Abstract
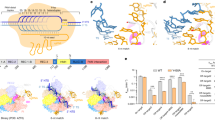
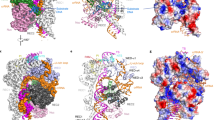
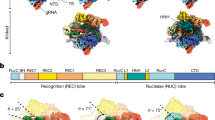
The RNA-programmable DNA-endonuclease Cas9 is widely used for genome engineering, where a high degree of specificity is required. To investigate which features of Cas9 determine the sensitivity to mismatches along the target DNA, we performed in vitro biochemical assays and bacterial survival assays in Escherichia coli. We demonstrate that arginines in the Cas9 bridge helix influence guide RNA, and target DNA binding and cleavage. They cluster in two groups that either increase or decrease the Cas9 sensitivity to mismatches. We show that the bridge helix is essential for R-loop formation and that R63 and R66 reduce Cas9 specificity by stabilizing the R-loop in the presence of mismatches. Additionally, we identify Q768 that reduces sensitivity of Cas9 to protospacer adjacent motif-distal mismatches. The Cas9_R63A/Q768A variant showed increased specificity in human cells. Our results provide a firm basis for function- and structure-guided mutagenesis to increase Cas9 specificity for genome engineering.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The datasets generated and analyzed during this study are available from the corresponding author upon request. Raw pictures for all in vitro experiments are available as Supplementary Note 1 online. Amplicon sequencing data are available online (accession code PRJNA577558).
References
Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9–crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl Acad. Sci. USA 109, E2579–E2586 (2012).
Jinek, M. et al. A programmable dual-RNA–Guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Deltcheva, E. et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471, 602 (2011).
Sternberg, S. H., Redding, S., Jinek, M., Greene, E. C. & Doudna, J. A. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature 507, 62 (2014).
Szczelkun, M. D. et al. Direct observation of R-loop formation by single RNA-guided Cas9 and cascade effector complexes. Proc. Natl Acad. Sci. USA 111, 9798–9803 (2014).
Adli, M. The CRISPR tool kit for genome editing and beyond. Nat. Commun. 9, 1911 (2018).
Doudna, J. A. & Charpentier, E. The new frontier of genome engineering with CRISPR-Cas9. Science 346, 1258096 (2014).
Komor, A. C., Badran, A. H. & Liu, D. R. CRISPR-based technologies for the manipulation of eukaryotic genomes. Cell 168, 20–36 (2017).
Wu, W. Y., Lebbink, J. H. G., Kanaar, R., Geijsen, N. & van der Oost, J. Genome editing by natural and engineered CRISPR-associated nucleases. Nat. Chem. Biol. 14, 642–651 (2018).
Chen, J. S. et al. Enhanced proofreading governs CRISPR–Cas9 targeting accuracy. Nature 550, 407 (2017).
Dagdas, Y. S., Chen, J. S., Sternberg, S. H., Doudna, J. A. & Yildiz, A. A conformational checkpoint between DNA binding and cleavage by CRISPR-Cas9. Sci. Adv. 3, eaao0027 (2017).
Tycko, J., Myer, VicE. & Hsu, PatrickD. Methods for optimizing CRISPR-Cas9 genome editing specificity. Mol. Cell 63, 355–370 (2016).
Casini, A. et al. A highly specific SpCas9 variant is identified by in vivo screening in yeast. Nat. Biotech. 36, 265 (2018).
Hu, J. H. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556, 57 (2018).
Kleinstiver, B. P. et al. High-fidelity CRISPR–Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490 (2016).
Kulcsár, P. I. et al. Crossing enhanced and high fidelity SpCas9 nucleases to optimize specificity and cleavage. Genome Biol. 18, 190 (2017).
Slaymaker, I. M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016).
Lee, J. K. et al. Directed evolution of CRISPR-Cas9 to increase its specificity. Nat. Commun. 9, 3048 (2018).
Vakulskas, C. A. et al. A high-fidelity Cas9 mutant delivered as a ribonucleoprotein complex enables efficient gene editing in human hematopoietic stem and progenitor cells. Nat. Med. 24, 1216–1224 (2018).
Singh, D. et al. Mechanisms of improved specificity of engineered Cas9s revealed by single-molecule FRET analysis. Nat. Struct. Mol. Biol. 25, 347–354 (2018).
Babu, K. et al. Bridge Helix of Cas9 modulates target DNA cleavage and mismatch tolerance. Biochemistry 58, 1905–1917 (2019).
Mekler, V., Minakhin, L. & Severinov, K. Mechanism of duplex DNA destabilization by RNA-guided Cas9 nuclease during target interrogation. Proc. Natl Acad. Sci. USA 114, 5443–5448 (2017).
Singh, D., Sternberg, S. H., Fei, J., Doudna, J. A. & Ha, T. Real-time observation of DNA recognition and rejection by the RNA-guided endonuclease Cas9. Nat. Commun. 7, 12778 (2016).
Kunne, T. et al. Role of nucleotide identity in effective CRISPR target escape mutations. Nucleic Acids Res. 46, 10395–10404 (2018).
Xue, C. et al. CRISPR interference and priming varies with individual spacer sequences. Nucleic Acids Res. 43, 10831–10847 (2015).
Cencic, R. et al. Protospacer adjacent motif (PAM)-distal sequences engage CRISPR Cas9 DNA target cleavage. PLoS One 9, e109213 (2014).
Sternberg, S. H., LaFrance, B., Kaplan, M. & Doudna, J. A. Conformational control of DNA target cleavage by CRISPR–Cas9. Nature 527, 110 (2015).
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014).
Anders, C., Niewoehner, O., Duerst, A. & Jinek, M. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 513, 569 (2014).
Jiang, F. et al. Structures of a CRISPR-Cas9 R-loop complex primed for DNA cleavage. Science 351, 867–871 (2016).
Jiang, F., Zhou, K., Ma, L., Gressel, S. & Doudna, J. A. A Cas9–guide RNA complex preorganized for target DNA recognition. Science 348, 1477–1481 (2015).
Zeng, Y. et al. The initiation, propagation and dynamics of CRISPR-SpyCas9 R-loop complex. Nucleic Acids Res. 46, 350–361 (2018).
Fu, Y., Sander, J. D., Reyon, D., Cascio, V. M. & Joung, J. K. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat. Biotech. 32, 279 (2014).
Datsenko, K. A. et al. Molecular memory of prior infections activates the CRISPR/Cas adaptive bacterial immunity system. Nat. Commun. 3, 945 (2012).
Fineran, P. C. et al. Degenerate target sites mediate rapid primed CRISPR adaptation. Proc. Natl Acad. Sci. USA 111, E1629–E1638 (2014).
Jinek, M. et al. Structures of Cas9 endonucleases reveal RNA-mediated conformational activation. Science 343, 1247997 (2014).
Bisaria, N., Jarmoskaite, I. & Herschlag, D. Lessons from enzyme kinetics reveal specificity principles for RNA-guided nucleases in RNA interference and CRISPR-based genome editing. Cell Syst. 4, 21–29 (2017).
Gong, S., Yu, H. H., Johnson, K. A. & Taylor, D. W. DNA Unwinding Is the Primary Determinant of CRISPR-Cas9 Activity. Cell Rep. 22, 359–371 (2018).
Maniatis, T., F Fritsch, E. & Sambrook, J. Molecular Cloning: A Laboratory Manual (Wiley, 2018).
Kirsch, R. D. & Joly, E. An improved PCR-mutagenesis strategy for two-site mutagenesis or sequence swapping between related genes. Nucleic Acids Res. 26, 1848–1850 (1998).
Langley, K. E., Villarejo, M. R., Fowler, A. V., Zamenhof, P. J. & Zabin, I. Molecular basis of beta-galactosidase alpha-complementation. Proc. Natl Acad. Sci. USA 72, 1254–1257 (1975).
Ullmann, A., Jacob, F. & Monod, J. Characterization by in vitro complementation of a peptide corresponding to an operator-proximal segment of the β-galactosidase structural gene of Escherichia coli. J. Mol. Biol. 24, 339–343 (1967).
Richter, F. et al. Engineering of temperature- and light-switchable Cas9 variants. Nucleic Acids Res. 44, 10003–10014 (2016).
Fonfara, I. et al. Phylogeny of Cas9 determines functional exchangeability of dual-RNA and Cas9 among orthologous type II CRISPR-Cas systems. Nucleic Acids Res. 42, 2577–2590 (2014).
Zuo, Z. et al. Engineering Pseudomonas putida KT2440 for simultaneous degradation of organophosphates and pyrethroids and its application in bioremediation of soil. Biodegradation 26, 223–233 (2015).
Datlinger, P. et al. Pooled CRISPR screening with single-cell transcriptome readout. Nat. Methods 14, 297 (2017).
Hanahan, D. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 166, 557–580 (1983).
Inoue, H., Nojima, H. & Okayama, H. High efficiency transformation of Escherichia coli with plasmids. Gene 96, 23–28 (1990).
McClelland, M., Hanish, J., Nelson, M. & Patel, Y. KGB: a single buffer for all restriction endonucleases. Nucleic Acids Res. 16, 364 (1988).
Labun, K. et al. Accurate analysis of genuine CRISPR editing events with ampliCan. Genome Res. 29, 843–847 (2019).
Acknowledgements
We thank F. Richter for plasmids (pRob12, pCDF_dCas9 and pACYC_sgRNA) and help with setting up the bacterial survival assay, K. Schmidt, S. Ressel and J. Boshart for technical help, M. Ugolini for help with flow cytometry, and members of the Charpentier laboratory for valuable discussions and critical reading of the manuscript. We thank S. Klages and N. Mages from the sequencing core facility of the Max Planck Institute for Molecular Genetics for help with amplicon sequencing. We thank the Alexander von Humboldt Foundation (AvH Professorship to E.C.), the Helmholtz Association (E.C.), the Max Planck Society (E.C.), the Max Planck Foundation (E.C.), the Göran Gustafsson Foundation (Göran Gustafsson Prize to E.C.), the Kempe Foundation (E.C.), Umeå University (E.C.) and the Swedish Research Council (E.C.) for funding this study. K.C. was a fellow of the Austrian Doctoral Program in RNA Biology.
Author information
Authors and Affiliations
Contributions
E.C. oversaw the study. M. Bratovič, I.F., K.C., M.Boettcher and E.C. designed experiments. M. Bratovič and I.F. performed all in vitro experiments and bacterial survival assays, and analyzed all data. M. Bratovič and K.C. performed experiments in eukaryotic cells. S.B. and B.T. performed library preparation and amplicon sequencing. E.J.C.G. and T.J.S. analyzed the sequencing data and performed statistical analysis. M. Bratovič, I.F. and E.C. wrote the manuscript, which all other authors commented on and approved.
Corresponding author
Ethics declarations
Competing interests
E.C. is cofounder of ERS Genomics and CRISPR Therapeutics and is a member of the scientific advisory board of CRISPR Therapeutics. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–9, Figs. 1–14 and Note 1
Rights and permissions
About this article
Cite this article
Bratovič, M., Fonfara, I., Chylinski, K. et al. Bridge helix arginines play a critical role in Cas9 sensitivity to mismatches. Nat Chem Biol 16, 587–595 (2020). https://doi.org/10.1038/s41589-020-0490-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0490-4
This article is cited by
-
Engineering Cas9: next generation of genomic editors
Applied Microbiology and Biotechnology (2024)
-
Sniper2L is a high-fidelity Cas9 variant with high activity
Nature Chemical Biology (2023)
-
Genome-wide CRISPR off-target prediction and optimization using RNA-DNA interaction fingerprints
Nature Communications (2023)
-
A cleavage rule for selection of increased-fidelity SpCas9 variants with high efficiency and no detectable off-targets
Nature Communications (2023)
-
CRISPR/Cas9 therapeutics: progress and prospects
Signal Transduction and Targeted Therapy (2023)