C1-based biomanufacturing.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Zhou, Y. J., Kerkhoven, E. J. & Nielsen, J. Barriers and opportunities in bio-based production of hydrocarbons. Nat. Energy 3, 925–935 (2018).
Clomburg, J. M., Crumbley, A. M. & Gonzalez, R. Industrial biomanufacturing: the future of chemical production. Science 355, aag0804 (2017).
Naik, S. N., Goud, V. V., Rout, P. K. & Dalai, A. K. Production of first and second generation biofuels: a comprehensive review. Renew. Sust. Energ. 14, 578–597 (2010).
Strong, P. J., Xie, S. & Clarke, W. P. Methane as a resource: can the methanotrophs add value? Environ. Sci. Technol. 49, 4001–4018 (2015).
Pfeifenschneider, J., Brautaset, T. & Wendisch, V. F. Methanol as carbon substrate in the bio-economy: Metabolic engineering of aerobic methylotrophic bacteria for production of value-added chemicals. Biofuel. Bioprod. Biorefin. 11, 719–731 (2017).
Dürre, P. & Eikmanns, B. J. C1-carbon sources for chemical and fuel production by microbial gas fermentation. Curr. Opin. Biotechnol. 35, 63–72 (2015).
Whitaker, W. B., Sandoval, N. R., Bennett, R. K., Fast, A. G. & Papoutsakis, E. T. Synthetic methylotrophy: engineering the production of biofuels and chemicals based on the biology of aerobic methanol utilization. Curr. Opin. Biotechnol. 33, 165–175 (2015).
Yurimoto, H., Sakai, Y. & Kato, N. Hansenula polymorpha: Biology and Applications Ch. 5. (Wiley-Blackwell, Hoboken, 2002).
Wang, Y., Fan, L., Tuyishime, P., Zheng, P. & Sun, J. Synthetic methylotrophy: a practical solution for methanol-based biomanufacturing. Trends Biotechnol. 38, 650–666 (2020).
Zhang, W. et al. Current advance in bioconversion of methanol to chemicals. Biotechnol. Biofuels 11, 260 (2018).
Gassler, T. et al. The industrial yeast Pichia pastoris is converted from a heterotroph into an autotroph capable of growth on CO2. Nat. Biotechnol. 38, 210–216 (2019). This work created a fully synthetic autotrophic eukaryote (P. pastoris), able to produce all biomass carbon from CO2.
Gleizer, S. et al. Conversion of Escherichia coli to generate all biomass carbon from CO2. Cell 179, 1255–1263.e12 (2019). This work created a fully synthetic autotrophic prokaryote (E. coli), able to produce all biomass carbon from CO2.
Chen, F. Y.-H., Jung, H.-W., Tsuei, C.-Y. & Liao, J. C. Converting Escherichia coli to a synthetic methylotroph growing solely on methanol. Cell 182, 933–946. e14 (2020). In this work, E. coli was successfully converted into a fully synthetic methylotroph growing solely on methanol.
Kim, S. et al. Growth of E. coli on formate and methanol via the reductive glycine pathway. Nat. Chem. Biol. 16, 538–545 (2020). This work generated an E. coli strain able to grow solely on formate as a carbon source by constructing a synthetic reductive glycine pathway.
Methanol: 2018 World Market Outlook and Forecast up to 2027 (Merchant Research & Consulting, 2018); https://mcgroup.co.uk/researches/methanol
Du, X. L., Jiang, Z., Su, D. S. & Wang, J. Q. Research progress on the indirect hydrogenation of carbon dioxide to methanol. ChemSusChem 9, 322–332 (2016).
Bertau, M., Offermanns, H., Plass, L., Schmidt, F. & Wernicke, H.-J. Methanol: The Basic Chemical and Energy Feedstock of the Future (Springer, 2014).
Schrader, J. et al. Methanol-based industrial biotechnology: current status and future perspectives of methylotrophic bacteria. Trends Biotechnol. 27, 107–115 (2009).
Yurimoto, H., Oku, M. & Sakai, Y. Yeast methylotrophy: metabolism, gene regulation and peroxisome homeostasis. Int. J. Microbiol. 2011, 101298 (2011).
Houard, S., Heinderyckx, M. & Bollen, A. Engineering of non-conventional yeasts for efficient synthesis of macromolecules: the methylotrophic genera. Biochimie 84, 1089–1093 (2002).
Ledeboer, A. et al. Molecular cloning and characterization of a gene coding for methanol oxidase in Hansenula polymorpha. Nucleic Acids Res. 13, 3063–3082 (1985).
Cregg, J. M., Madden, K., Barringer, K., Thill, G. & Stillman, C. Functional characterization of the two alcohol oxidase genes from the yeast Pichia pastoris. Mol. Cell. Biol. 9, 1316–1323 (1989).
Sakai, Y. & Tani, Y. Cloning and sequencing of the alcohol oxidase-encoding gene (AOD1) from the formaldehyde-producing asporogeneous methylotrophic yeast, Candida boidinii S2. Gene 114, 67–73 (1992).
Yurimoto, H., Kato, N. & Sakai, Y. Assimilation, dissimilation, and detoxification of formaldehyde, a central metabolic intermediate of methylotrophic metabolism. Chem. Rec. 5, 367–375 (2005).
Rußmayer, H. et al. Systems-level organization of yeast methylotrophic lifestyle. BMC Biol. 13, 1–25 (2015).
Keltjens, J. T., Pol, A., Reimann, J. & den Camp, H. J. O. PQQ-dependent methanol dehydrogenases: rare-earth elements make a difference. Appl. Microbiol. Biotechnol. 98, 6163–6183 (2014).
Lee, J.-Y. et al. Discovery and biochemical characterization of a methanol dehydrogenase from Lysinibacillus xylanilyticus. Front. Bioeng. Biotechnol. 8, 67 (2020).
Orita, I., Sakamoto, N., Kato, N., Yurimoto, H. & Sakai, Y. Bifunctional enzyme fusion of 3-hexulose-6-phosphate synthase and 6-phospho-3-hexuloisomerase. Appl. Microbiol. Biotechnol. 76, 439–445 (2007).
Kallen, R. G. & Jencks, W. P. The mechanism of the condensation of formaldehyde with tetrahydrofolic acid. J. Biol. Chem. 241, 5851–5863 (1966).
Lindén, P., Keech, O., Stenlund, H., Gardeström, P. & Moritz, T. Reduced mitochondrial malate dehydrogenase activity has a strong effect on photorespiratory metabolism as revealed by 13C labelling. J. Exp. Bot. 67, 3123–3135 (2016).
Cotton, C. A., Claassens, N. J., Benito-Vaquerizo, S. & Bar-Even, A. Renewable methanol and formate as microbial feedstocks. Curr. Opin. Biotechnol. 62, 168–180 (2020).
Vorholt, J. A. Cofactor-dependent pathways of formaldehyde oxidation in methylotrophic bacteria. Arch. Microbiol. 178, 239–249 (2002).
Wang, X. et al. Biological conversion of methanol by evolved Escherichia coli carrying a linear methanol assimilation pathway. Bioresour. Bioprocess. 4, 41 (2017).
Whitaker, W. B. et al. Engineering the biological conversion of methanol to specialty chemicals in Escherichia coli. Metab. Eng. 39, 49–59 (2017).
Bennett, R. K., Gonzalez, J. E., Whitaker, W. B., Antoniewicz, M. R. & Papoutsakis, E. T. Expression of heterologous non-oxidative pentose phosphate pathway from Bacillus methanolicus and phosphoglucose isomerase deletion improves methanol assimilation and metabolite production by a synthetic Escherichia coli methylotroph. Metab. Eng. 45, 75–85 (2018).
De Simone, A. et al. Mixing and matching methylotrophic enzymes to design a novel methanol utilization pathway in E. coli. Metab. Eng. 61, 315–325 (2020).
Müller, J. E. et al. Engineering Escherichia coli for methanol conversion. Metab. Eng. 28, 190–201 (2015). The first report of the transplantation of the methanol assimilation pathway to the industrial host E. coli, which paved the way towards synthetic methylotrophic organisms.
Price, J. V., Chen, L., Whitaker, W. B., Papoutsakis, E. & Chen, W. Scaffoldless engineered enzyme assembly for enhanced methanol utilization. Proc. Natl Acad. Sci. USA 113, 12691–12696 (2016).
Woolston, B. M., King, J. R., Reiter, M., Van Hove, B. & Stephanopoulos, G. Improving formaldehyde consumption drives methanol assimilation in engineered E. coli. Nat. Commun. 9, 1–12 (2018). The authors of this work revealed that methanol assimilation is kinetically limited by methanol dehydrogenase, which provided the direction for future engineering strategies.
Meyer, F. et al. Methanol-essential growth of Escherichia coli. Nat. Commun. 9, 1508 (2018). In this work, methanol utilization by E. coli was achieved using a synthetic pathway and by coupling co-consumption of methanol to growth, which was a first step to complete synthetic methylotrophic E. coli.
Chen, C.-T. et al. Synthetic methanol auxotrophy of Escherichia coli for methanol-dependent growth and production. Metab. Eng. 49, 257–266 (2018).
Portnoy, V. A., Bezdan, D. & Zengler, K. Adaptive laboratory evolution—harnessing the power of biology for metabolic engineering. Curr. Opin. Biotechnol. 22, 590–594 (2011).
Tuyishime, P. et al. Engineering Corynebacterium glutamicum for methanol-dependent growth and glutamate production. Metab. Eng. 49, 220–231 (2018).
Espinosa, M.I. et al. Engineering and evolution of methanol assimilation in Saccharomyces cerevisiae. Preprint at bioRxiv https://doi.org/10.1101/717942v2 (2020).
Espinosa, M. I. et al. Adaptive laboratory evolution of native methanol assimilation in Saccharomyces cerevisiae. Nat. Commun. 11, 5564 (2020).
Hwang, I. Y. et al. Biocatalytic conversion of methane to methanol as a key step for development of methane-based biorefineries. J. Microbiol. Biotechnol. 24, 1597–1605 (2014).
Hanson, R. S. & Hanson, T. E. Methanotrophic bacteria. Microbiol. Mol. Biol. Rev. 60, 439–471 (1996).
Semrau, J. D., DiSpirito, A. A. & Yoon, S. Methanotrophs and copper. FEMS Microbiol. Rev. 34, 496–531 (2010).
de la Torre, A. et al. Genome-scale metabolic reconstructions and theoretical investigation of methane conversion in Methylomicrobium buryatense strain 5G (B1). Microb. Cell Fact. 14, 188 (2015).
Zilly, F. E. et al. Tuning a P450 enzyme for methane oxidation. Angew. Chem. Int. Ed. 50, 2720–2724 (2011).
Meinhold, P., Peters, M. W., Chen, M. M., Takahashi, K. & Arnold, F. H. Direct conversion of ethane to ethanol by engineered cytochrome P450 BM3. ChemBioChem 6, 1765–1768 (2005).
Balasubramanian, R. et al. Oxidation of methane by a biological dicopper centre. Nature 465, 115–119 (2010).
Kim, H. J. et al. Biological conversion of methane to methanol through genetic reassembly of native catalytic domains. Nat. Catal. 2, 342–353 (2019). In this work, the catalytic domains of a methane monooxygenases were assembled on apoferritin, resulting in a stable and soluble enzyme expression in the non- methanotrophic host E. coli.
Liew, F. et al. Gas fermentation—a flexible platform for commercial scale production of low-carbon-fuels and chemicals from waste and renewable feedstocks. Front. Microbiol. 7, 694 (2016).
Ernst, A. & Zibrak, J. D. Carbon monoxide poisoning. N. Engl. J. Med. 339, 1603–1608 (1998).
Alonso, J. R., Cardellach, F., López, S., Casademont, J. & Miró, Ò. Carbon monoxide specifically inhibits cytochrome c oxidase of human mitochondrial respiratory chain. Pharmacol. Toxicol. 93, 142–146 (2003).
Meyer, O. & Schlegel, H. G. Biology of aerobic carbon monoxide-oxidizing bacteria. Annu. Rev. Microbiol. 37, 277–310 (1983).
Oelgeschläger, E. & Rother, M. Carbon monoxide-dependent energy metabolism in anaerobic bacteria and archaea. Arch. Microbiol. 190, 257–269 (2008).
King, G. M. & Weber, C. F. Distribution, diversity and ecology of aerobic CO-oxidizing bacteria. Nat. Rev. Microbiol. 5, 107–118 (2007).
Diender, M., Stams, A. J. & Sousa, D. Z. Pathways and bioenergetics of anaerobic carbon monoxide fermentation. Front. Microbiol. 6, 1275 (2015).
Can, M., Armstrong, F. A. & Ragsdale, S. W. Structure, function, and mechanism of the nickel metalloenzymes, CO dehydrogenase, and acetyl-CoA synthase. Chem. Rev. 114, 4149–4174 (2014).
Ragsdale, S. W. & Pierce, E. Acetogenesis and the Wood–Ljungdahl pathway of CO2 fixation. Biochim. Biophys. Acta 1784, 1873–1898 (2008).
Roberts, D. L. et al. Cloning and expression of the gene cluster encoding key proteins involved in acetyl-CoA synthesis in Clostridium thermoaceticum: CO dehydrogenase, the corrinoid/Fe-S protein, and methyltransferase. Proc. Natl Acad. Sci. USA 86, 32–36 (1989).
Fast, A. G. & Papoutsakis, E. T. Functional expression of the Clostridium ljungdahlii acetyl-coenzyme A synthase in Clostridium acetobutylicum as demonstrated by a novel in vivo CO exchange activity en route to heterologous installation of a functional Wood-Ljungdahl pathway. Appl. Environ. Microbiol. 84, 7 (2018).
Carlson, E. D. & Papoutsakis, E. T. Heterologous expression of the clostridium carboxidivorans CO dehydrogenase alone or together with the acetyl coenzyme a synthase enables both reduction of CO2 and oxidation of CO by clostridium acetobutylicum. Appl. Environ. Microbiol. 83, 16 (2017).
Takors, R. et al. Using gas mixtures of CO, CO2 and H2 as microbial substrates: the do’s and don’ts of successful technology transfer from laboratory to production scale. Microb. Biotechnol. 11, 606–625 (2018).
Fuchs, G. Alternative pathways of carbon dioxide fixation: insights into the early evolution of life? Annu. Rev. Microbiol. 65, 631–658 (2011).
Ducat, D. C. & Silver, P. A. Improving carbon fixation pathways. Curr. Opin. Chem. Biol. 16, 337–344 (2012).
Sánchez-Andrea, I. et al. The reductive glycine pathway allows autotrophic growth of Desulfovibrio desulfuricans. Nat. Commun. 11, 5090 (2020).
Schwander, T., von Borzyskowski, L. S., Burgener, S., Cortina, N. S. & Erb, T. J. A synthetic pathway for the fixation of carbon dioxide in vitro. Science 354, 900–904 (2016).
Gong, F., Cai, Z. & Li, Y. Synthetic biology for CO2 fixation. Sci. China Life Sci. 59, 1106–1114 (2016).
Raines, C. A. The Calvin cycle revisited. Photosynth. Res. 75, 1–10 (2003).
Claassens, N. J. A warm welcome for alternative CO2 fixation pathways in microbial biotechnology. Microb. Biotechnol. 10, 31 (2017).
Andersson, I. & Backlund, A. Structure and function of Rubisco. Plant Physiol. Biochem. 46, 275–291 (2008).
Bar-Even, A., Noor, E., Lewis, N. E. & Milo, R. Design and analysis of synthetic carbon fixation pathways. Proc. Natl Acad. Sci. USA 107, 8889–8894 (2010).
Erb, T. J. & Zarzycki, J. Biochemical and synthetic biology approaches to improve photosynthetic CO2-fixation. Curr. Opin. Chem. Biol. 34, 72–79 (2016).
Davidi, D. et al. Highly active rubiscos discovered by systematic interrogation of natural sequence diversity. EMBO J. 39, e104081 (2020).
Gong, F., Zhu, H., Zhang, Y. & Li, Y. Biological carbon fixation: from natural to synthetic. J. CO2 Util. 28, 221–227 (2018).
Antonovsky, N. et al. Sugar synthesis from CO2 in Escherichia coli. Cell 166, 115–125 (2016). This work achieved a fully functional carbon fixation cycle in E. coli, capable of synthetizing sugars solely form CO2.
von Borzyskowski, L. S. et al. An engineered Calvin-Benson-Bassham cycle for carbon dioxide fixation in Methylobacterium extorquens AM1. Metab. Eng. 47, 423–433 (2018).
Flamholz, A. I. et al. Functional reconstitution of a bacterial CO2 concentrating mechanism in Escherichia coli. eLife 9, e59882 (2020).
Liu, Z., Wang, K., Chen, Y., Tan, T. & Nielsen, J. Third-generation biorefineries as the means to produce fuels and chemicals from CO2. Nat. Catal. 3, 274–288 (2020).
Guo, L. et al. Enhancement of malate production through engineering of the periplasmic rTCA pathway in Escherichia coli. Biotechnol. Bioeng. 115, 1571–1580 (2018).
Keller, M. W. et al. Exploiting microbial hyperthermophilicity to produce an industrial chemical, using hydrogen and carbon dioxide. Proc. Natl Acad. Sci. USA 110, 5840–5845 (2013).
Liu, Z. & Liu, T. Production of acrylic acid and propionic acid by constructing a portion of the 3-hydroxypropionate/4-hydroxybutyrate cycle from Metallosphaera sedula in Escherichia coli. J. Ind. Microbiol. 43, 1659–1670 (2016).
d Mattozzi, M., Ziesack, M., Voges, M. J., Silver, P. A. & Way, J. C. Expression of the sub-pathways of the Chloroflexus aurantiacus 3-hydroxypropionate carbon fixation bicycle in E. coli: toward horizontal transfer of autotrophic growth. Metab. Eng. 16, 130–139 (2013).
Tashiro, Y., Hirano, S., Matson, M. M., Atsumi, S. & Kondo, A. Electrical-biological hybrid system for CO2 reduction. Metab. Eng. 47, 211–218 (2018).
Yishai, O., Bouzon, M., Doring, V. & Bar-Even, A. In vivo assimilation of one-carbon via a synthetic reductive glycine pathway in Escherichia coli. ACS Synth. Biol. 7, 2023–2028 (2018).
Gonzalez de la Cruz, J., Machens, F., Messerschmidt, K. & Bar-Even, A. Core catalysis of the reductive glycine pathway demonstrated in yeast. ACS Synth. Biol. 8, 911–917 (2019).
Claassens, N. J. et al. Replacing the Calvin cycle with the reductive glycine pathway in Cupriavidus necator. Metab. Eng. 62, 30–41 (2020).
Satanowski, A. et al. Awakening a latent carbon fixation cycle in Escherichia coli. Nat. Commun. 11, 5812–5812 (2020).
Yishai, O., Lindner, S. N., Gonzalez de la Cruz, J., Tenenboim, H. & Bar-Even, A. The formate bio-economy. Curr. Opin. Chem. Biol. 35, 1–9 (2016).
Mao, W. et al. Recent progress in metabolic engineering of microbial formate assimilation. Appl. Microbiol. Biotechnol. 104, 6905–6917 (2020).
Bar-Even, A. Formate assimilation: the metabolic architecture of natural and synthetic pathways. Biochemistry 55, 3851–3863 (2016).
Siegel, J. B. et al. Computational protein design enables a novel one-carbon assimilation pathway. Proc. Natl Acad. Sci. USA 112, 3704–3709 (2015).
Bang, J. & Lee, S. Y. Assimilation of formic acid and CO2 by engineered Escherichia coli equipped with reconstructed one-carbon assimilation pathways. Proc. Natl Acad. Sci. USA 115, E9271–E9279 (2018).
Bang, J., Hwang, C. H., Ahn, J. H., Lee, J. A. & Lee, S. Y. Escherichia coli is engineered to grow on CO2 and formic acid. Nat. Microbiol. 5, 1459–1463 (2020). This work created an E. coli strain able to grow on CO2 and formic acid as sole carbon sources at improved cell densities.
Sandberg, T. E., Salazar, M. J., Weng, L. L., Palsson, B. O. & Feist, A. M. The emergence of adaptive laboratory evolution as an efficient tool for biological discovery and industrial biotechnology. Metab. Eng. 56, 1–16 (2019).
McCarty, N. S. & Ledesma-Amaro, R. Synthetic biology tools to engineer microbial communities for biotechnology. Trends Biotechnol. 37, 181–197 (2019).
Chistoserdova, L. Applications of methylotrophs: can single carbon be harnessed for biotechnology? Curr. Opin. Biotechnol. 50, 189–194 (2018).
Acknowledgements
W.J. is supported by Monash University under a Monash Graduate Scholarship (MGS), a Monash International Tuition Scholarship (MITS), and a Graduate Research International Travel Award (GRITA). R.L.-A. and H.P. received funding from the Biotechnology and Biological Sciences Research Council (BBSRC; BB/R01602X/1). R.L.-A. received funding from UK research and Innovation (19-ERACoBioTech- 33 SyCoLim BB/T011408/1), the BBSRC (BB/T013176/1), the British Council 527429894, and the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (DEUSBIO - 949080). D.H-V. is supported by Erasmus+ (E MADRID03 – UK LONDON015). R.L.-A.: Newton Advanced Fellowship (NAF\R1\201187). In addition, the authors thank A. Graham for improving the figures.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Chemical Biology thanks Benjamin Woolston, Yongjin Zhou and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Table 1
Rights and permissions
About this article
Cite this article
Jiang, W., Hernández Villamor, D., Peng, H. et al. Metabolic engineering strategies to enable microbial utilization of C1 feedstocks. Nat Chem Biol 17, 845–855 (2021). https://doi.org/10.1038/s41589-021-00836-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00836-0
This article is cited by
-
Increasing lysine level improved methanol assimilation toward butyric acid production in Butyribacterium methylotrophicum
Biotechnology for Biofuels and Bioproducts (2023)
-
Thermophilic Moorella thermoacetica as a platform microorganism for C1 gas utilization: physiology, engineering, and applications
Bioresources and Bioprocessing (2023)
-
The potential of CO2-based production cycles in biotechnology to fight the climate crisis
Nature Communications (2023)
-
Synergistic investigation of natural and synthetic C1-trophic microorganisms to foster a circular carbon economy
Nature Communications (2023)
-
Methanol-fuelled yeast synthesizes anticancer drugs
Nature Synthesis (2023)