Abstract
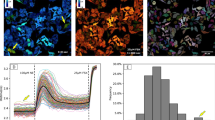
Here we generate fluorescence resonance energy transfer biosensors for guanine exchange factors (GEFs) by inserting a fluorescent protein pair in a structural ‘hinge’ common to many GEFs. Fluorescent biosensors can map the activation of signaling molecules in space and time, but it has not been possible to quantify how different activation events affect one another or contribute to a specific cell behavior. By imaging the GEF biosensors in the same cells as red-shifted biosensors of Rho GTPases, we can apply partial correlation analysis to parse out the extent to which each GEF contributes to the activation of a specific GTPase in regulating cell movement. Through analysis of spontaneous cell protrusion events, we identify when and where the GEF Asef regulates the GTPases Cdc42 and Rac1 to control cell edge dynamics. This approach exemplifies a powerful means to elucidate the real-time connectivity of signal transduction networks.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Code availability
All code used for the data analysis was written in MATLAB 2014b. All the code can be downloaded from the Danuser Lab github: https://github.com/DanuserLab under Biosensor, Windowing-Protrusion and Time-Series-Tools repositories.
References
Devreotes, P. & Horwitz, A. R. Signaling networks that regulate cell migration. Cold Spring Harb. Perspect. Biol. 7, a005959 (2015).
Komatsu, N. et al. Development of an optimized backbone of FRET biosensors for kinases and GTPases. Mol. Biol. Cell 22, 4647–4656 (2011).
Machacek, M. et al. Coordination of Rho GTPase activities during cell protrusion. Nature 461, 99–103 (2009).
Yang, H. W., Collins, S. R. & Meyer, T. Locally excitable Cdc42 signals steer cells during chemotaxis. Nat. Cell Biol. 18, 191–201 (2016).
Zawistowski, J. S., Sabouri-Ghomi, M., Danuser, G., Hahn, K. M. & Hodgson, L. A RhoC biosensor reveals differences in the activation kinetics of RhoA and RhoC in migrating cells. PLoS ONE 8, e79877 (2013).
Hodgson, L. et al. FRET binding antenna reports spatiotemporal dynamics of GDI-Cdc42 GTPase interactions. Nat. Chem. Biol. 12, 802–809 (2016).
Rossman, K. L., Der, C. J. & Sondek, J. GEF means go: turning on Rho GTPases with guanine nucleotide-exchange factors. Nat. Rev. Mol. Cell Biol. 6, 167–180 (2005).
Hall, A. Rho family GTPases. Biochem. Soc. Trans. 40, 1378–1382 (2012).
Mitin, N. et al. Release of autoinhibition of ASEF by APC leads to Cdc42 activation and tumor suppression. Nat. Struct. Mol. Biol. 14, 814–823 (2007).
Slattery, S. D. & Hahn, K. M. A high-content assay for biosensor validation and for examining stimuli that affect biosensor activity. Curr. Protoc. Cell Biol. 65, 11–31 (2014).
Rizzo, M. A., Springer, G. H., Granada, B. & Piston, D. W. An improved cyan fluorescent protein variant useful for FRET. Nat. Biotechnol. 22, 445–449 (2004).
Nagai, T. et al. A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat. Biotechnol. 20, 87–90 (2002).
Markwardt, M. L. et al. An improved cerulean fluorescent protein with enhanced brightness and reduced reversible photoswitching. PLoS ONE 6, e17896 (2011).
Xia, N. S. et al. Bioluminescence of Aequorea macrodactyla, a common jellyfish species in the East China Sea. Mar. Biotechnol. 4, 155–162 (2002).
Ai, H. W., Henderson, J. N., Remington, S. J. & Campbell, R. E. Directed evolution of a monomeric, bright and photostable version of Clavularia cyan fluorescent protein: structural characterization and applications in fluorescence imaging. Biochem. J. 400, 531–540 (2006).
Nguyen, A. W. & Daugherty, P. S. Evolutionary optimization of fluorescent proteins for intracellular FRET. Nat. Biotechnol. 23, 355–360 (2005).
Itoh, R. E. et al. Phosphorylation and activation of the Rac1 and Cdc42 GEF Asef in A431 cells stimulated by EGF. J. Cell Sci. 121, 2635–2642 (2008).
Yu, B. et al. Structural and energetic mechanisms of cooperative autoinhibition and activation of Vav1. Cell 140, 246–256 (2010).
Aghazadeh, B., Lowry, W. E., Huang, X. Y. & Rosen, M. K. Structural basis for relief of autoinhibition of the Dbl homology domain of proto-oncogene Vav by tyrosine phosphorylation. Cell 102, 625–633 (2000).
Crespo, P., Schuebel, K. E., Ostrom, A. A., Gutkind, J. S. & Bustelo, X. R. Phosphotyrosine-dependent activation of Rac-1 GDP/GTP exchange by the vav proto-oncogene product. Nature 385, 169–172 (1997).
Barreira, M. et al. The C-terminal SH3 domain contributes to the intramolecular inhibition of Vav family proteins. Sci. Signal 7, ra35 (2014).
Yohe, M. E. et al. Auto-inhibition of the Dbl family protein Tim by an N-terminal helical motif. J. Biol. Chem. 282, 13813–13823 (2007).
Yohe, M. E., Rossman, K. & Sondek, J. Role of the C-terminal SH3 domain and N-terminal tyrosine phosphorylation in regulation of Tim and related Dbl-family proteins. Biochemistry 47, 6827–6839 (2008).
Xu, Z., Gakhar, L., Bain, F. E., Spies, M. & Fuentes, E. J. The Tiam1 guanine nucleotide exchange factor is auto-inhibited by its pleckstrin homology coiled-coil extension domain. J. Biol. Chem. 292, 17777–17793 (2017).
Chen, Z., Guo, L., Sprang, S. R. & Sternweis, P. C. Modulation of a GEF switch: autoinhibition of the intrinsic guanine nucleotide exchange activity of p115-RhoGEF. Protein Sci. 20, 107–117 (2011).
Lambert, J. M. et al. Tiam1 mediates Ras activation of Rac by a PI(3)K-independent mechanism. Nat. Cell Biol. 4, 621–625 (2002).
Miyamoto, Y., Yamauchi, J., Tanoue, A., Wu, C. & Mobley, W. C. TrkB binds and tyrosine-phosphorylates Tiam1, leading to activation of Rac1 and induction of changes in cellular morphology. Proc. Natl Acad. Sci. USA 103, 10444–10449 (2006).
Servitja, J. M., Marinissen, M. J., Sodhi, A., Bustelo, X. R. & Gutkind, J. S. Rac1 function is required for Src-induced transformation. Evidence of a role for Tiam1 and Vav2 in Rac activation by Src. J. Biol. Chem. 278, 34339–34346 (2003).
Tolias, K. F. et al. The Rac1 guanine nucleotide exchange factor Tiam1 mediates EphB receptor-dependent dendritic spine development. Proc. Natl Acad. Sci. USA 104, 7265–7270 (2007).
Suzuki, N., Nakamura, S., Mano, H. & Kozasa, T. Galpha 12 activates Rho GTPase through tyrosine-phosphorylated leukemia-associated RhoGEF. Proc. Natl Acad. Sci. USA 100, 733–738 (2003).
Feng, Q. et al. Cool-1 functions as an essential regulatory node for EGF receptor- and Src-mediated cell growth. Nat. Cell Biol. 8, 945–956 (2006).
Jaiswal, M. et al. Mechanistic insights into specificity, activity, and regulatory elements of the regulator of G-protein signaling (RGS)-containing Rho-specific guanine nucleotide exchange factors (GEFs) p115, PDZ-RhoGEF (PRG), and leukemia-associated RhoGEF (LARG). J. Biol. Chem. 286, 18202–18212 (2011).
Nalbant, P., Hodgson, L., Kraynov, V., Toutchkine, A. & Hahn, K. M. Activation of endogenous Cdc42 visualized in living cells. Science 305, 1615–1619 (2004).
Herrington, K. A. et al. Spatial analysis of Cdc42 activity reveals a role for plasma membrane-associated Cdc42 in centrosome regulation. Mol. Biol. Cell 28, 2135–2145 (2017).
Mendoza, M. C., Vilela, M., Juarez, J. E., Blenis, J. & Danuser, G. ERK reinforces actin polymerization to power persistent edge protrusion during motility. Sci. Signal 8, ra47 (2015).
Lee, K. et al. Functional hierarchy of redundant actin assembly factors revealed by fine-grained registration of intrinsic image fluctuations. Cell Syst. 1, 37–50 (2015).
Ji, L., Lim, J. & Danuser, G. Fluctuations of intracellular forces during cell protrusion. Nat. Cell Biol. 10, 1393–1400 (2008).
Shcherbakova, D. M., Hink, M. A., Joosen, L., Gadella, T. W. & Verkhusha, V. V. An orange fluorescent protein with a large Stokes shift for single-excitation multicolor FCCS and FRET imaging. J. Am. Chem. Soc. 134, 7913–7923 (2012).
Shaner, N. C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat. Biotechnol. 22, 1567–1572 (2004).
Nishimura, T. et al. PAR-6-PAR-3 mediates Cdc42-induced Rac activation through the Rac GEFs STEF/Tiam1. Nat. Cell Biol. 7, 270–277 (2005).
Martin, K. et al. Spatio-temporal co-ordination of RhoA, Rac1 and Cdc42 activation during prototypical edge protrusion and retraction dynamics. Sci. Rep. 6, 21901 (2016).
Kuo, J. C., Han, X., Hsiao, C. T., Yates, J. R. 3rd & Waterman, C. M. Analysis of the myosin-II-responsive focal adhesion proteome reveals a role for beta-Pix in negative regulation of focal adhesion maturation. Nat. Cell Biol. 13, 383–393 (2011).
Goicoechea, S. M., Awadia, S. & Garcia-Mata, R. I’m coming to GEF you: regulation of RhoGEFs during cell migration. Cell Adh. Migr. 8, 535–549 (2014).
Lawson, C. D. & Ridley, A. J. Rho GTPase signaling complexes in cell migration and invasion. J. Cell Biol. 217, 447–457 (2018).
Azoitei, M. L. et al. Spatiotemporal dynamics of GEF-H1 activation controlled by microtubule- and Src-mediated pathways. J. Cell Biol. 218, 3077–3097 (2019).
Whitlow, M. et al. An improved linker for single-chain Fv with reduced aggregation and enhanced proteolytic stability. Protein Eng. 6, 989–995 (1993).
Kuhlman, B., Yang, H. Y., Boice, J. A., Fairman, R. & Raleigh, D. P. An exceptionally stable helix from the ribosomal protein L9: implications for protein folding and stability. J. Mol. Biol. 270, 640–647 (1997).
Kim, J.H. et al. High cleavage efficiency of a 2A peptide derived from porcine Teschovirus-1 in human cell lines, zebrafish and MICE. PLoS ONE 6, e18556 (2011).
Lindenburg, L. H., Hessels, A. M., Ebberink, E. H., Arts, R. & Merkx, M. Robust red FRET sensors using self-associating fluorescent domains. ACS Chem. Biol. 8, 2133–2139 (2013).
Fellmann, C. et al. An optimized microRNA backbone for effective single-copy RNAi. Cell Rep. 5, 1704–1713 (2013).
Shcherbakova, D. M. & Verkhusha, V. V. Near-infrared fluorescent proteins for multicolor in vivo imaging. Nat. Methods 10, 751–754 (2013).
Vilela, M. et al. Fluctuation analysis of activity biosensor images for the study of information flow in signaling pathways. Methods Enzymol. 519, 253–276 (2013).
Huang, N. E. et al. The empirical mode decomposition and the Hilbert spectrum for nonlinear and non-stationary time series analysis. Proc. R. Soc. A. 454, 903–995 (1998).
Zoubir, A. M. & Iskander, D. R. Bootstrap Techniques for Signal Processing (Cambridge Univ. Press, 2004).
Bailey, N. T. J. Statistical Methods in Biology 3rd edn (Cambridge Univ. Press, 1995).
Acknowledgements
Grant nos. R01-GM062299 (J.S.), R01 GM071868 (G.D.) and R35GM122596 (K.M.H.) supported this work. D.J.M. was supported by funding from the Leukemia and Lymphoma Society. J.H. was supported by a Human Frontiers in Sciences Program. G.G. was supported by the ‘Integrated Training in Cancer Model Systems’ Training Program, T32CA009156. We thank the Calabrese laboratory and F. Pimenta for the shRNA expression vector.
Author information
Authors and Affiliations
Contributions
D.J.M., J.S. and K.M.H. designed the biosensors. D.J.M. carried out all experiments, except for work by J.R. in building Tiam1, Tim, B-Pix and LARG biosensors. G.G. and M.A. contributed to initial design of Tim and LARG biosensors, respectively. M.V., J.H. and G.D. developed and carried out computational analysis. G.D., J.S. and K.M.H. provided intellectual input in all phases of the study and directed the work. The paper was written by D.J.M., M.V., G.D., J.S. and K.M.H., with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10.
Supplementary Video 1
Cdc42 (left) and Rac1 (right) activation reported by biosensors in MDA-MB-231 cells undergoing random edge motion.
Supplementary Video 2
Asef (left, middle) and Vav2 (right) activation reported by biosensors in MDA-MB-231 cells undergoing random edge motion.
Supplementary Video 3
Simultaneously imaged RhoGEF and Rho GTPase biosensors in MDA-MB-231 cells undergoing constitutive edge motion.
Supplementary Video 4
Simultaneously imaged Cdc42 and Rac1 biosensors in MDA-MB-231 cells undergoing constitutive edge motion.
Rights and permissions
About this article
Cite this article
Marston, D.J., Vilela, M., Huh, J. et al. Multiplexed GTPase and GEF biosensor imaging enables network connectivity analysis. Nat Chem Biol 16, 826–833 (2020). https://doi.org/10.1038/s41589-020-0542-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0542-9
This article is cited by
-
Pick-up single-cell proteomic analysis for quantifying up to 3000 proteins in a Mammalian cell
Nature Communications (2024)
-
COVID-19 Virus Structural Details: Optical and Electrochemical Detection
Journal of Fluorescence (2024)
-
Live imaging of apoptotic signaling flow using tunable combinatorial FRET-based bioprobes for cell population analysis of caspase cascades
Scientific Reports (2022)
-
A computational model of mutual antagonism in the mechano-signaling network of RhoA and nitric oxide
BMC Molecular and Cell Biology (2021)
-
Spatiotemporal development of coexisting wave domains of Rho activity in the cell cortex
Scientific Reports (2021)