Abstract
Caenorhabditis elegans serves as a model for understanding adiposity and its connections to aging. Current methodologies do not distinguish between fats serving the energy needs of the parent, akin to mammalian adiposity, from those that are distributed to the progeny, making it difficult to accurately interpret the physiological implications of fat content changes induced by external perturbations. Using spectroscopic coherent Raman imaging, we determine the protein content, chemical profiles and dynamics of lipid particles in live animals. We find fat particles in the adult intestine to be diverse, with most destined for the developing progeny. In contrast, the skin-like epidermis contains fats that are the least heterogeneous, the least dynamic and have high triglyceride content. These attributes are most consistent with stored somatic energy reservoirs. These results challenge the prevailing practice of assessing C. elegans adiposity by measurements that are dominated by the intestinal fat content.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The C. elegans accession codes (strains, developmental stages and sex) include the following: wild-type (Bristol N2), from L3 to gravid adult hermaphrodites; DH1390 rme-2(b1008), adult hermaphrodites; LIU1 ldrls1 [dhs-3p::dhs-3::GFP + unc-76(+)], adult hermaphrodites; CB4088 him-5(e1490), adult males; and a transgenic strain bls1(vit-2p::vit-2::GFP) expressing a fusion of YP170, adult hermaphrodites. Primary datasets generated and analyzed in this study consist of spectral image data. Each data file is in HDF5 format, and is ~5 GB in size. Approximately 50 such files were generated during the study. These data are available from the corresponding author on reasonable request. Intermediate data, such as spectral peak position and amplitude data extracted from spectral images after watershed, are also available upon request.
Code availability
Primary analysis of the coherent Raman spectra was performed using custom-built software that is publicly available and can be found at https://github.com/CCampJr/crikit2. Routine image analysis, such as watershed, was performed using Fiji ImageJ’s watershed method (v 1.52p, the latest version can be downloaded at https://imagej.net/Fiji). Statistical analysis was performed using NbClust (v 3.0), fitdistrplus (v 1.0–14) and ggplot2 (v 3.2.1) under R programming v 3.5.1, and SciKit-Learn (v 0.18.1) under Python v 3.4.
References
Delhaye, M. & Dhamelincourt, P. Raman microprobe and microscope with laser excitation. J. Raman Spectrosc. 3, 33–43 (1975).
Shipp, D. W., Sinjab, F. & Notingher, I. Raman spectroscopy: techniques and applications in the life sciences. Adv. Opt. Photon. 9, 315–428 (2017).
Camp, C. H. Jr & Cicerone, M. T. Chemically sensitive bioimaging with coherent Raman scattering. Nat. Photon. 9, 295–305 (2015).
Wang, P. et al. Imaging lipid metabolism in live Caenorhabditis elegans using fingerprint vibrations. Angew. Chem. Int. Ed. 53, 11787–11792 (2014).
Fu, D. et al. In vivo metabolic fingerprinting of neutral lipids with hyperspectral stimulated Raman scattering microscopy. J. Am. Chem. Soc. 136, 8820–8828 (2014).
Shi, L. et al. Optical imaging of metabolic dynamics in animals. Nat. Commun. 9, 2995 (2018).
Camp, C. H. Jr et al. High-speed coherent Raman fingerprint imaging of biological tissues. Nat. Photon. 8, 627–634 (2014).
Liu, Y., Lee, Y. J. & Cicerone, M. T. Broadband CARS spectral phase retrieval using a time-domain Kramers–Kronig transform. Opt. Lett. 34, 1363–1365 (2009).
Freudiger, C. W. et al. Label-free biomedical imaging with high sensitivity by stimulated Raman scattering microscopy. Science 322, 1857–1861 (2008).
Camp, C. H. Jr, Lee, Y. J. & Cicerone, M. T. Quantitative, comparable coherent anti-Stokes Raman scattering (CARS) spectroscopy: correcting errors in phase retrieval. J. Raman Spectrosc. 47, 408–415 (2016).
Hellerer, T. et al. Monitoring of lipid storage in Caenorhabditis elegans using coherent anti-Stokes Raman scattering (CARS) microscopy. Proc. Natl Acad. Sci. USA 104, 14658–14663 (2007).
Yen, K. et al. A comparative study of fat storage quantitation in nematode Caenorhabditis elegans using label and label-free methods. PLoS ONE 5, e12810 (2010).
Le, T. T., Duren, H. M., Slipchenko, M. N., Hu, C.-D. & Cheng, J.-X. Label-free quantitative analysis of lipid metabolism in living Caenorhabditis elegans. J. Lipid Res. 51, 672–677 (2010).
Klapper, M. et al. Fluorescence based fixative and vital staining of lipid droplets in C. elegans reveal fat stores using microscopic and flow cytometry approaches. J. Lipid Res. 52, 1281–1293 (2011).
Wang, M. C., Min, W., Freudiger, C. W., Ruvkun, G. & Xie, X. S. RNAi screening for fat regulatory genes with SRS microscopy. Nat. Methods 8, 135–138 (2011).
Li, X. et al. Quantitative imaging of lipid synthesis and lipolysis dynamics in Caenorhabditis elegans by stimulated Raman scattering microscopy. Anal. Chem. 91, 2279–2287 (2019).
Chen, W.-W. et al. Specific polyunsaturated fatty acids modulate lipid delivery and oocyte development in C. elegans revealed by molecular-selective label-free imaging. Sci. Rep. 6, 32021 (2016).
Srinivasan, S. Regulation of body fat in Caenorhabditis elegans. Annu. Rev. Physiol. 77, 161–178 (2015).
Watts, J. L. & Ristow, M. Lipid and carbohydrate metabolism in Caenorhabditis elegans. Genetics 207, 413–446 (2017).
Watts, J. L. Fat synthesis and adiposity regulation in Caenorhabditis elegans. Trends Endocrinol. Metab. 20, 58–65 (2009).
Soukas, A. A., Kane, E. A., Carr, C. E., Melo, J. A. & Ruvkun, G. Rictor/TORC2 regulates fat metabolism, feeding, growth and life span in Caenorhabditis elegans. Genes Dev. 23, 496–511 (2009).
O’Rourke, E. J., Soukas, A. A., Carr, C. E. & Ruvkun, G. C. elegans major fats are stored in vesicles distinct from lysosome-related organelles. Cell Metab. 10, 430–435 (2009).
Perez, M. F. & Lehner, B. Vitellogenins—yolk gene function and regulation in Caenorhabditis elegans. Front. Physiol. 10, 1067 (2019).
Fukumoto, S. & Fujimoto, T. Deformation of lipid droplets in fixed samples. Histochem. Cell Biol. 118, 423–428 (2002).
Lemieux, G. A. & Ashrafi, K. Insights and challenges in using C. elegans for investigation of fat metabolism. Crit. Rev. Biochem. Mol. Biol. 50, 69–84 (2015).
Vincent, L. & Soille, P. Watersheds in digital spaces: an efficient algorithm based on immersion simulations. IEEE Trans. Pattern Anal. Mach. Intell. 13, 583–598 (1991).
Hubbard, E. J. & Greenstein, D. Introduction to the Germ Line (WormBook, 2005); https://doi.org/10.1895/wormbook.1.18.1.
Dowen, R. H., Breen, P. C., Tullius, T., Conery, A. L. & Ruvkun, G. A microRNA program in the C. elegans hypodermis couples to intestinal mTORC2/PQM-1 signaling to modulate fat transport. Genes Dev. 30, 1515–1528 (2016).
Grant, B. & Hirsh, D. Receptor-mediated endocytosis in the Caenorhabditis elegans oocyte. Mol. Biol. Cell 10, 4311–4326 (1999).
Kimble, J. & Sharrock, W. J. Tissue-specific synthesis of yolk proteins in Caenorhabditis elegans. Dev. Biol. 96, 189–196 (1983).
Zhang, P. et al. Proteomic study and marker protein identification of Caenorhabditis elegans lipid droplets. Mol. Cell. Proteom. 11, 317–328 (2012).
Schafer, W. Egg-laying (WormBook 2006); https://doi.org/10.1895/wormbook.1.38.1.
Czamara, K. et al. Raman spectroscopy of lipids: a review. J. Raman Spectrosc. 46, 4–20 (2015).
Kubagawa, H. M. et al. Oocyte signals derived from polyunsaturated fatty acids control sperm recruitment in vivo. Nat. Cell Biol. 8, 1143–1148 (2006).
Vrablik, T. L., Petyuk, V. A., Larson, E. M., Smith, R. D. & Watts, J. L. Lipidomic and proteomic analysis of Caenorhabditis elegans lipid droplets and identification of ACS-4 as a lipid droplet-associated protein. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1851, 1337–1345 (2015).
Kurzchalia, T. V. & Ward, S. Why do worms need cholesterol? Nat. Cell Biol. 5, 684–688 (2003).
Charrad, M., Ghazzali, N., Boiteau, V. & Niknafs, A. NbClust: an R package for determining the relevant number of clusters in a data set. J. Stat. Softw. 61, 1–36 (2014).
Nascimento, J. M. P. & Dias, J. M. B. Vertex component analysis: a fast algorithm to unmix hyperspectral data. IEEE Trans. Geosci. Remote Sens. 43, 898–910 (2005).
Sakamoto, K., Göransson, O., Hardie, D. G. & Alessi, D. R. Activity of LKB1 and AMPK-related kinases in skeletal muscle: effects of contraction, phenformin and AICAR. Am. J. Physiol. Endocrinol. Metab. 287, E310–E317 (2004).
McKay, R. M., McKay, J. P., Avery, L. & Graff, J. M. C. elegans: a model for exploring the genetics of fat storage. Dev. Cell 4, 131–142 (2003).
O’Rourke, E. J. & Ruvkun, G. MXL-3 and HLH-30 transcriptionally link lipolysis and autophagy to nutrient availability. Nat. Cell Biol. 15, 668–676 (2013).
Lee, J. H. et al. Lipid droplet protein LID-1 mediates ATGL-1-dependent lipolysis during fasting in Caenorhabditis elegans. Mol. Cell. Biol. 34, 4165–4176 (2014).
Lee, Y. J., Moon, D., Migler, K. B. & Cicerone, M. T. Quantitative image analysis of broadband CARS hyperspectral images of polymer blends. Anal. Chem. 83, 2733–2739 (2011).
Hollon, T. C. et al. Near real-time intraoperative brain tumor diagnosis using stimulated Raman histology and deep neural networks. Nat. Med. 26, 52–58 (2020).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning. Data Mining, Inference and Prediction 2nd edn (Springer, 2009).
Acknowledgements
K.A. and G.A.L. acknowledge support from NIH/NIA (R01AG046400) and BWF Innovations in Regulatory Sciences. T.C. acknowledges support from MOST-106-2119-M-001-030-MY3 of the Republic of China (Taiwan).
Author information
Authors and Affiliations
Contributions
K.A., T.-C.C., M.T.C. and W.-W.C. initiated the project. G.A.L., K.A. and W.-W.C. conceived and designed the study. W.-W.C. conducted all experiments. C.H.C. and W.-W.C. developed the ensemble machine-learning workflow. W.-W.C. analyzed all data with discussions and contributions from G.A.L., K.A., M.T.C., C.H.C. and T.-C.C. G.A.L. and K.A. conceived and drew Fig. 6. W.-W.C., G.A.L., K.A. and M.T.C. wrote the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The analysis results after applying ensemble machine-learning method for fed and short-term starved (2–3 hr) adult hermaphrodites as well as adult male (him-5) worms.
a, The analysis results of gonad (oocytes) and skin-like epidermis near pharynx in the fed wild-type adult hermaphrodites. b, The analysis results of the intestine in the fed / short-term starved adult hermaphrodite and male (him-5) worms. c, The analysis results of gonad and epidermis in the short-term starved adult hermaphrodites. Scale bar =10 μm for (a) to (c). The experiments were repeated at least four times independently with similar results for (a) and (b), and were repeated two times independently with similar results for (c). The region of pharyngeal neurons was excluded for the analysis of skin-like epidermis near pharynx.
Extended Data Fig. 2 The effect of long-term starvation and phenformin.
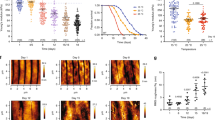
a, The CARS images of control, long-term starved (18–20 hr), and phenformin-treated worms. The young adult worms that had been fed dFA since the late L4 stage (12 h) were placed on the OP50 lawns that lacked the deuterium label (for control), on the plates lacked OP50 (for 18–20 h long-term starvation), and on the OP50 lawns that lacked the deuterium label but contained phenformin (20 h treatment with final concentration = 7.5 mM), respectively. After that, the worms were imaged by BCARS. The right column is the results after applied the ensemble machine-learning method. Scale bar =10μm. The experiments were repeated at least three times independently with similar results. The region of pharyngeal neurons was excluded for the analysis of skin-like epidermis near pharynx. b, The normalized lipid content (or normalized mean 2845 cm-1 intensity) of total lipid-rich particles in the tissue (normalized to a fixed tissue area). c, The average number of the dFA-retaining observed in the tissue (normalized to a fixed tissue area). Total ~4000 particles were analyzed; the data were collected from n = 3–6 biologically independent animals for each condition, where each measurement is presented as a dot; The error bars represent the standard error of the mean.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10.
Rights and permissions
About this article
Cite this article
Chen, WW., Lemieux, G.A., Camp, C.H. et al. Spectroscopic coherent Raman imaging of Caenorhabditis elegans reveals lipid particle diversity. Nat Chem Biol 16, 1087–1095 (2020). https://doi.org/10.1038/s41589-020-0565-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0565-2
This article is cited by
-
Computational coherent Raman scattering imaging: breaking physical barriers by fusion of advanced instrumentation and data science
eLight (2023)
-
Lipid droplets and peroxisomes are co-regulated to drive lifespan extension in response to mono-unsaturated fatty acids
Nature Cell Biology (2023)
-
Raman microscopy-based quantification of the physical properties of intracellular lipids
Communications Biology (2021)
-
Toward spatially resolved metabolomics
Nature Chemical Biology (2020)